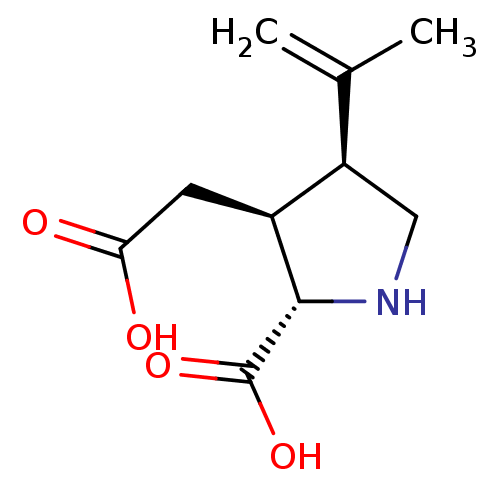
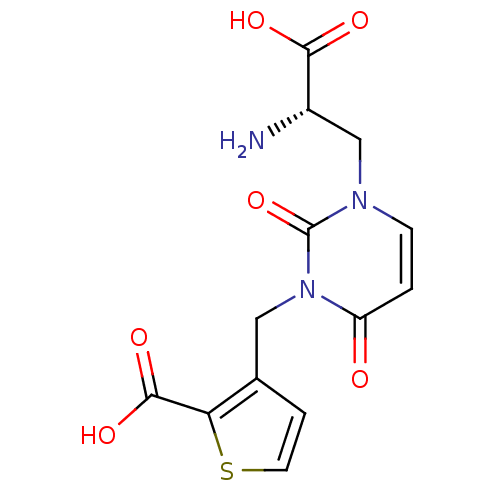
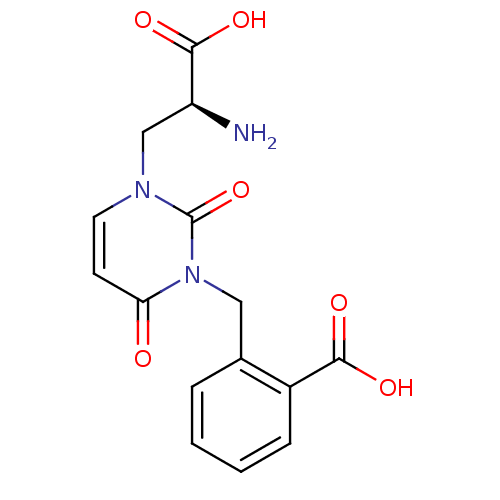
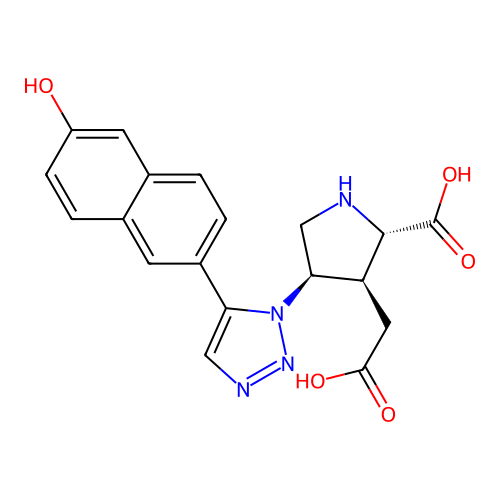
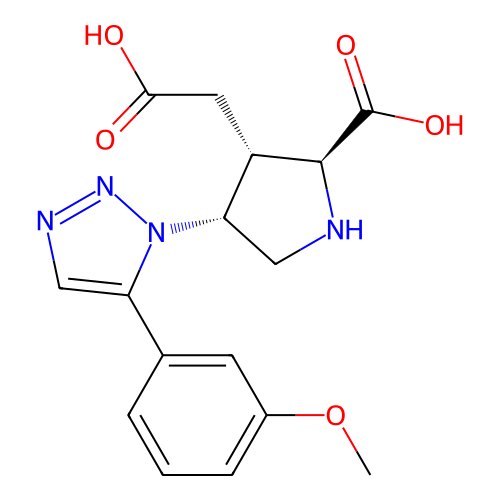
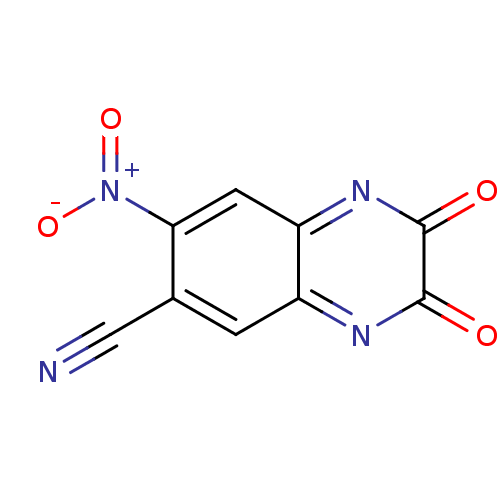
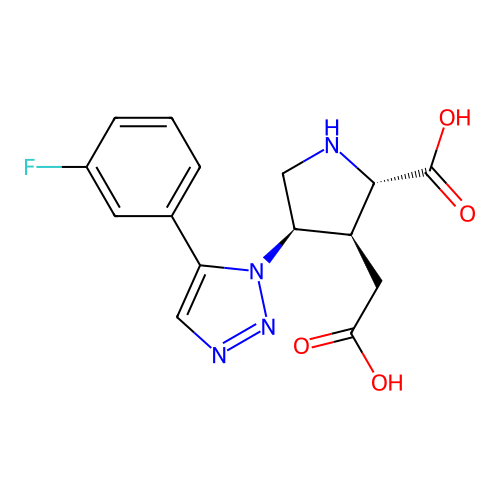
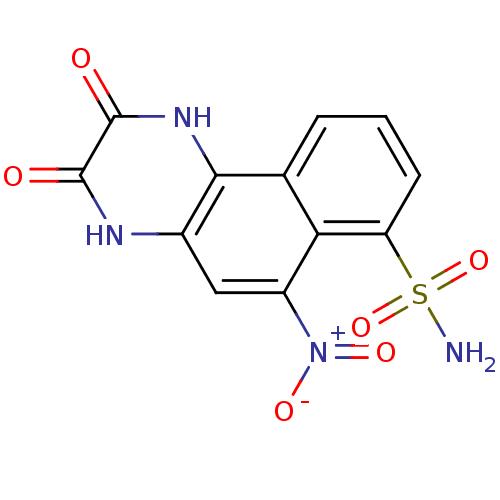
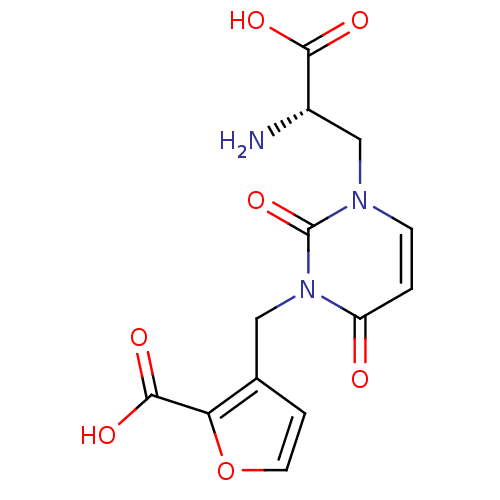
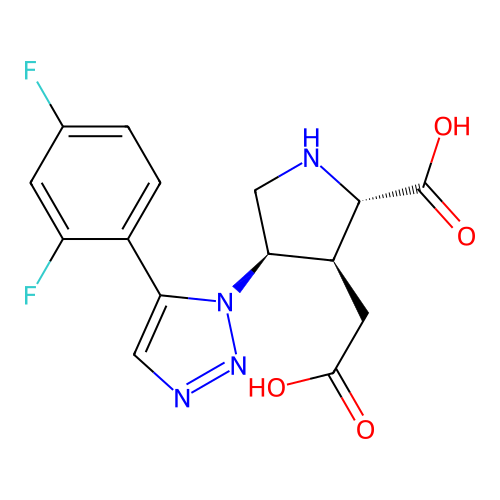
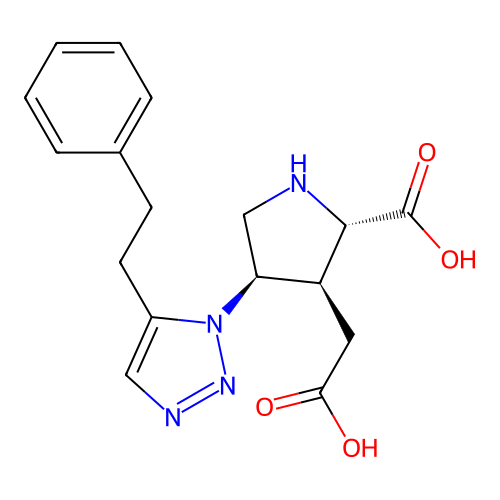
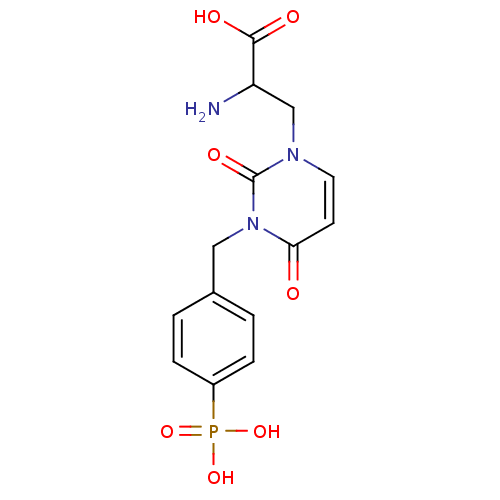
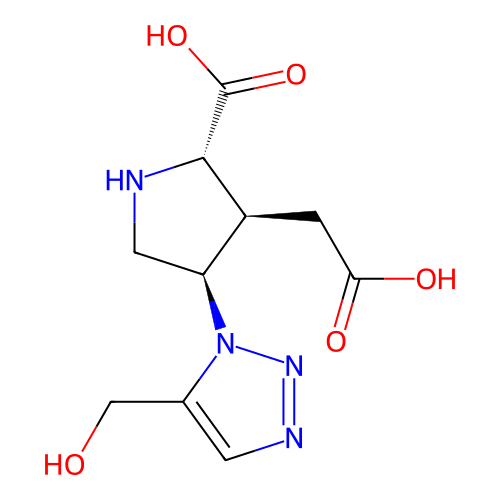
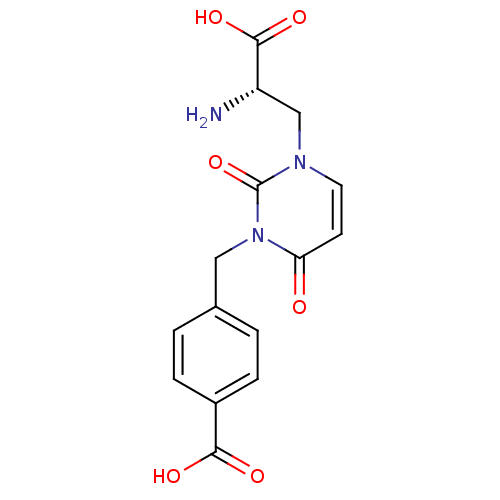
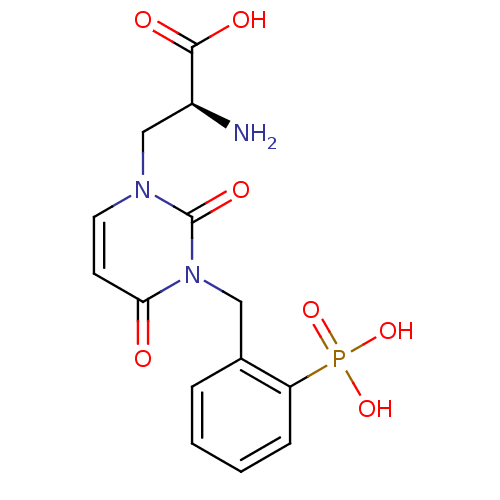
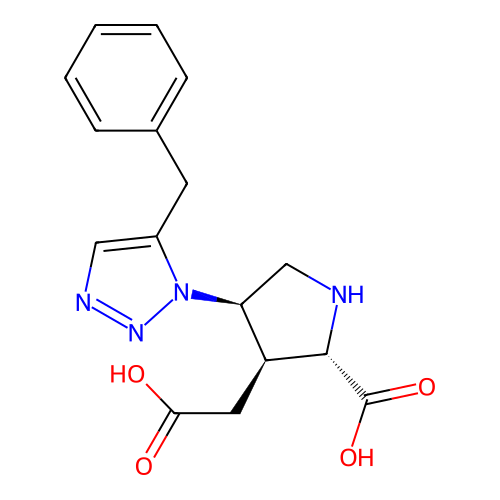
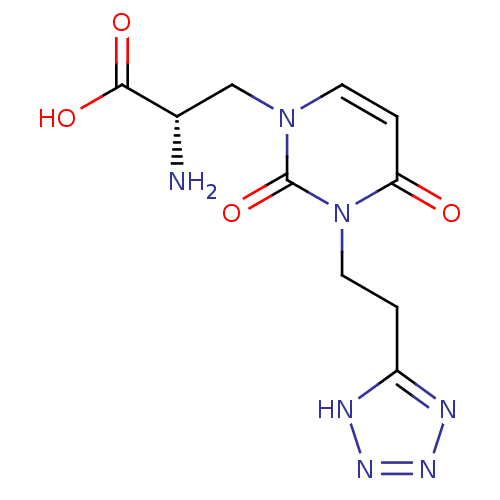
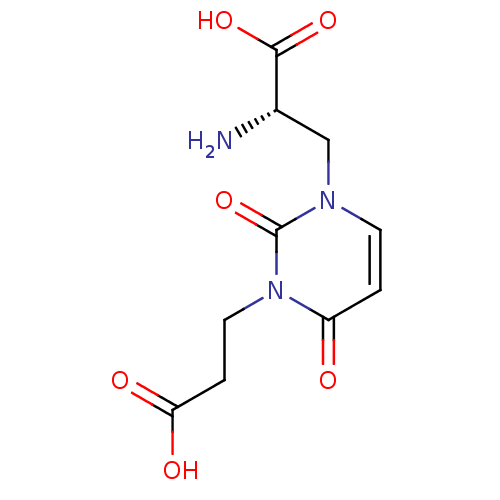
Affinity DataKi: 15nMAssay Description:Binding affinity to rat cloned KA2 receptorMore data for this Ligand-Target Pair
Affinity DataKd: 105nMAssay Description:Antagonist activity against GLUK5-containing kainate receptor by inhibition of kainate-induced depolarization of neonatal rat dorsal root fibersMore data for this Ligand-Target Pair
Affinity DataKi: 258nMAssay Description:Displacement of [3H]-kainate from mouse GluK5 expressed in Sf9 cell membrane incubated for 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKd: 402nMAssay Description:Antagonist activity against GLUK5-containing kainate receptor by inhibition of kainate-induced depolarization of neonatal rat dorsal root fibersMore data for this Ligand-Target Pair
Affinity DataKi: 420nMAssay Description:Displacement of [3H]-kainate from mouse GluK5 expressed in Sf9 cell membrane incubated for 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 432nMAssay Description:Displacement of [3H]-kainate from mouse GluK5 expressed in Sf9 cell membrane incubated for 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 483nMAssay Description:Displacement of [3H]-kainate from mouse GluK5 expressed in Sf9 cell membrane incubated for 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 499nMAssay Description:Displacement of [3H]-kainate from mouse GluK5 expressed in Sf9 cell membrane incubated for 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 1.02E+3nMAssay Description:Displacement of [3H]-kainate from mouse GluK5 expressed in Sf9 cell membrane incubated for 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKd: 1.66E+3nMAssay Description:Antagonist activity against GLUK5-containing kainate receptor by inhibition of kainate-induced depolarization of neonatal rat dorsal root fibersMore data for this Ligand-Target Pair
Affinity DataKi: 1.76E+3nMAssay Description:Displacement of [3H]-kainate from mouse GluK5 expressed in Sf9 cell membrane incubated for 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKd: 1.78E+3nMAssay Description:Antagonist activity against GLUK5-containing kainate receptor by inhibition of kainate-induced depolarization of neonatal rat dorsal root fibersMore data for this Ligand-Target Pair
Affinity DataKd: 1.88E+3nMAssay Description:Antagonist activity against GLUK5-containing kainate receptor by inhibition of kainate-induced depolarization of neonatal rat dorsal root fibersMore data for this Ligand-Target Pair
Affinity DataKi: 2.15E+3nMAssay Description:Displacement of [3H]-kainate from mouse GluK5 expressed in Sf9 cell membrane incubated for 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 3.25E+3nMAssay Description:Displacement of [3H]-kainate from mouse GluK5 expressed in Sf9 cell membrane incubated for 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKd: 4.68E+3nMAssay Description:Antagonist activity against GLUK5-containing kainate receptor by inhibition of kainate-induced depolarization of neonatal rat dorsal root fibersMore data for this Ligand-Target Pair
Affinity DataKi: 5.35E+3nMAssay Description:Displacement of [3H]-kainate from mouse GluK5 expressed in Sf9 cell membrane incubated for 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 5.57E+3nMAssay Description:Displacement of [3H]-kainate from mouse GluK5 expressed in Sf9 cell membrane incubated for 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKd: 5.80E+3nMAssay Description:Antagonist activity against GLUK5-containing kainate receptor by inhibition of kainate-induced depolarization of neonatal rat dorsal root fibersMore data for this Ligand-Target Pair
Affinity DataKi: 7.97E+3nMAssay Description:Displacement of [3H]-kainate from mouse GluK5 expressed in Sf9 cell membrane incubated for 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKd: 9.25E+3nMAssay Description:Antagonist activity against GLUK5-containing kainate receptor by inhibition of kainate-induced depolarization of neonatal rat dorsal root fibersMore data for this Ligand-Target Pair
Affinity DataKd: 1.05E+4nMAssay Description:Antagonist activity against GLUK5-containing kainate receptor by inhibition of kainate-induced depolarization of neonatal rat dorsal root fibersMore data for this Ligand-Target Pair
Affinity DataKi: 1.11E+4nMAssay Description:Displacement of [3H]-kainate from mouse GluK5 expressed in Sf9 cell membrane incubated for 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKd: 5.37E+4nMAssay Description:Antagonist activity against GLUK5-containing kainate receptor by inhibition of kainate-induced depolarization of neonatal rat dorsal root fibersMore data for this Ligand-Target Pair
Affinity DataKd: 6.02E+4nMAssay Description:Antagonist activity against GLUK5-containing kainate receptor by inhibition of kainate-induced depolarization of neonatal rat dorsal root fibersMore data for this Ligand-Target Pair
Affinity DataKd: 7.31E+4nMAssay Description:Antagonist activity against GLUK5-containing kainate receptor by inhibition of kainate-induced depolarization of neonatal rat dorsal root fibersMore data for this Ligand-Target Pair