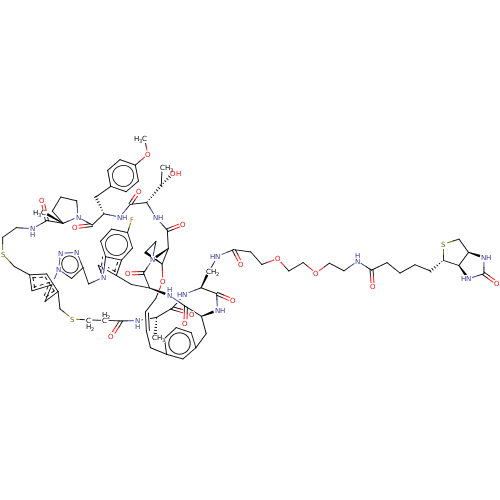
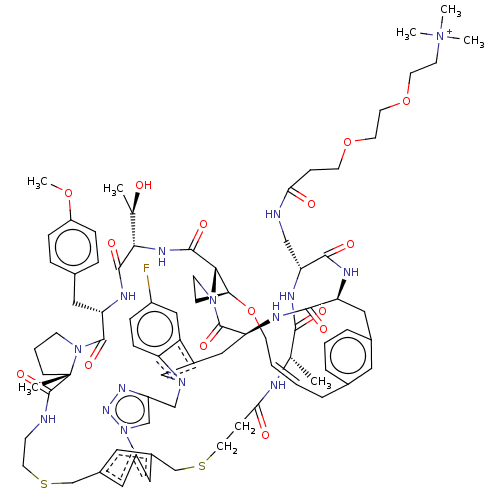
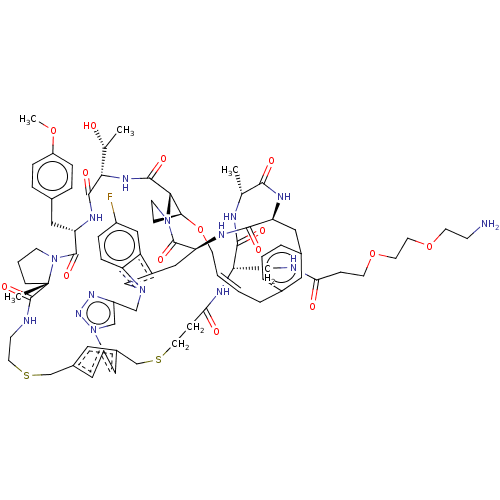
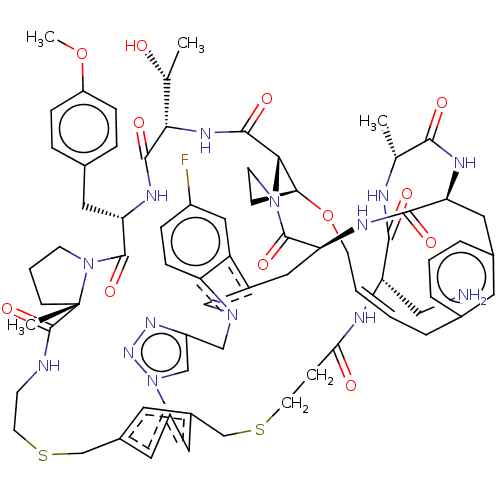
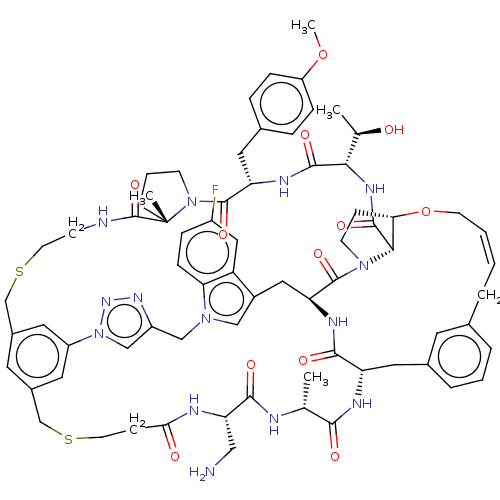
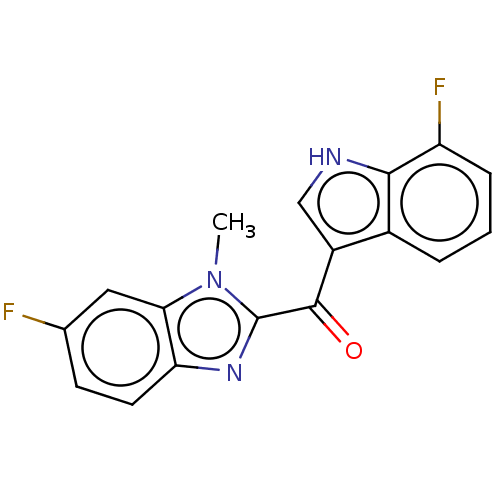
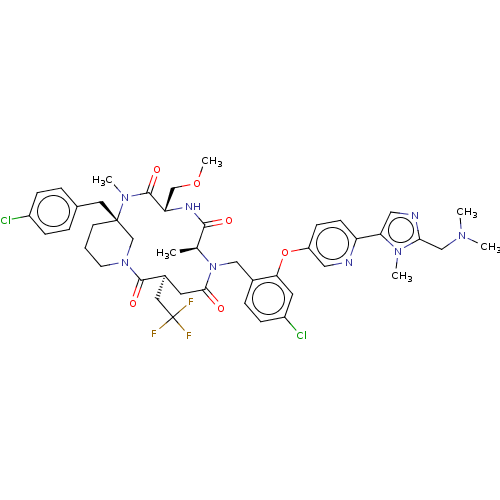
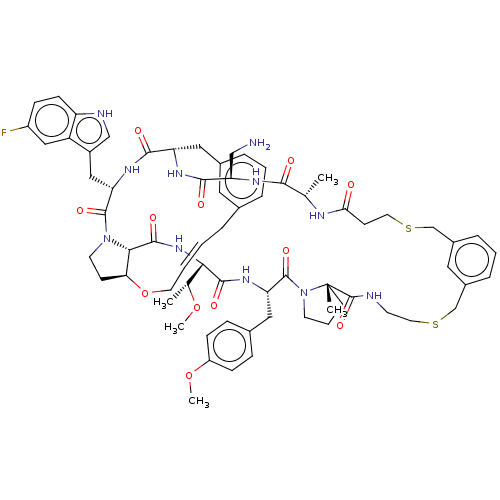
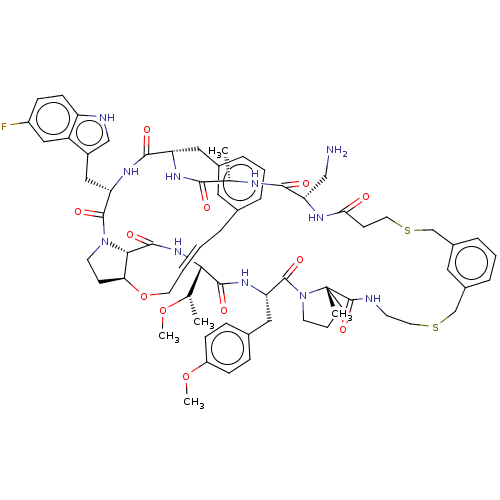
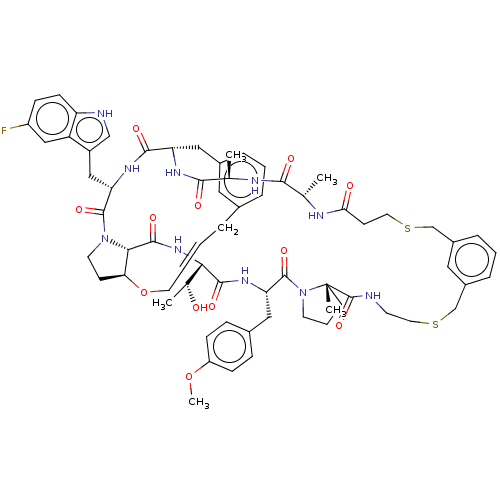
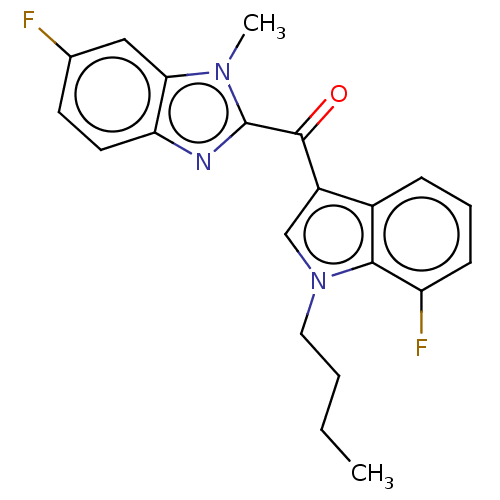
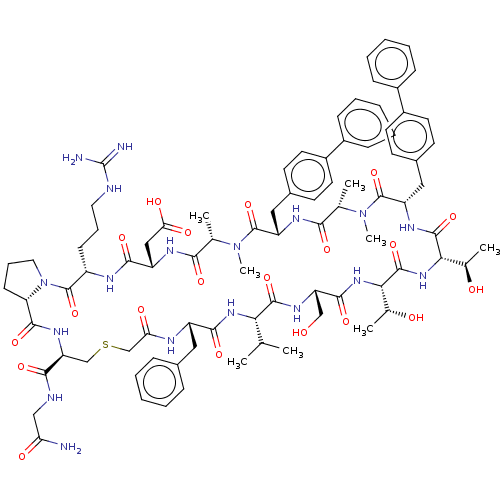
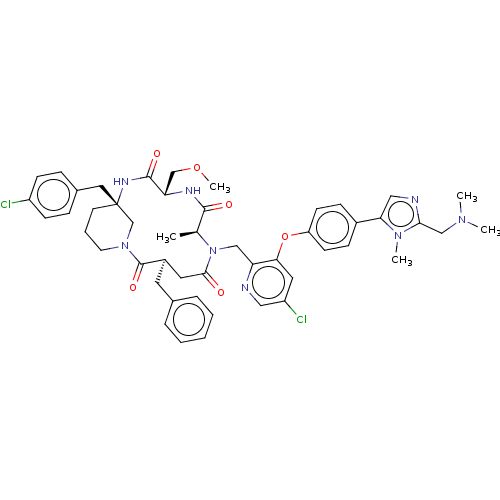
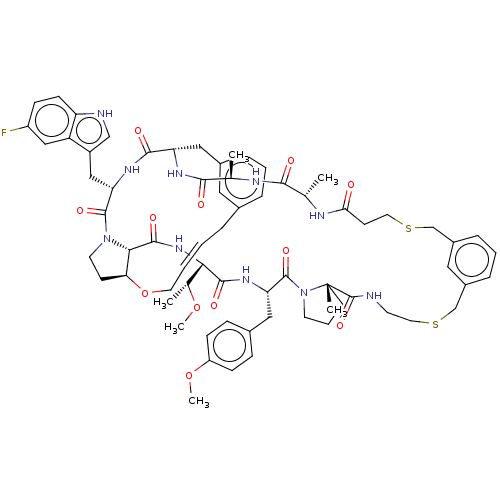
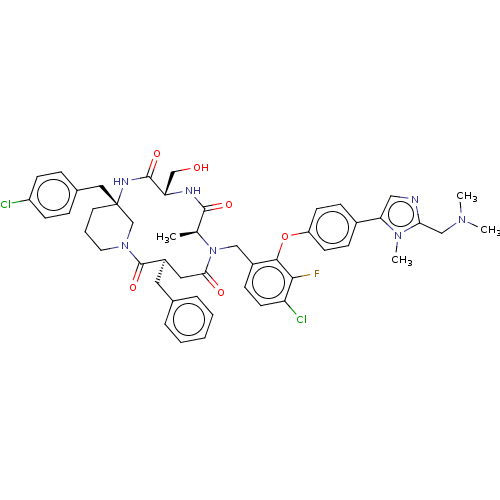
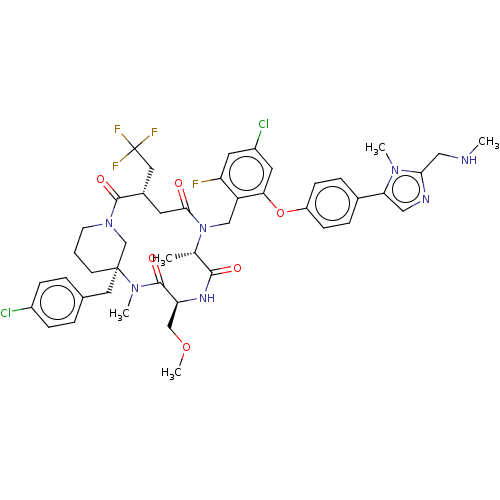
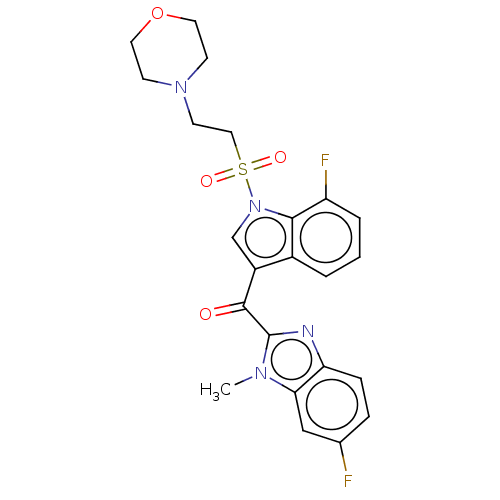
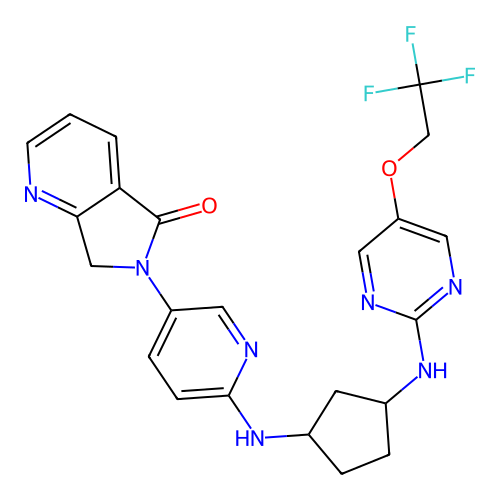
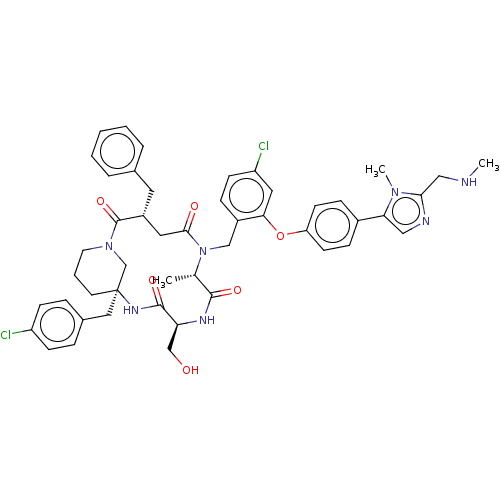
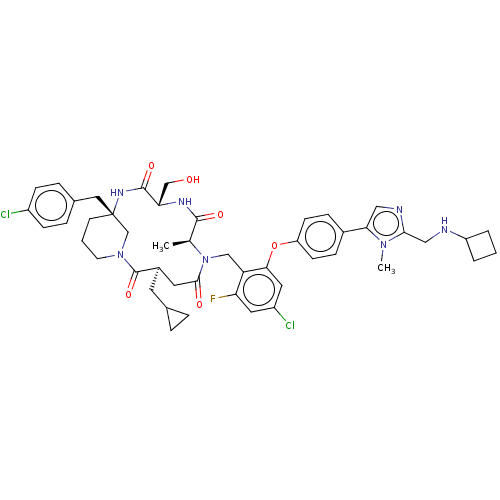
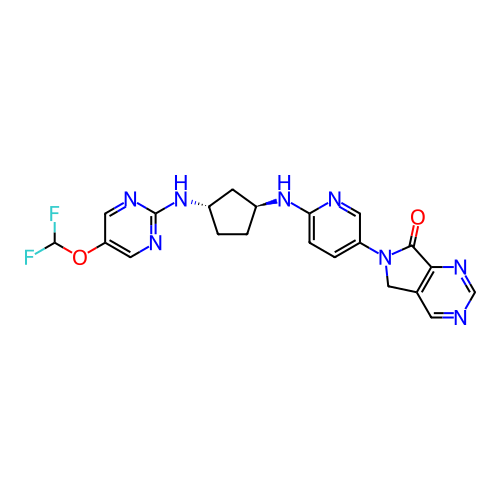
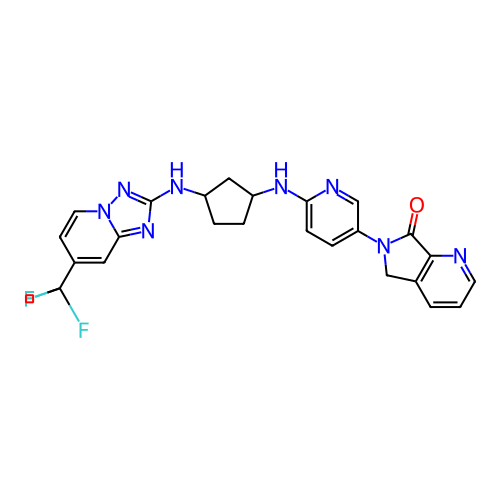
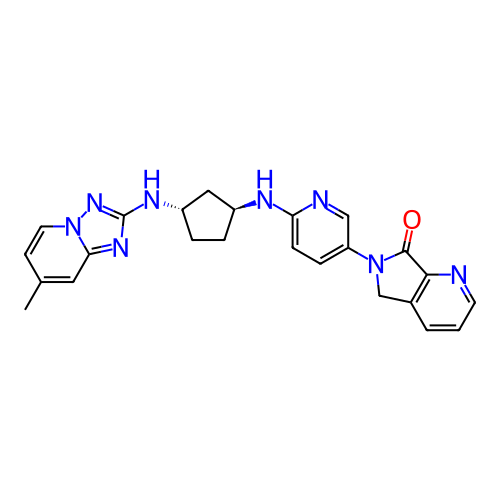
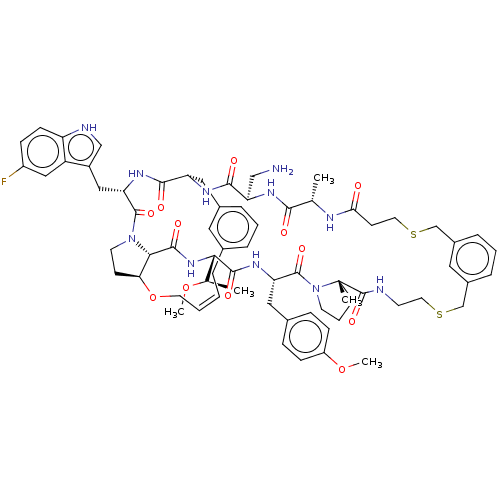
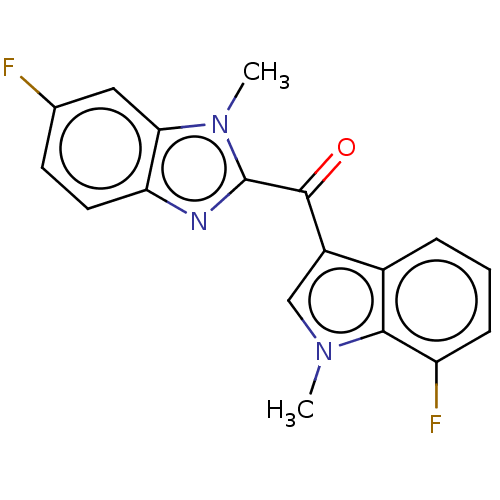
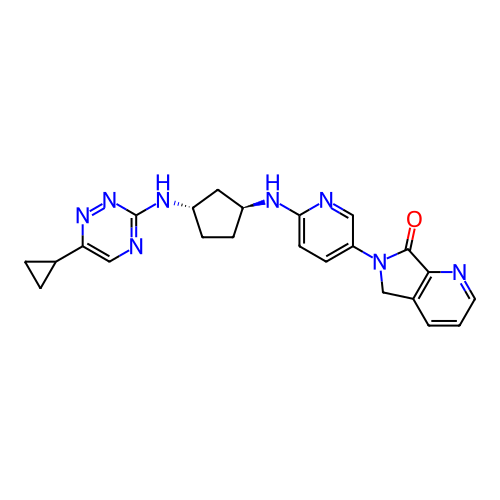
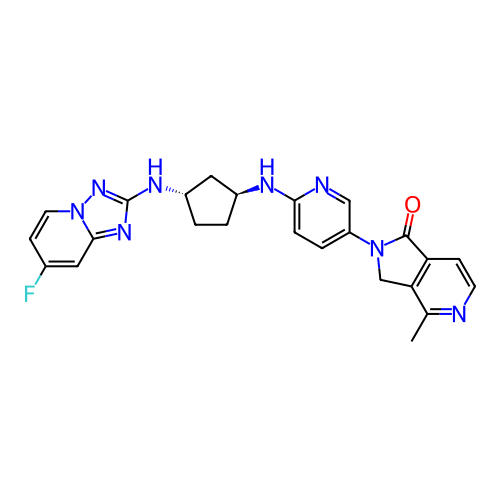
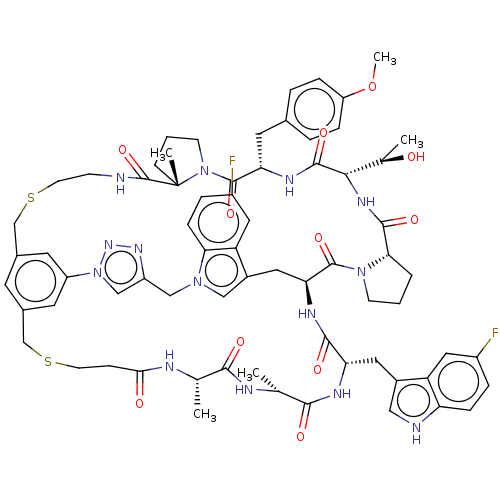
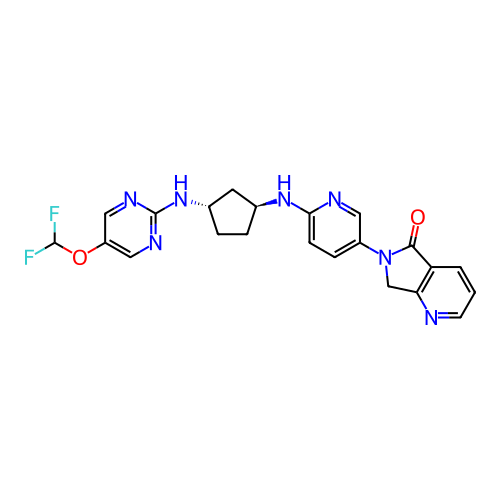
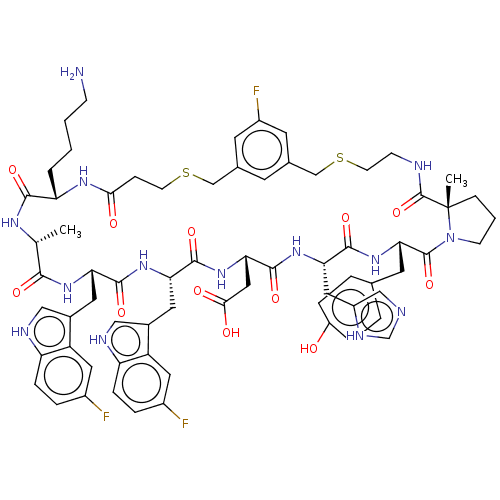
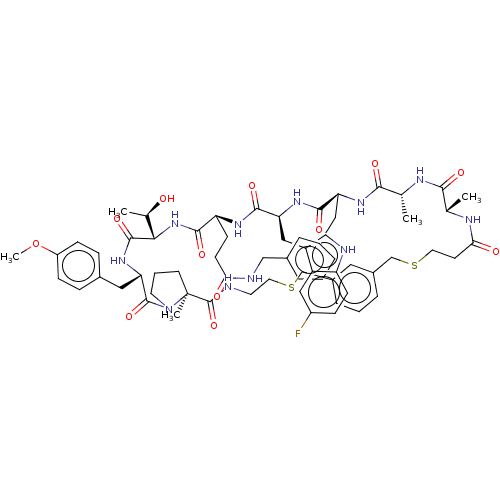
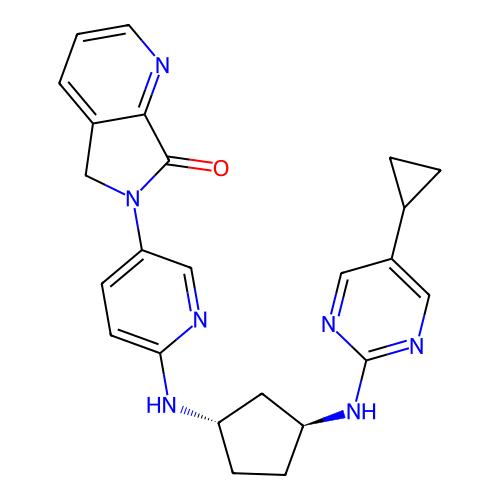
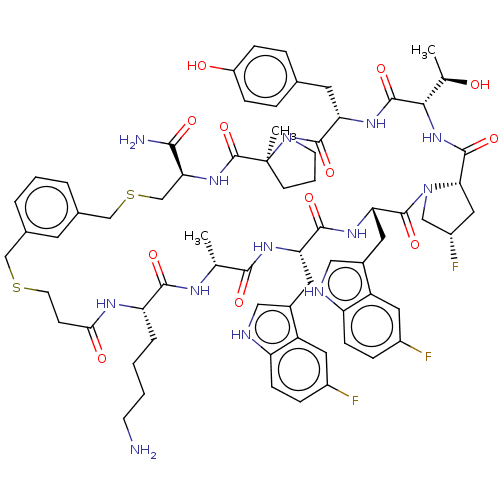
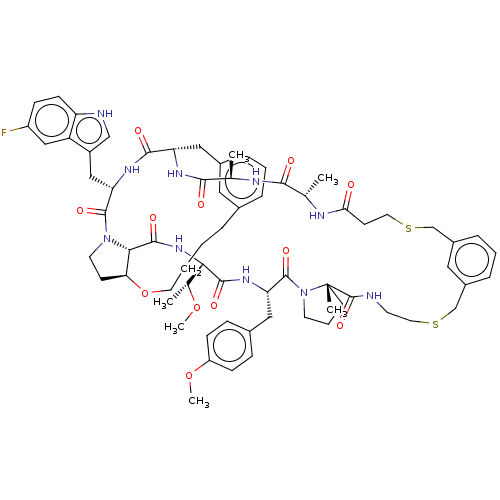
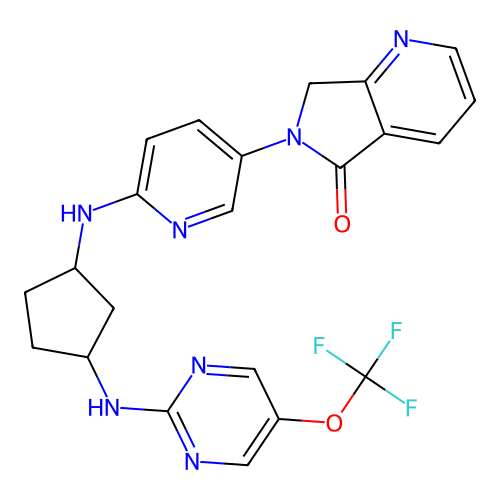
Affinity DataKi: 0.000600nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET ultra assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.000930nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET ultra assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.00239nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET ultra assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.00736nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET plus assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.00813nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET ultra assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.00826nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET plus assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.00940nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET ultra assayMore data for this Ligand-Target Pair
Affinity DataKi: <0.0100nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET plus assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0114nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET plus assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0370nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET plus assayMore data for this Ligand-Target Pair
Affinity DataKi: >0.129nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: >0.129nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: >0.129nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: >0.129nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: >0.129nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataEC50: 0.150nMAssay Description:Modulation of PCSK9 in human HepG2 cells assessed as reduction in PCSK9 expression incubated for 24 hrs prior to compound washout followed by compoun...More data for this Ligand-Target Pair
Affinity DataIC50: 0.152nMAssay Description:Inhibition of human PCSK9 by Alexa Fluor 647 staining based TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.160nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET plus assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.230nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET plus assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.25nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET plus assayMore data for this Ligand-Target Pair
Affinity DataEC50: 0.280nMAssay Description:Modulation of PCSK9 in human HepG2 cells assessed as reduction in PCSK9 expression incubated for 24 hrs prior to compound washout followed by compoun...More data for this Ligand-Target Pair
Affinity DataKd: 0.300nMAssay Description:Binding affinity to human PCSK9More data for this Ligand-Target Pair
Affinity DataIC50: 0.327nMAssay Description:Inhibition of human PCSK9 by Alexa Fluor 647 staining based TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.390nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET plus assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.430nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.438nMAssay Description:Inhibition of human PCSK9 by Alexa Fluor 647 staining based TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.450nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.480nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.570nMAssay Description:Inhibition of human PCSK9 by Alexa Fluor 647 staining based TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataEC50: 0.690nMAssay Description:Modulation of PCSK9 in human HepG2 cells assessed as reduction in PCSK9 expression incubated for 24 hrs prior to compound washout followed by compoun...More data for this Ligand-Target Pair
Affinity DataIC50: 0.850nMAssay Description:Inhibition of human PCSK9 by Alexa Fluor 647 staining based TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.930nMAssay Description:Inhibition of human PCSK9 by Alexa Fluor 647 staining based TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataEC50: 1.20nMAssay Description:Modulation of PCSK9 in human HepG2 cells assessed as reduction in PCSK9 expression incubated for 24 hrs prior to compound washout followed by compoun...More data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:Inhibition of AlexaFluor647-tagged cyclic peptide binding to avi-tagged-biotinylated human PCSK9 measured after 2 hrs by Lance Streptavidin Europium ...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:Inhibition of AlexaFluor647-tagged cyclic peptide binding to avi-tagged-biotinylated human PCSK9 measured after 2 hrs by Lance Streptavidin Europium ...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:Inhibition of AlexaFluor647-tagged cyclic peptide binding to avi-tagged-biotinylated human PCSK9 measured after 2 hrs by Lance Streptavidin Europium ...More data for this Ligand-Target Pair
Affinity DataKi: 2.10nMAssay Description:Inhibition of AlexaFluor647-tagged cyclic peptide binding to avi-tagged-biotinylated human PCSK9 measured after 2 hrs by Lance Streptavidin Europium ...More data for this Ligand-Target Pair
Affinity DataKi: 2.20nMAssay Description:Inhibition of human PCSK9 using Alexa Fluor 674 as substrate incubated for 2 hrs by FRET assayMore data for this Ligand-Target Pair