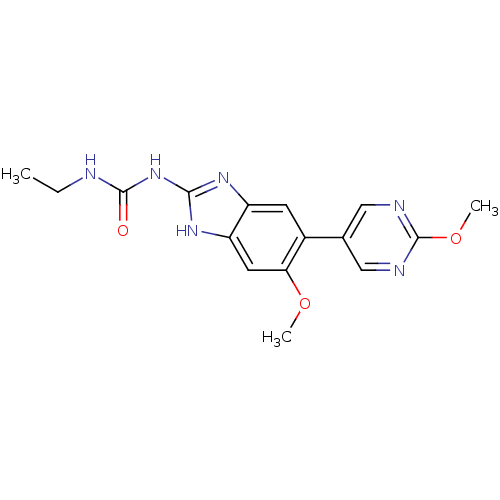
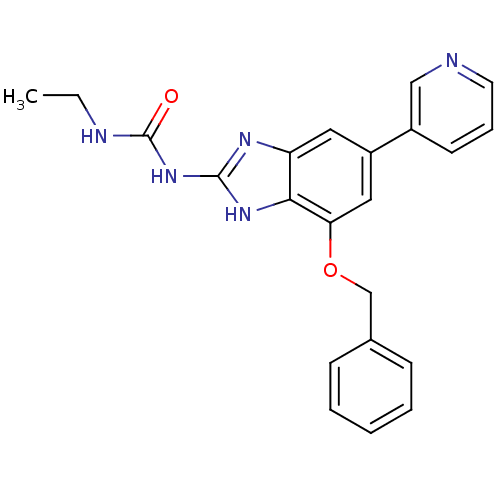
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Purdue University
Curated by ChEMBL
Purdue University
Curated by ChEMBL
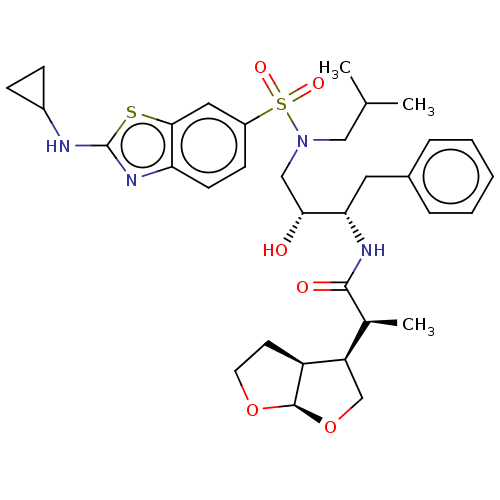
Affinity DataKi: 0.0220nMAssay Description:Inhibition of HIV1 protease by fluorescence based assayMore data for this Ligand-Target Pair
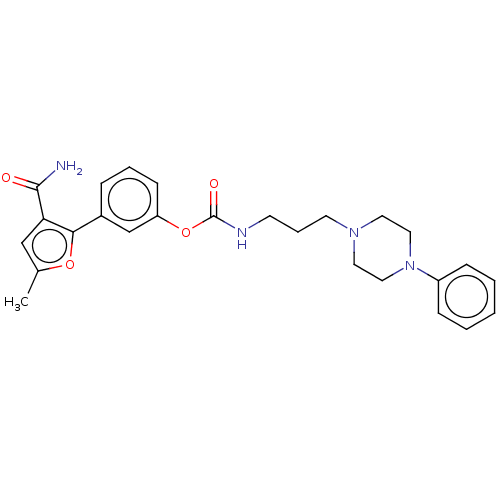
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Purdue University
Curated by ChEMBL
Purdue University
Curated by ChEMBL
Affinity DataKi: 0.0300nMAssay Description:Inhibition of HIV1 protease by fluorescence based assayMore data for this Ligand-Target Pair
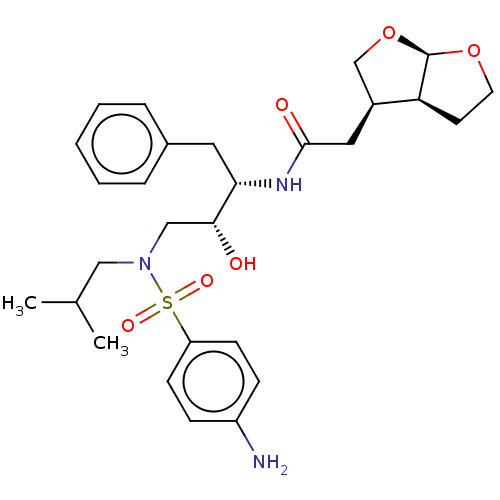
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Purdue University
Curated by ChEMBL
Purdue University
Curated by ChEMBL
Affinity DataKi: 0.0320nMAssay Description:Inhibition of HIV1 protease by fluorescence based assayMore data for this Ligand-Target Pair
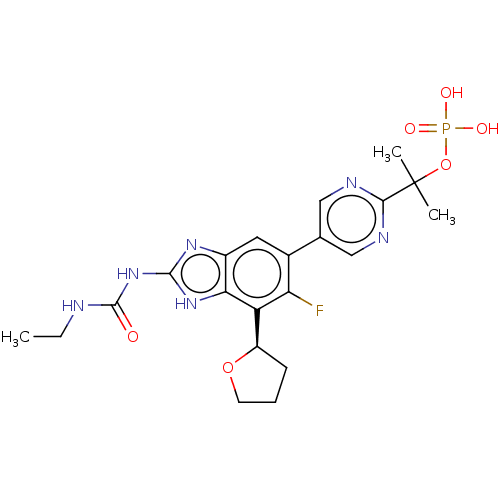
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Purdue University
Curated by ChEMBL
Purdue University
Curated by ChEMBL
Affinity DataKi: 0.0660nMAssay Description:Inhibition of HIV1 protease by fluorescence based assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.180nMAssay Description:Displacement of [3H]-7-OH-DPAT from rat brain membrane D3 receptor expressed in Sf9 cells incubated for 60 mins by liquid scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataKi: 0.230nMAssay Description:Inhibition of FAAH in mouse brain membranes assessed as inhibitory constant using [14C]-AEA as substrate incubated for 15 mins by scintillation count...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Purdue University
Curated by ChEMBL
Purdue University
Curated by ChEMBL
Affinity DataKi: 0.280nMAssay Description:Inhibition of HIV1 protease by fluorescence based assayMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Purdue University
Curated by ChEMBL
Purdue University
Curated by ChEMBL
Affinity DataKi: 1.5nMAssay Description:Inhibition of HIV1 protease by fluorescence based assayMore data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Inhibition of aureus Escherichia coli DNA gyrase A2B2 using pBR322 plasmid DNA as substrate by coupled enzyme reaction assayMore data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Inhibition of aureus Escherichia coli DNA gyrase A2B2 using pBR322 plasmid DNA as substrate by coupled enzyme reaction assayMore data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Inhibition of aureus Escherichia coli DNA gyrase A2B2 using pBR322 plasmid DNA as substrate by coupled enzyme reaction assayMore data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 6.90nMAssay Description:Inhibition of FAAH in mouse brain membranes assessed as inhibitory constant using [14C]-AEA as substrate incubated for 15 mins by scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 7nM ΔG°: -47.3kJ/molepH: 7.5 T: 2°CAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 8nM ΔG°: -47.0kJ/molepH: 7.5 T: 2°CAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase using pBR322 plasmid DNA as substrate by coupled enzyme reaction assayMore data for this Ligand-Target Pair
Affinity DataKi: 10nM ΔG°: -46.4kJ/molepH: 7.5 T: 2°CAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 10nM ΔG°: -46.4kJ/molepH: 7.5 T: 2°CAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 10nM ΔG°: -46.4kJ/molepH: 7.5 T: 2°CAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 12nM ΔG°: -46.0kJ/molepH: 7.5 T: 2°CAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 14nM ΔG°: -45.6kJ/molepH: 7.5 T: 2°CAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 15nM ΔG°: -45.4kJ/molepH: 7.5 T: 2°CAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Inhibition of FAAH in mouse brain membranes assessed as inhibitory constant using [14C]-AEA as substrate incubated for 15 mins by scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Inhibition of FAAH in mouse brain membranes assessed as inhibitory constant using [14C]-AEA as substrate incubated for 15 mins by scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 16nM ΔG°: -45.2kJ/molepH: 7.5 T: 2°CAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Purdue University
Curated by ChEMBL
Purdue University
Curated by ChEMBL
Affinity DataKi: 17nMAssay Description:Inhibition of HIV1 protease by fluorescence based assayMore data for this Ligand-Target Pair
Affinity DataKi: 17nM ΔG°: -45.1kJ/molepH: 7.5 T: 2°CAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: <17.7nMAssay Description:Inhibition of Staphylococcus aureus DNA gyraseMore data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:Displacement of [3H]-spiperone from human D3 receptor expressed in CHO cells co-expressing Galpha16 incubated for 120 mins by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:Displacement of [3H]-spiperone from human D3 receptor expressed in CHO cells co-expressing Galpha16 incubated for 120 mins by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: <21nMAssay Description:Inhibition of Staphylococcus aureus DNA gyraseMore data for this Ligand-Target Pair
Affinity DataKi: 23nM ΔG°: -44.3kJ/molepH: 7.5 T: 2°CAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair

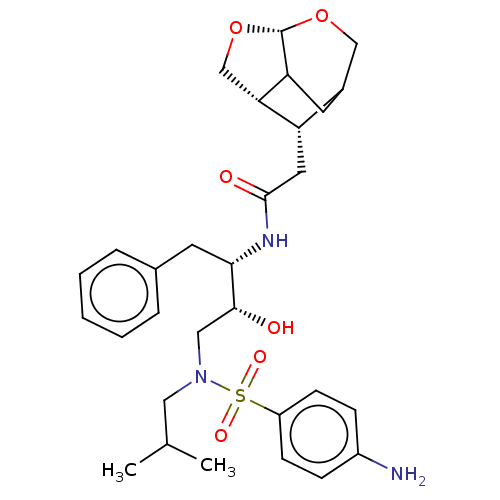
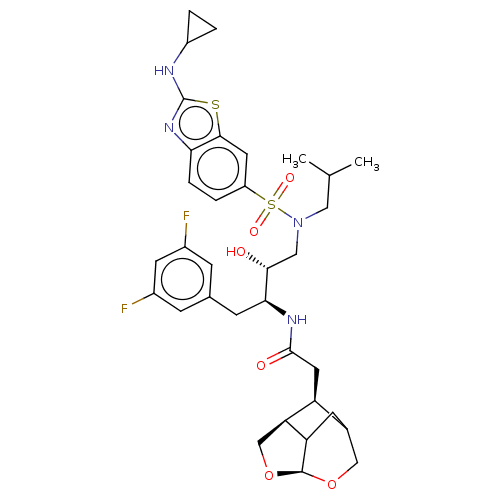
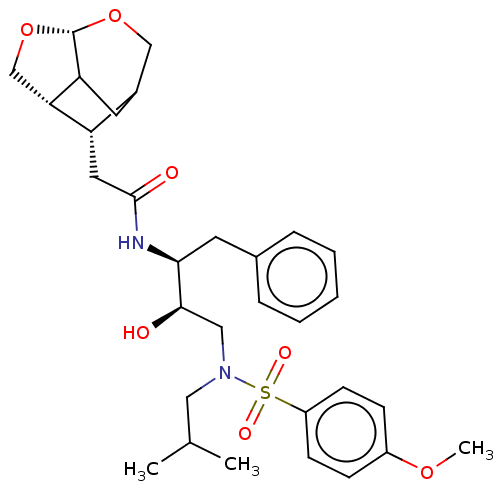
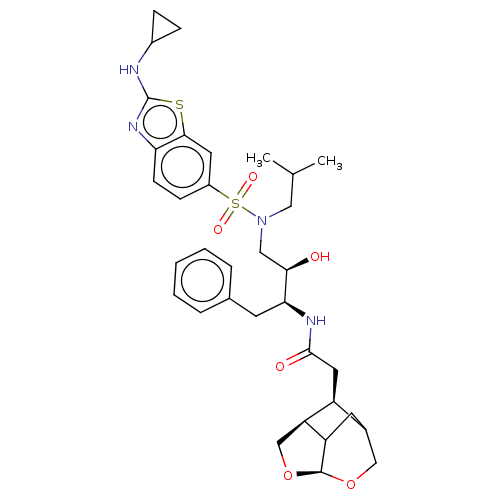
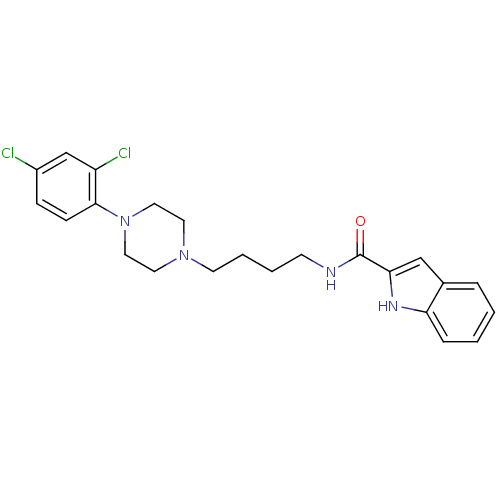
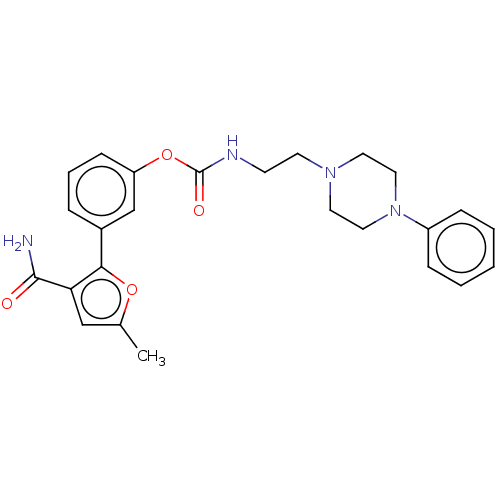
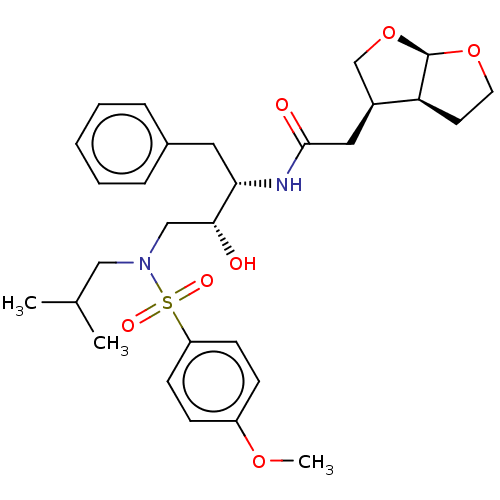
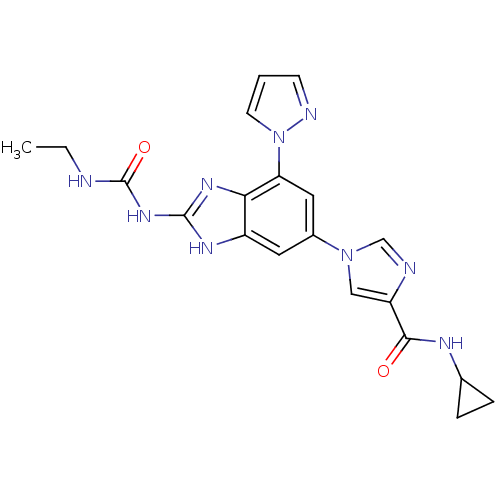
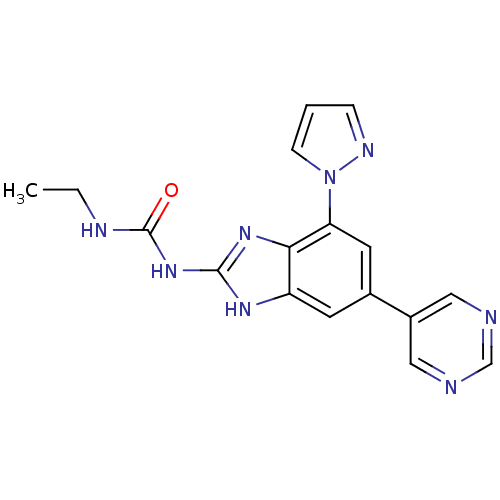
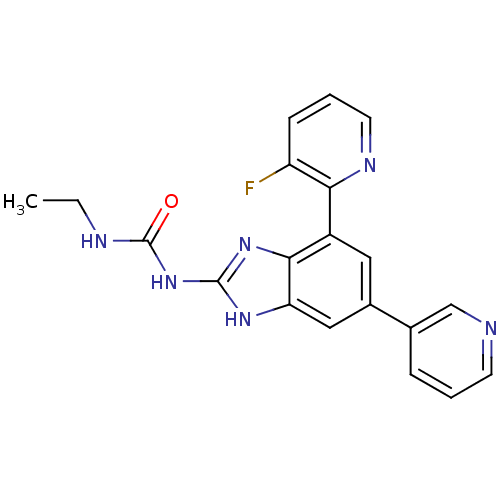

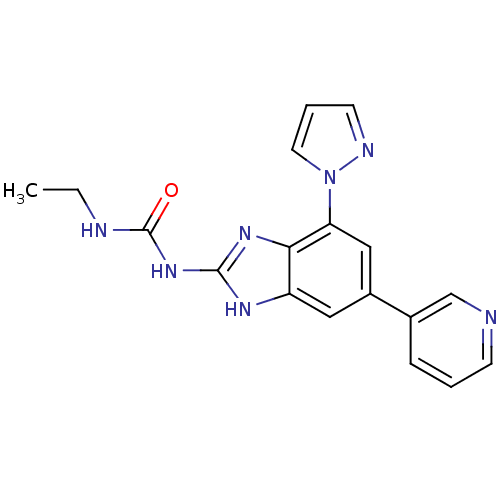
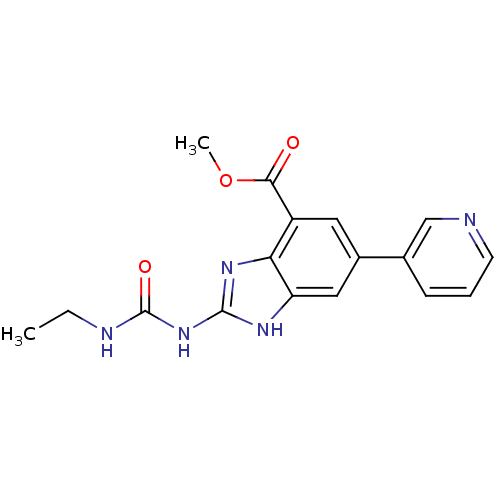
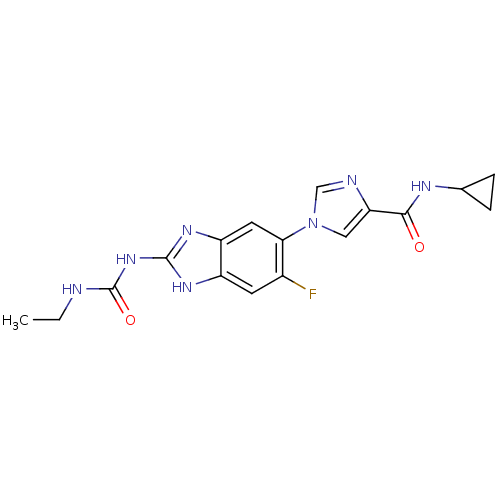
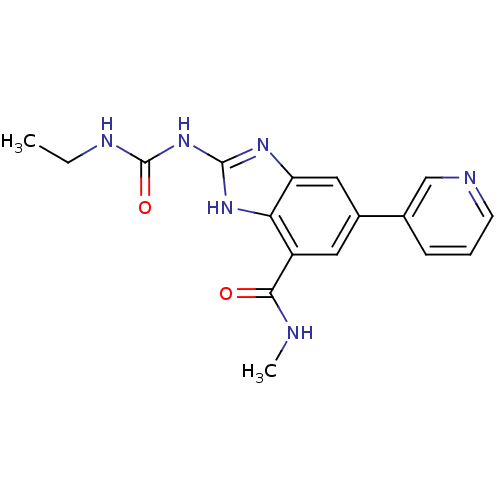
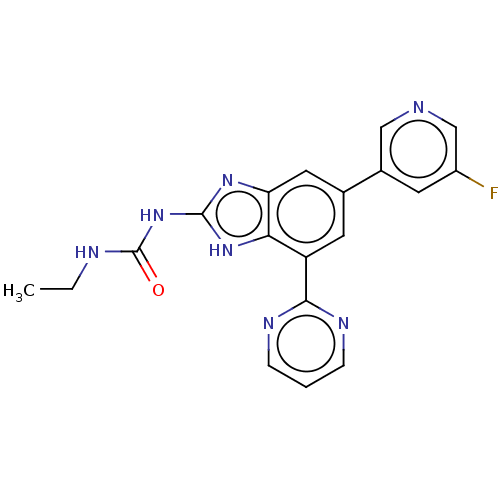
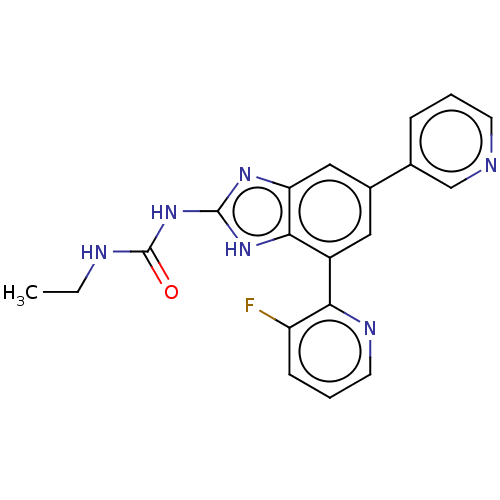
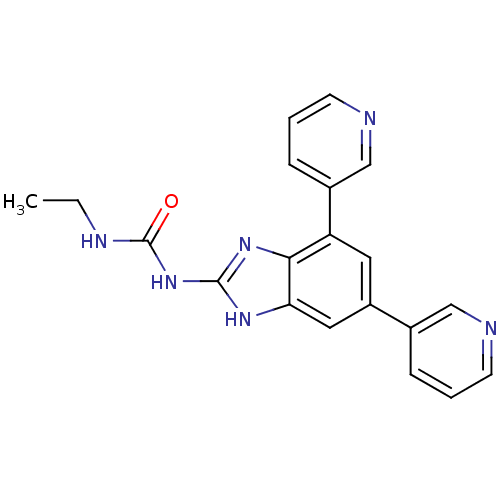
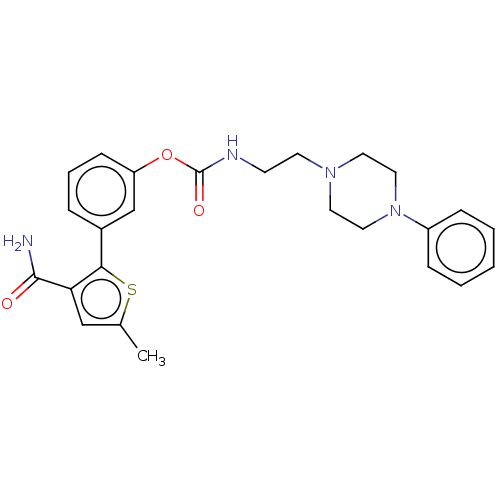
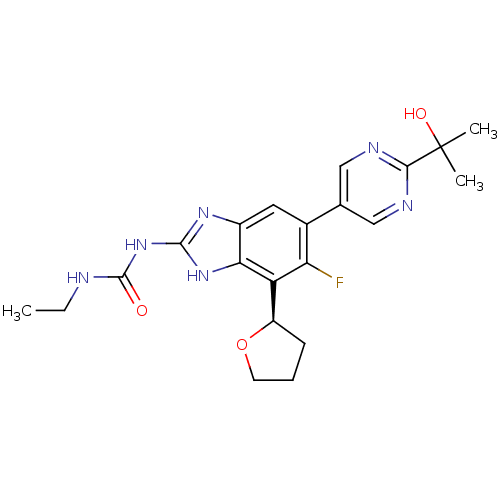
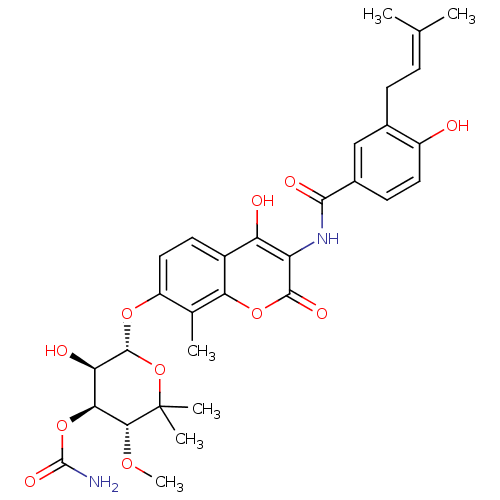
 3D Structure (crystal)
3D Structure (crystal)