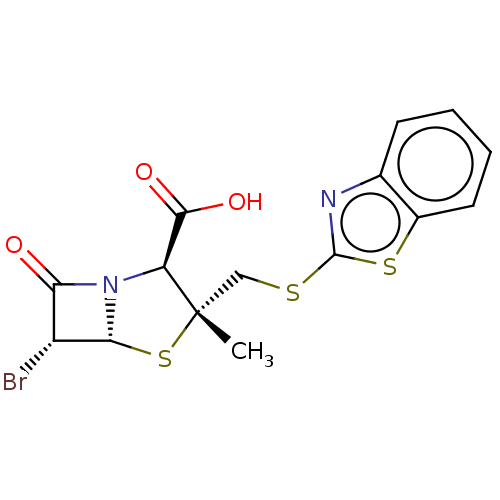
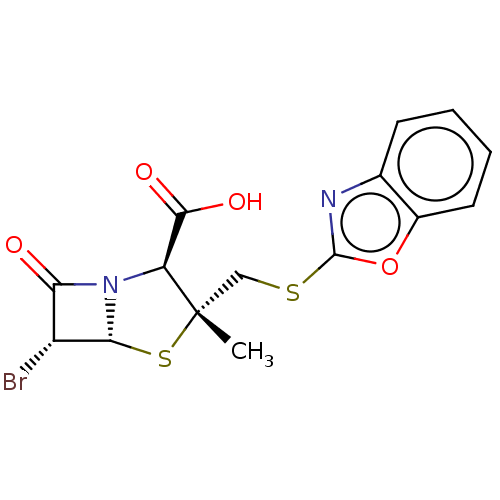
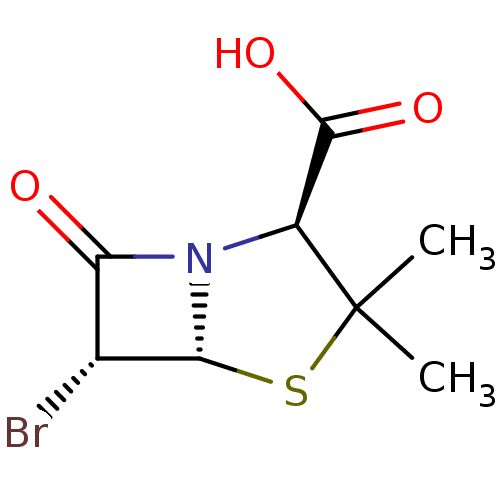
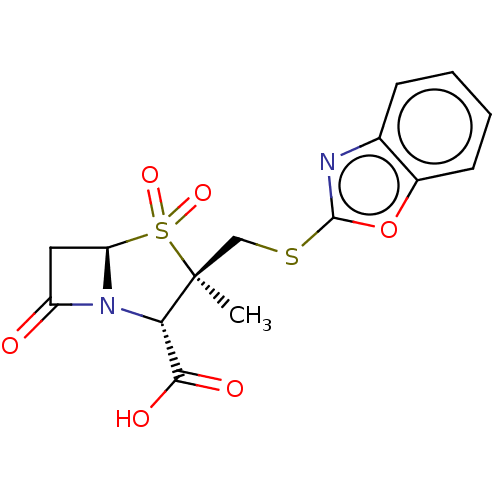
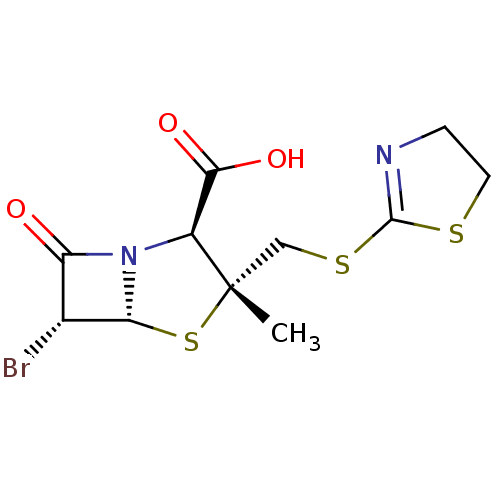
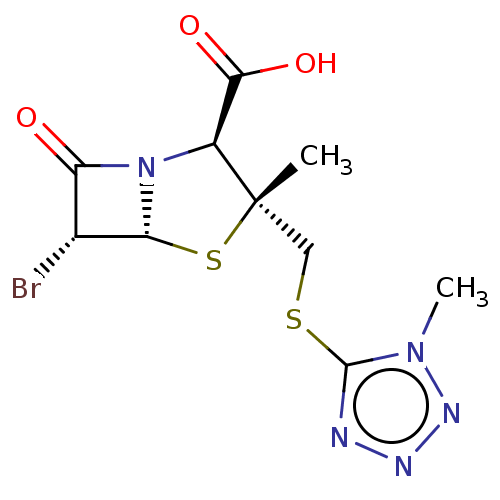
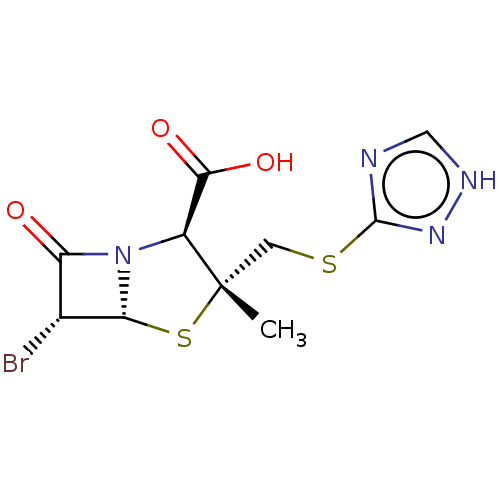
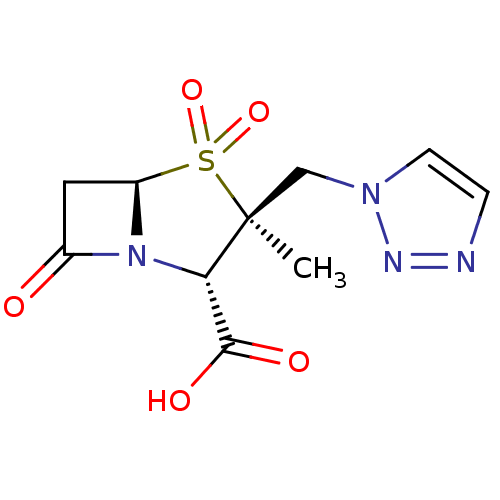
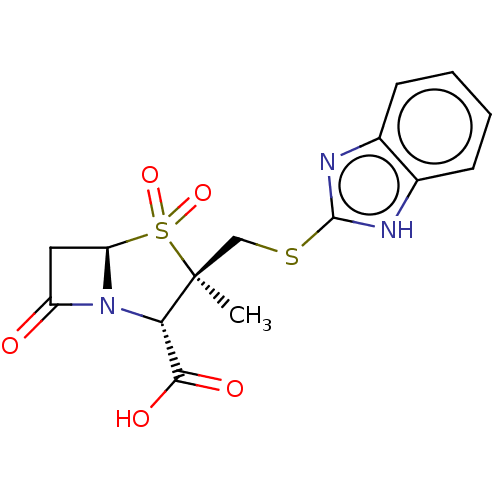
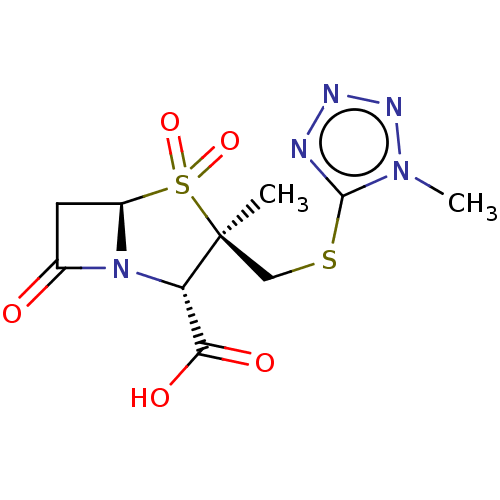
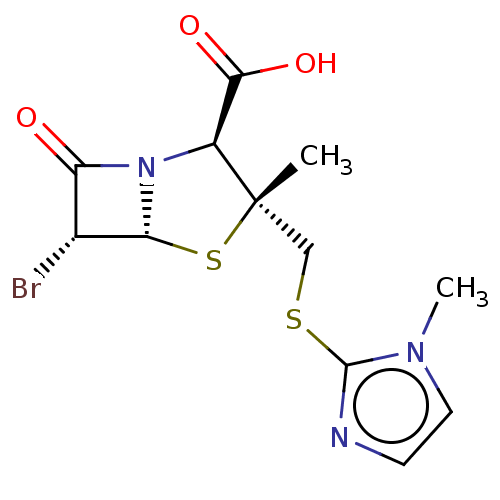
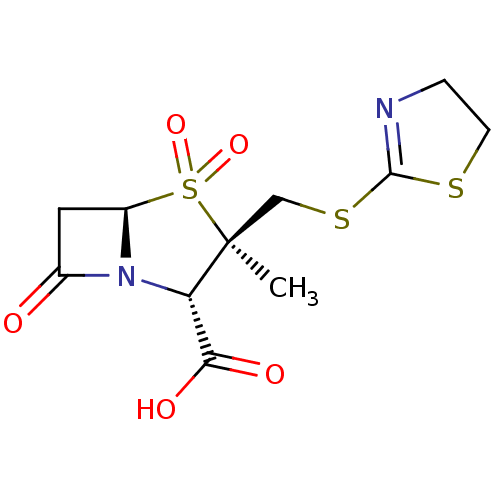
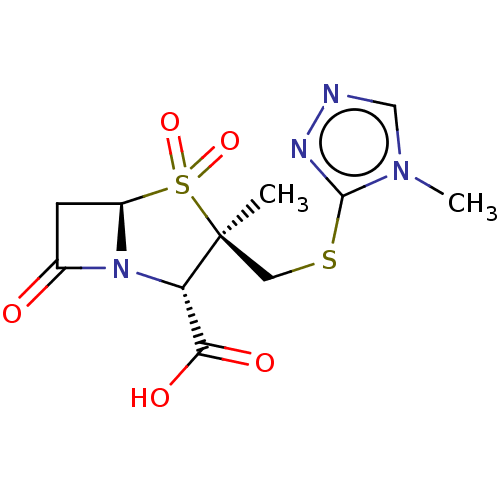
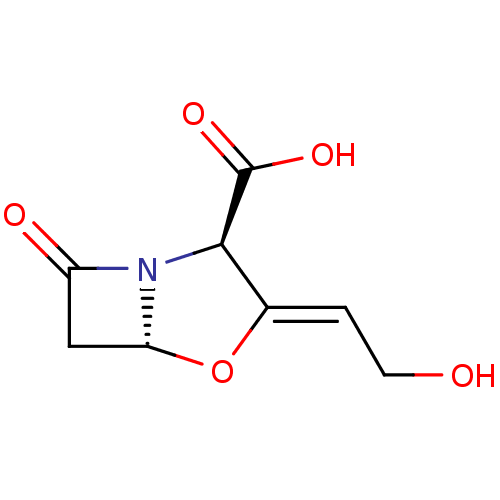
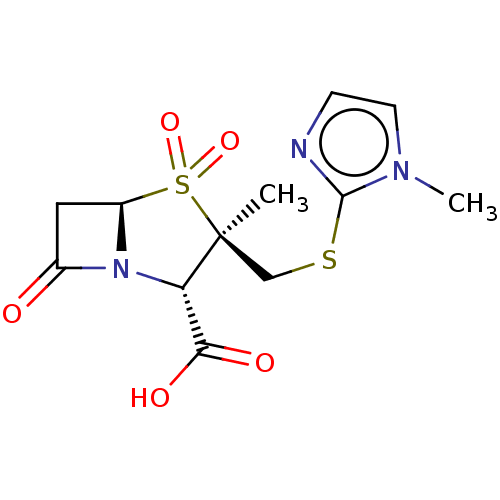
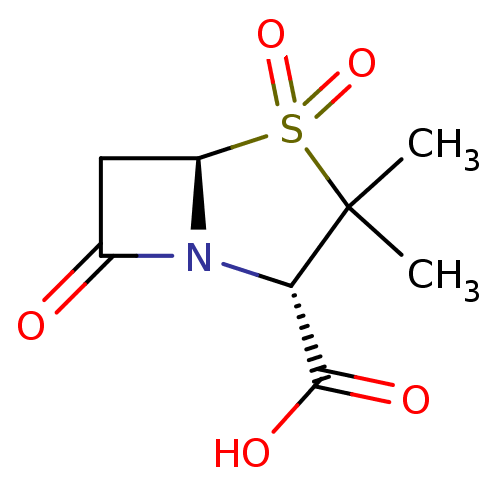
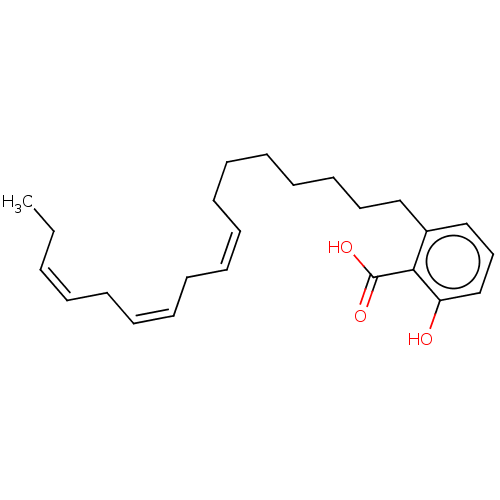
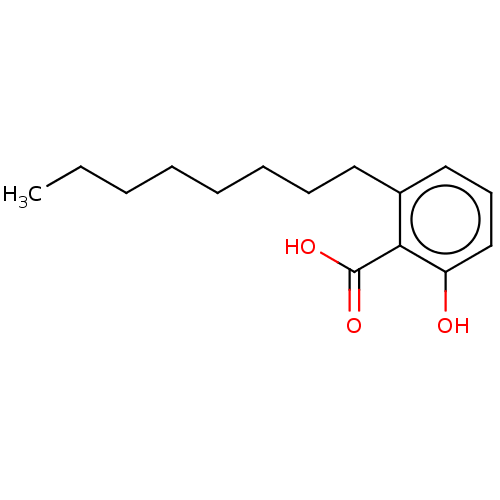
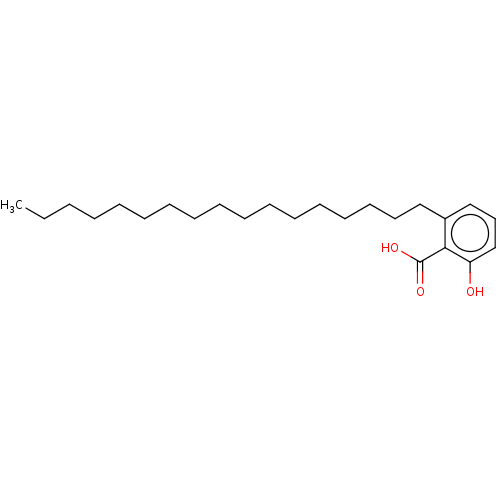
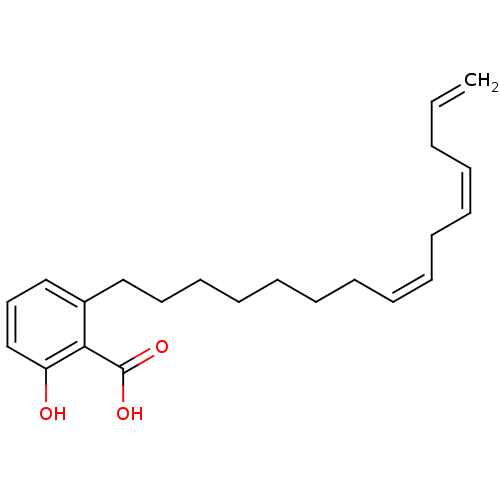
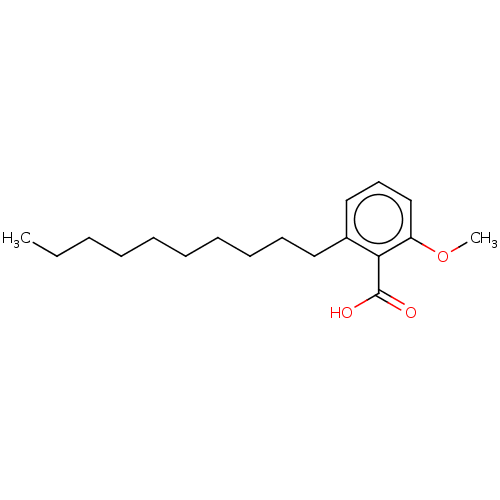
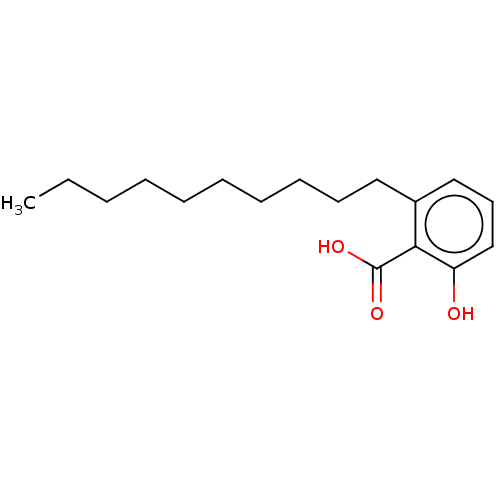
Affinity DataIC50: 2.25nMAssay Description:Inhibitory activity against Proteus vulgaris HJ33C Class-I beta-lactamase enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 2.33nMAssay Description:Tested for the inhibitory activity against Proteus vulgaris HJ33C Class-I beta-lactamase enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 3.57nMAssay Description:Inhibitory activity against Proteus vulgaris HJ33C Class-I beta-lactamase enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 5.23nMAssay Description:Inhibitory activity against Proteus vulgaris HJ33C Class-I beta-lactamase enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 7.55nMAssay Description:Inhibitory activity against Proteus vulgaris HJ33C Class-I beta-lactamase enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 7.61nMAssay Description:Inhibitory activity against Proteus vulgaris HJ33C Class-I beta-lactamase enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 7.91nMAssay Description:Inhibitory activity against Proteus vulgaris HJ33C Class-I beta-lactamase enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Inhibitory activity against Proteus vulgaris HJ33C Class-I beta-lactamase enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 21nMAssay Description:Inhibitory activity against Proteus vulgaris HJ33C Class-I beta-lactamase enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Inhibitory activity against Proteus vulgaris HJ33C Class-I beta-lactamase enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 25.5nMAssay Description:Inhibitory activity against Proteus vulgaris HJ33C Class-I beta-lactamase enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 37.1nMAssay Description:Inhibitory activity against Proteus vulgaris HJ33C Class-I beta-lactamase enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 37.5nMAssay Description:Inhibitory activity against Proteus vulgaris HJ33C Class-I beta-lactamase enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 65.3nMAssay Description:Inhibitory activity against Proteus vulgaris HJ33C Class-I beta-lactamase enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 72.4nMAssay Description:Inhibitory activity against Proteus vulgaris HJ33C Class-I beta-lactamase enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 95.4nMAssay Description:Inhibition of Proteus mirabilis C889 beta-lactamase assessed as inhibition of nitrocefin hydrolysis preincubated for 5 mins by microtiter plate assayMore data for this Ligand-Target Pair
Affinity DataIC50: 342nMAssay Description:Inhibitory activity against Proteus vulgaris HJ33C Class-I beta-lactamase enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 9.54E+4nMAssay Description:Concentration required for its inhibitory activity against Proteus mirabilis C889 class A beta-lactamaseMore data for this Ligand-Target Pair
Affinity DataIC50: 2.99E+5nMAssay Description:Inhibition of Proteus mirabilis C889 beta-lactamase assessed as inhibition of nitrocefin hydrolysis preincubated for 5 mins by microtiter plate assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.55E+7nMAssay Description:Concentration required for its inhibitory activity against Proteus mirabilis C889 class A beta-lactamaseMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+7nMAssay Description:Concentration required for its inhibitory activity against Proteus mirabilis C889 class A beta-lactamaseMore data for this Ligand-Target Pair
Affinity DataIC50: 6.06E+7nMAssay Description:Concentration required for its inhibitory activity against Proteus mirabilis C889 class A beta-lactamaseMore data for this Ligand-Target Pair
Affinity DataIC50: 9.05E+7nMAssay Description:Concentration required for its inhibitory activity against Proteus mirabilis C889 class A beta-lactamaseMore data for this Ligand-Target Pair
Affinity DataIC50: 2.96E+8nMAssay Description:Concentration required for its inhibitory activity against Proteus mirabilis C889 class A beta-lactamaseMore data for this Ligand-Target Pair
Affinity DataIC50: 6.15E+8nMAssay Description:Concentration required for its inhibitory activity against Proteus mirabilis C889 class A beta-lactamaseMore data for this Ligand-Target Pair
Affinity DataIC50: 6.46E+8nMAssay Description:Concentration required for its inhibitory activity against Proteus mirabilis C889 class A beta-lactamaseMore data for this Ligand-Target Pair