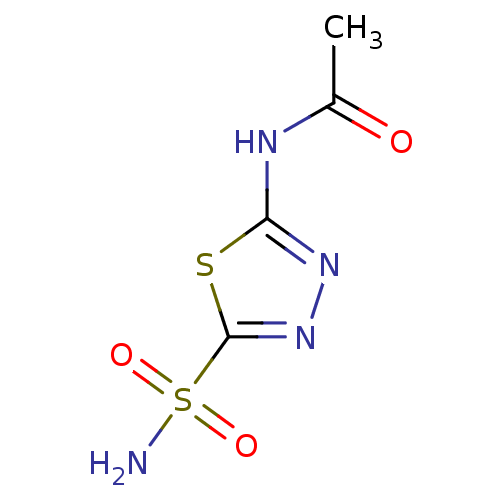
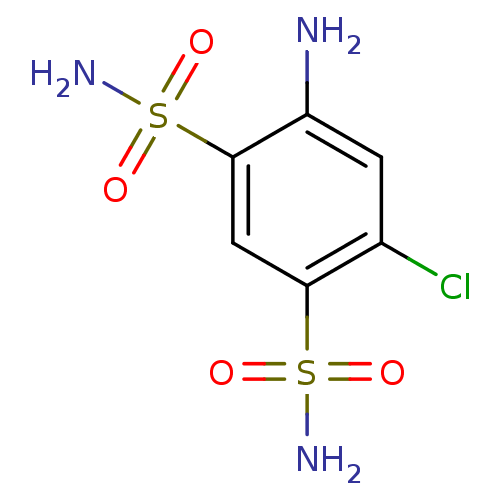
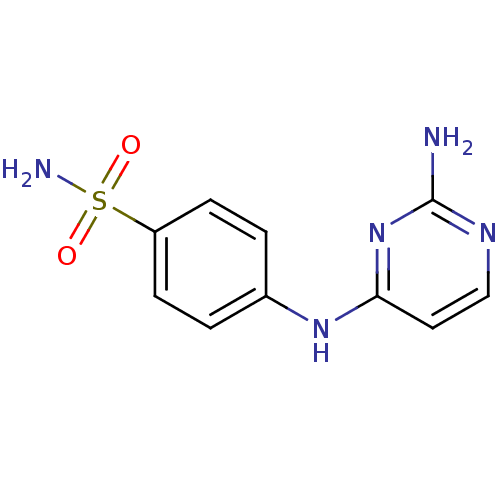
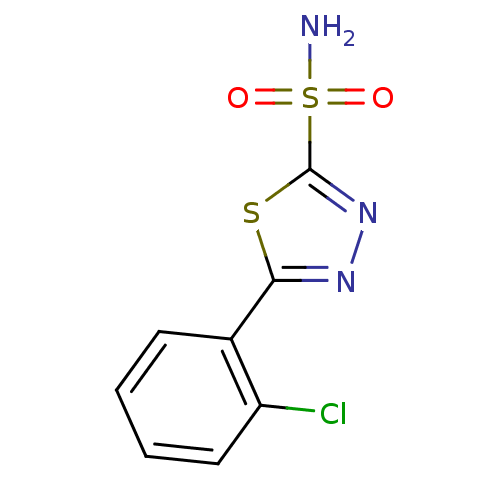
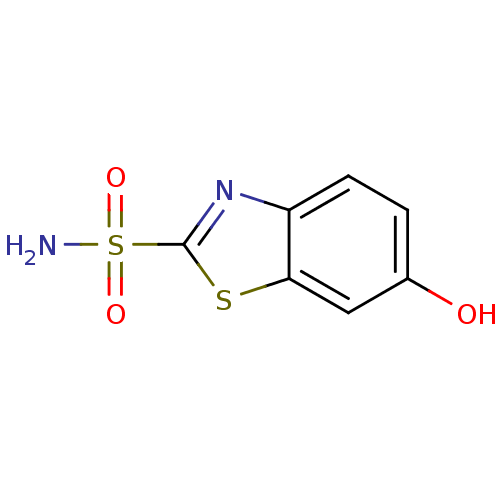
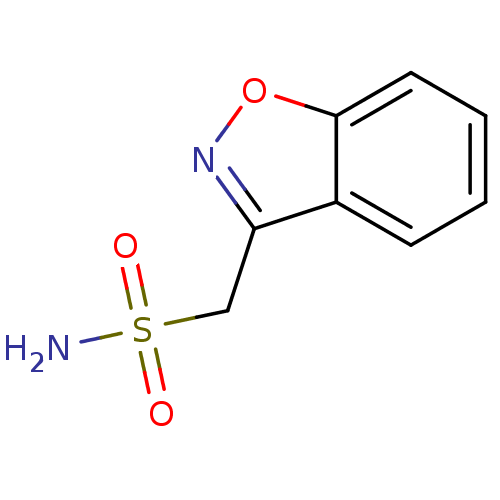
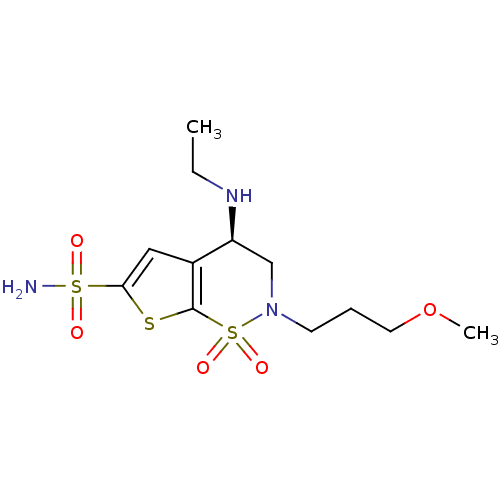
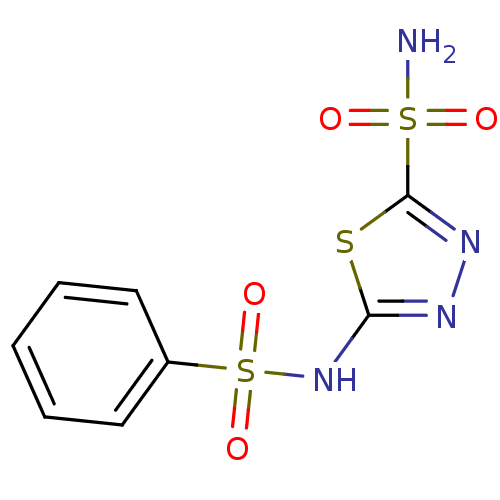
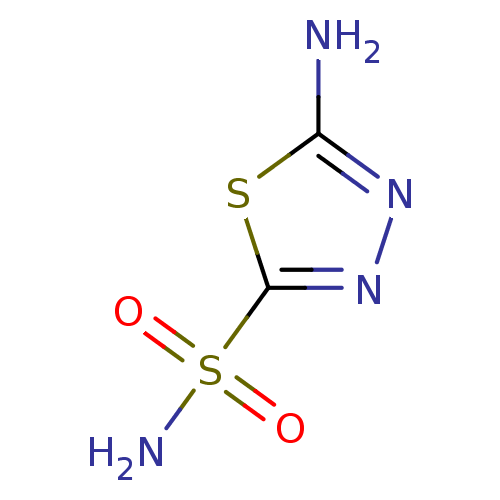
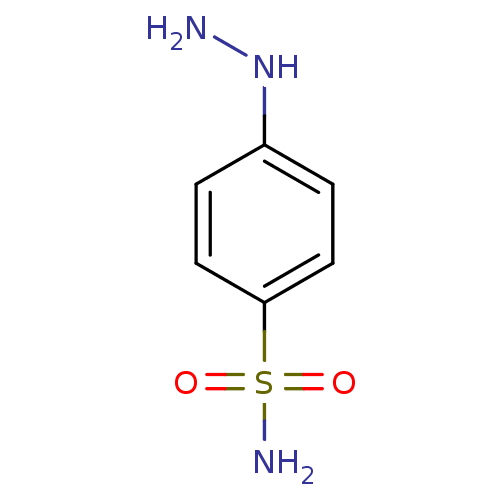
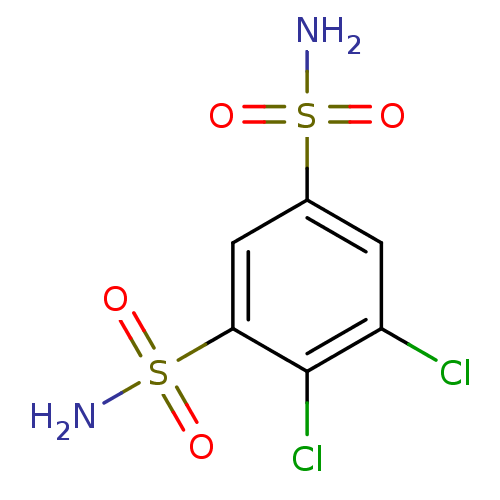
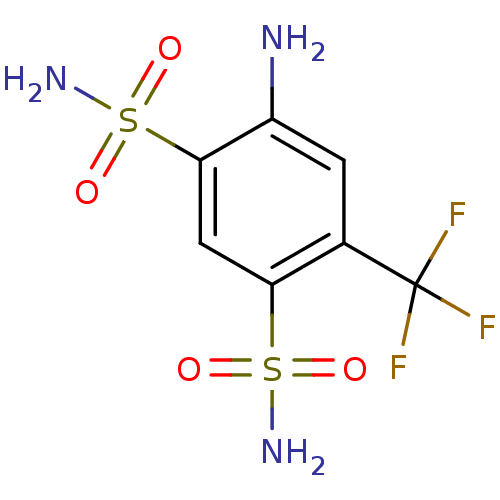
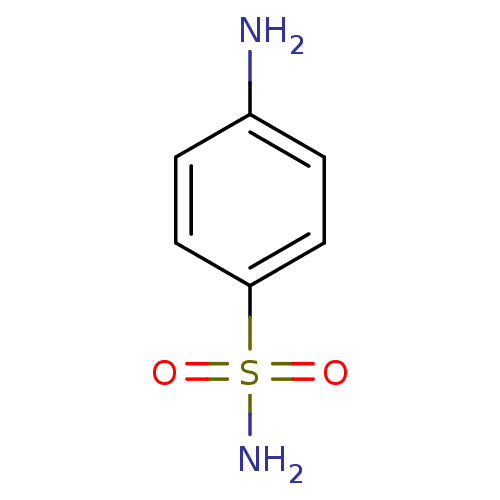
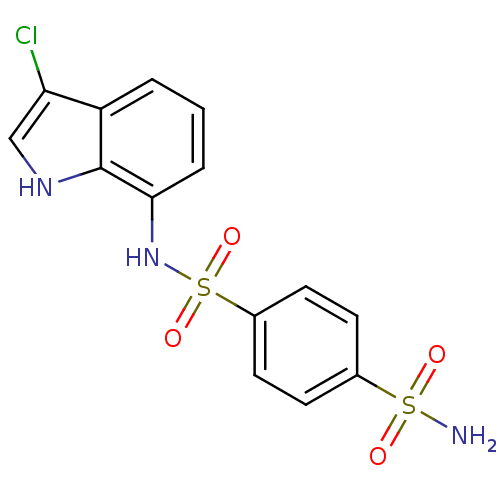
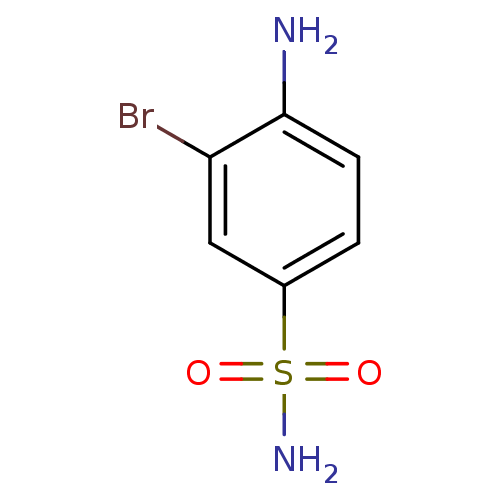
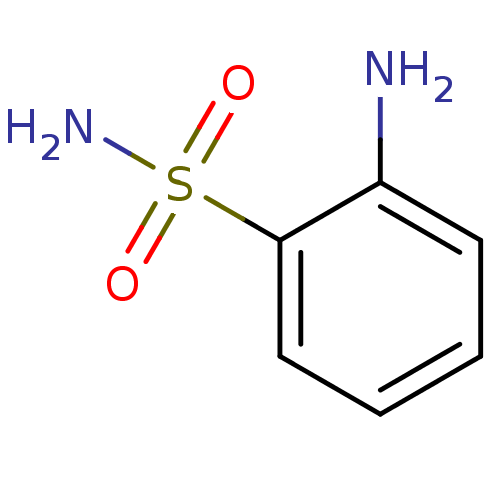
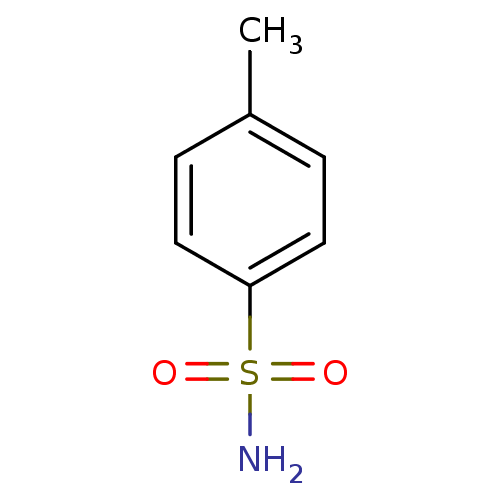
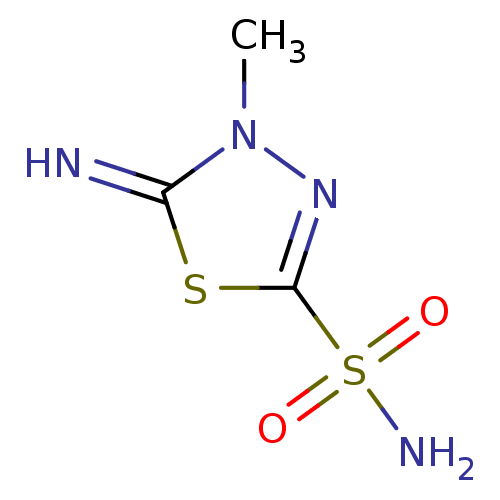
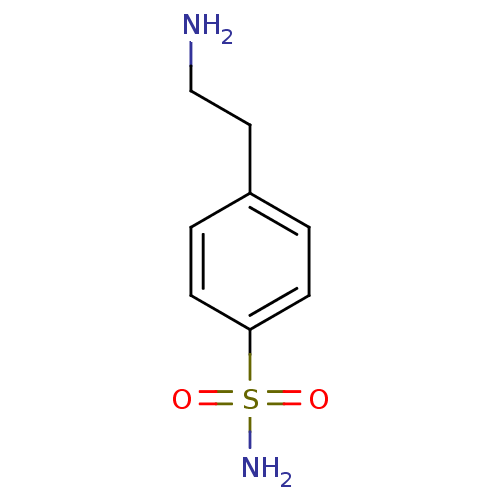
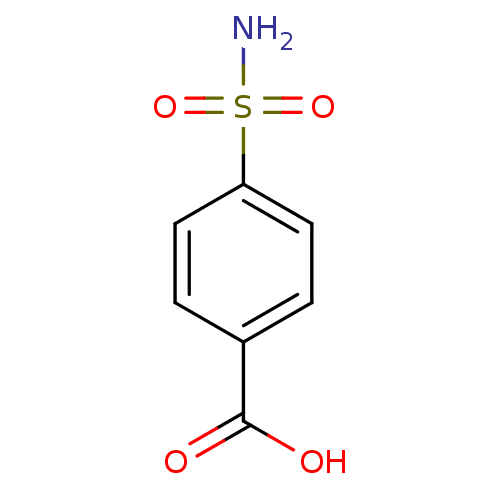
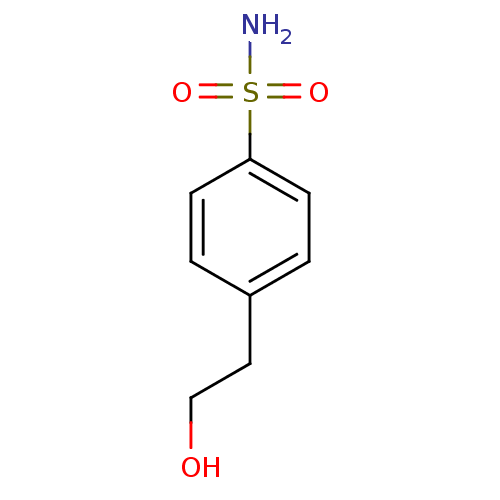
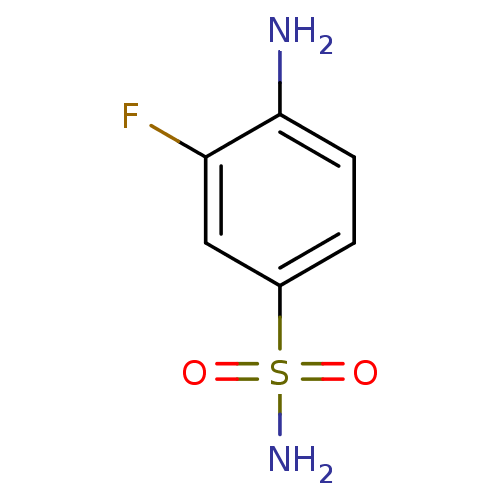
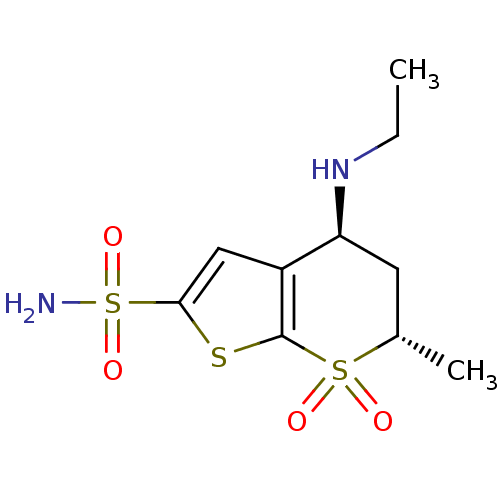
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 20nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
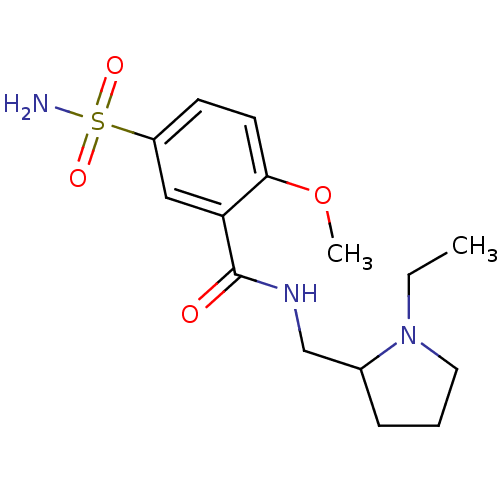
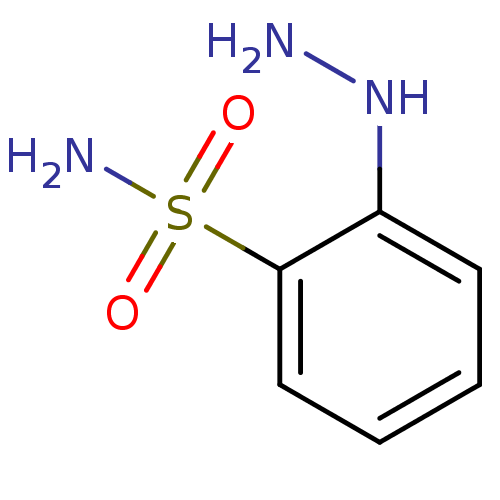
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 41nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
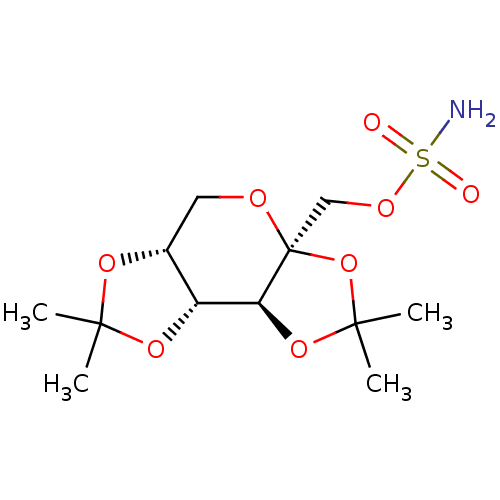
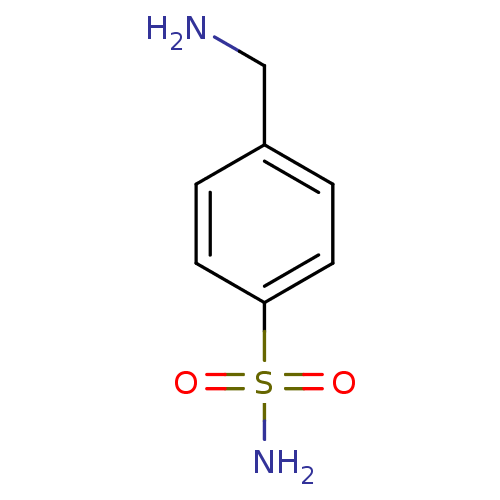
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 72nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
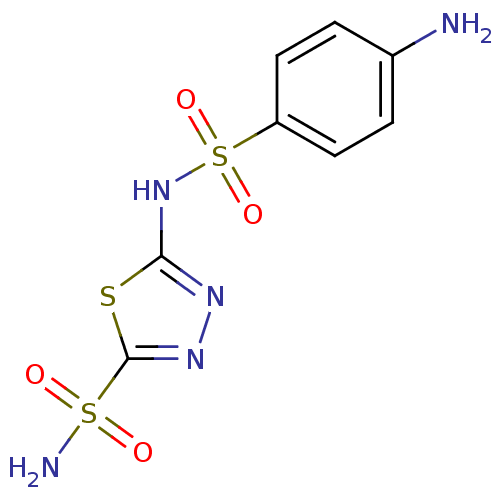
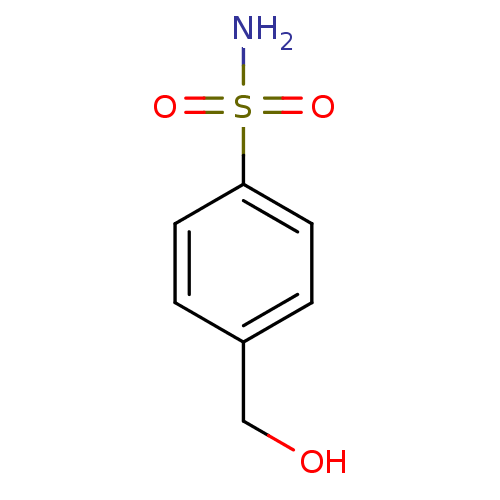
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 84nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 96nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 124nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 155nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 171nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 174nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 179nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 205nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 213nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 218nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 246nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 287nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 310nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 334nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 363nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 395nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 402nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 431nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 434nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 463nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 465nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 473nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 494nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 504nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 562nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 830nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 870nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 1.04E+3nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 1.08E+3nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 1.16E+3nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 1.31E+3nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Kochi Medical School
Curated by ChEMBL
Kochi Medical School
Curated by ChEMBL
Affinity DataKi: 4.31E+3nMAssay Description:Inhibition of Helicobacter pylori recombinant CAMore data for this Ligand-Target Pair