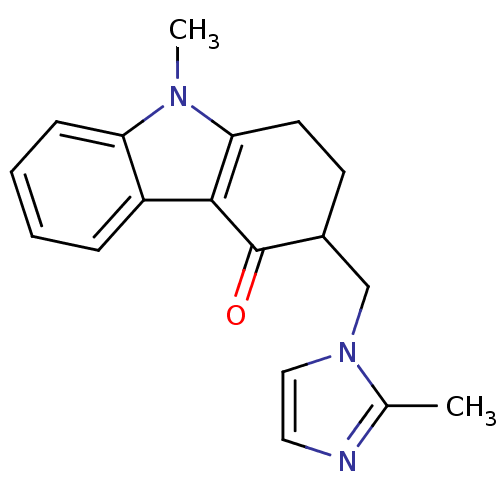
BDBM85330 CAS_68647::NSC_68647::ONDANSETRON::Ondansetron hydrochloride
SMILES Cc1nccn1CC2CCc3c(c4ccccc4n3C)C2=O
InChI Key InChIKey=FELGMEQIXOGIFQ-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 117 hits for monomerid = 85330
Found 117 hits for monomerid = 85330
Affinity DataIC50: 0.0900nMAssay Description:Antagonist activity at human 5HT3A receptor expressed in xenopus oocytes assessed as inhibition of 5HT-induced effect by electrophysiological methodMore data for this Ligand-Target Pair
Affinity DataIC50: 0.160nMAssay Description:Binding affinity to rat 5HT3A receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.770nMAssay Description:Compound was evaluated for its in vitro affinity at serotonergic 5-hydroxytryptamine 3 receptor by radioligand binding assay, using [3H]-LY 278584 in...More data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:In vitro affinity for 5-hydroxytryptamine 3 (5-HT3) receptor by displacement of [3H]BRL-43694 from rat entorhinal cortexMore data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Binding affinity for 5-hydroxytryptamine 3 receptor from rat cortex using [3H]BRL-43694 as radioligandMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Binding affinity against 5-hydroxytryptamine 3 receptor in rat cortical membrane using [3H]GR-65630 as radioligandMore data for this Ligand-Target Pair
Affinity DataIC50: 1.54nMAssay Description:Concentration required to inhibit the binding of radioligand [3H]GR-65630 to serotonin 5-hydroxytryptamine-3 receptor (5-HT 3 receptor)in rat brain c...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:The binding affinity was measured on 5-hydroxytryptamine 3 receptor using [3H]GR-65630 as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Displacement of [3H]GR-65630 binding to 5-hydroxytryptamine 3 receptor in rat brain cortical membranesMore data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Displacement of [3H]GR-65630 binding to 5-hydroxytryptamine 3 receptor in rat brain cortical membranesMore data for this Ligand-Target Pair
Affinity DataIC50: 1.90nMAssay Description:Compound was evaluated for the displacement of [3H]Q-ICS-205-930 from 5-HT3 recognition sites in rat brain membranesMore data for this Ligand-Target Pair
Affinity DataIC50: 1.90nMAssay Description:Compound was evaluated for the displacement of [3H]-Q-ICS 205-930 binding to 5-hydroxytryptamine 3 recognition sites in rat brain membranesMore data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:pKi value for inhibition of [3H]LY-278584 binding to 5-hydroxytryptamine 3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:Binding affinity to 5-HT2B receptor (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:Inhibitory activity against 5-hydroxytryptamine 3 receptor in rat cortical membranes using [3H]-1-Methyl-1H-indazole-3-carboxylic acid (8-methyl-8-az...More data for this Ligand-Target Pair
Affinity DataKi: 2.80nMAssay Description:Displacement of [3H]granisetron from human 5HT3A receptor expressed in HEK293 cells by filter binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.20nMAssay Description:Binding affinity against 5-hydroxytryptamine 3 (5-HT3) receptor in rat brain cortical membranes using radioligand [3H]quipazineMore data for this Ligand-Target Pair
Affinity DataKi: 3.20nMAssay Description:Binding affinity to 5-hydroxytryptamine 3 receptor was determined in rat cerebro cortical membranes using [3H]quipazine.More data for this Ligand-Target Pair
Affinity DataKi: 3.30nMAssay Description:Compound was evaluated for its binding affinity for 5-hydroxytryptamine 3 receptor by measuring displacement [3H]GR-65630 in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 3.30nMAssay Description:Binding affinity to 5-hydroxytryptamine 3 receptor using [3H]GR-65630 as radioligand in rat cortexMore data for this Ligand-Target Pair
Affinity DataKi: 3.31nMAssay Description:Binding affinity to 5-hydroxytryptamine 3 receptor using [3H]GR-65630 as radioligand in rat cortexMore data for this Ligand-Target Pair
Affinity DataKi: 3.40nMAssay Description:Binding affinity to human HT3A receptorMore data for this Ligand-Target Pair
Target5-hydroxytryptamine receptor 3A(Mouse)
Institut De Recherches Servier
Curated by PDSP Ki Database
Institut De Recherches Servier
Curated by PDSP Ki Database
Affinity DataKi: 6.20nMAssay Description:Displacement of the 5-hydroxytryptamine 3 receptor ligand [3H]GR-65630 from rat brain cortical membranes.More data for this Ligand-Target Pair
Affinity DataKi: 6.5nMAssay Description:Displacement of binding of [3H]-BRL 43694 to 5-hydroxytryptamine 3 receptor in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 6.80nMAssay Description:Binding affinity against 5-hydroxytryptamine 3 (5-HT3) receptor from rat cortical homogenate using [3H]zacopride as radioligandMore data for this Ligand-Target Pair
Affinity DataIC50: 7.10nMAssay Description:Binding affinity was evaluated in vitro by displacement of [3H]zacopride radioligand from 5-hydroxytryptamine 3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 7.60nMAssay Description:Binding affinity against radioligand [3H]quipazine labeled 5-hydroxytryptamine 3 receptor sites in neuroblastoma-glioma (NG108-15) cells.More data for this Ligand-Target Pair
Affinity DataKi: 7.60nMAssay Description:Compound was evaluated for its ability to displace [3H]quipazine binding to 5-hydroxytryptamine 3 receptor sites in NG 108-15. More data for this Ligand-Target Pair
Target5-hydroxytryptamine receptor 3A(Guinea pig)
Syntex Discovery Research
Curated by PDSP Ki Database
Syntex Discovery Research
Curated by PDSP Ki Database
Target5-hydroxytryptamine receptor 3A(Mouse)
Institut De Recherches Servier
Curated by PDSP Ki Database
Institut De Recherches Servier
Curated by PDSP Ki Database
Affinity DataKi: 12nMAssay Description:The binding affinity was measured for 5-hydroxytryptamine 3 receptor on NG 108-15 cell line of mouse neuroblastoma-glioma cells in presence of [3H]5 ...More data for this Ligand-Target Pair
Affinity DataKd: 13nMAssay Description:Binding affinity towards 5-hydroxytryptamine 3 receptor was determined by using [3H]-ICS 205-930 as radioligand in mouse N1E 115 cellsMore data for this Ligand-Target Pair
Target5-hydroxytryptamine receptor 3A(Mouse)
Institut De Recherches Servier
Curated by PDSP Ki Database
Institut De Recherches Servier
Curated by PDSP Ki Database
Affinity DataKi: 16nMAssay Description:In vitro displacement of [3H]ICS-205-930 from 5-hydroxytryptamine 3 receptor in cultured NG-108-15 rat glioma cellsMore data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:In vitro binding affinity for the 5-hydroxytryptamine 3 receptor was determined with NG-108-15 mouse neuroblastoma-glioma cellsMore data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Binding affinity towards 5-hydroxytryptamine 3 receptor by displacement of [3H]2 in Neuroblastoma-Glioma NG-108-15 cellsMore data for this Ligand-Target Pair
Affinity DataKd: 20nMAssay Description:Compound was evaluated for binding to 5-hydroxytryptamine 3 receptor in N1E cells using [3H]- -Tropisetron as radioligandMore data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of human MATE1-mediated [14]-metformin uptake expressed in HEK293 cells after 1.5 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 46nMAssay Description:Inhibition of [3H]BRL-43694 binding to rat 5-hydroxytryptamine 3 receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Inhibition of human MATE1 expressed in HEK-293-Flp-In cells incubated for 3 mins by ASP+ substrate uptake assayMore data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:Inhibition of human MATE2K-mediated ASP+ uptake expressed in HEK293 cells up to 500 uM after 1.5 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:Inhibition of human MATE1-mediated ASP+ uptake expressed in HEK293 cells after 1.5 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Inhibition of human MATE2K-mediated ASP+ uptake expressed in HEK293 cells after 1.5 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 400nMAssay Description:Inhibition of human ERG expressed in HEK cells by patch clamp techniqueMore data for this Ligand-Target Pair
Affinity DataKi: 680nMAssay Description:The binding affinity was measured on sigma receptor using [3H]- (+)-3-PPP as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 680nMAssay Description:The binding affinity was measured on cholecystokinin type B receptor using [3H]- CCK-8 as radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 813nMAssay Description:Inhibition of human ERGMore data for this Ligand-Target Pair
Affinity DataIC50: 813nMAssay Description:Inhibition of human ERG expressed in CHO cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
Affinity DataIC50: 813nMAssay Description:Inhibitory concentration against potassium channel HERGMore data for this Ligand-Target Pair