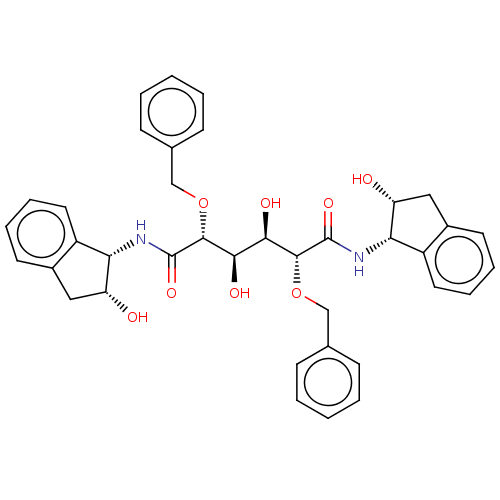
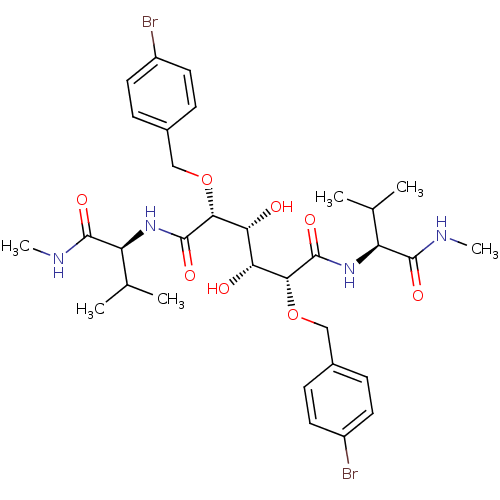
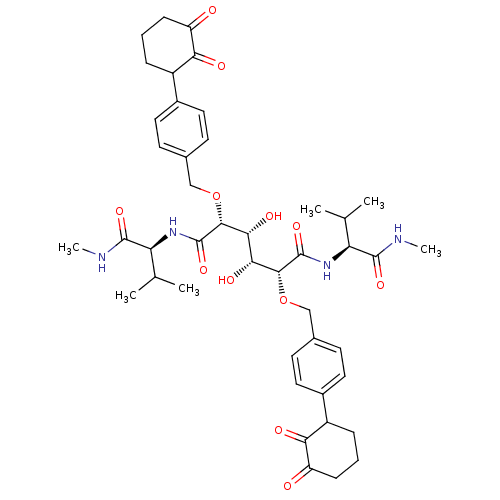
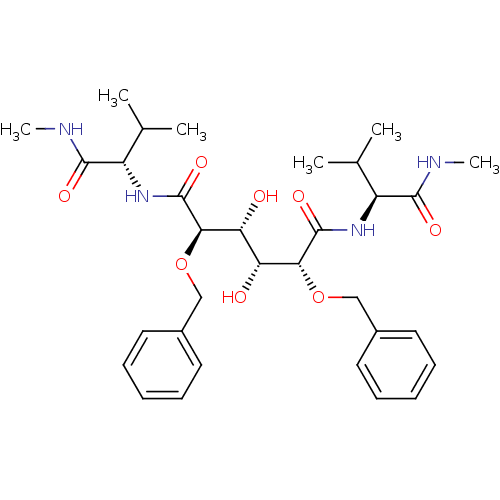
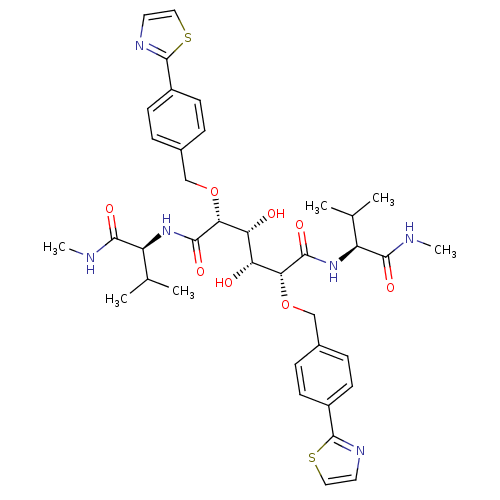
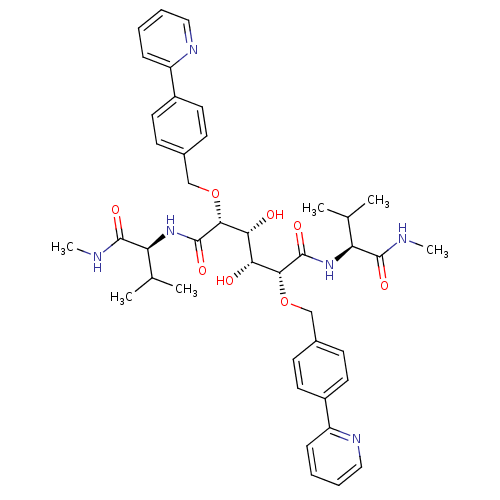
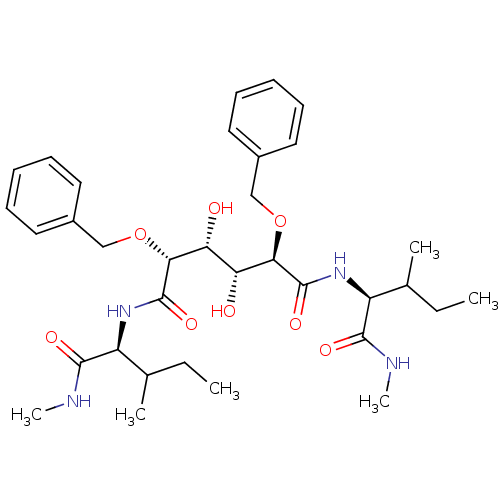
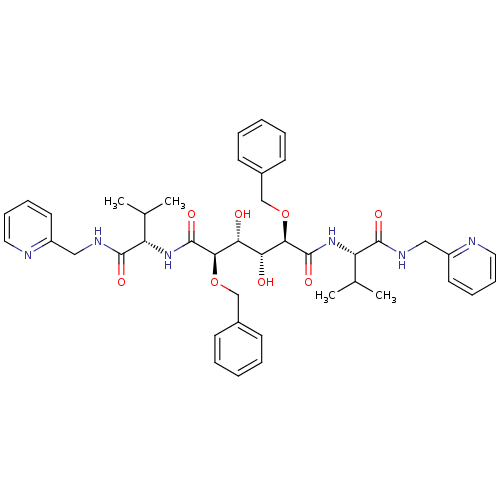
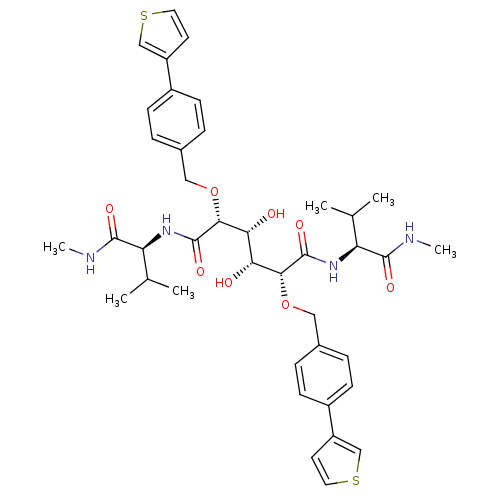
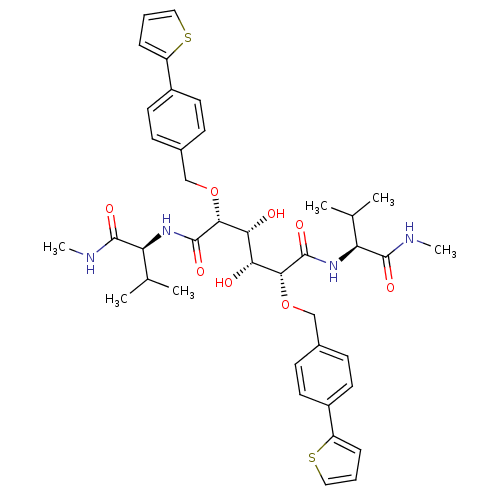
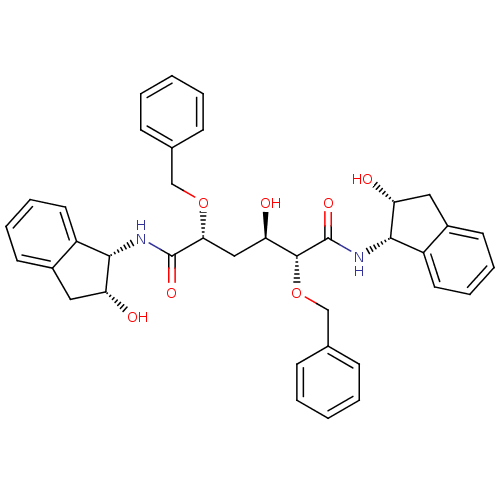
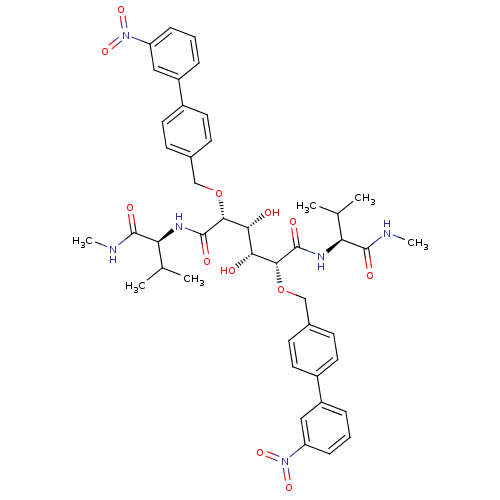
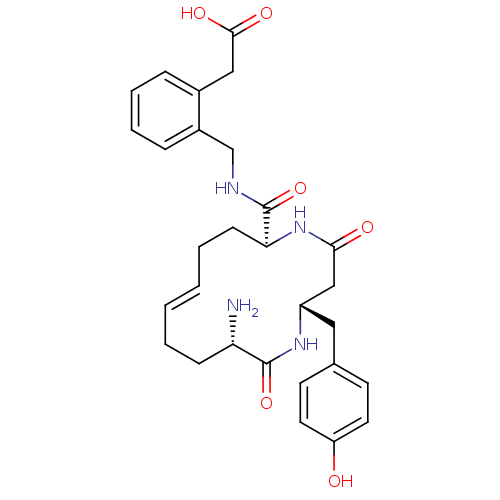
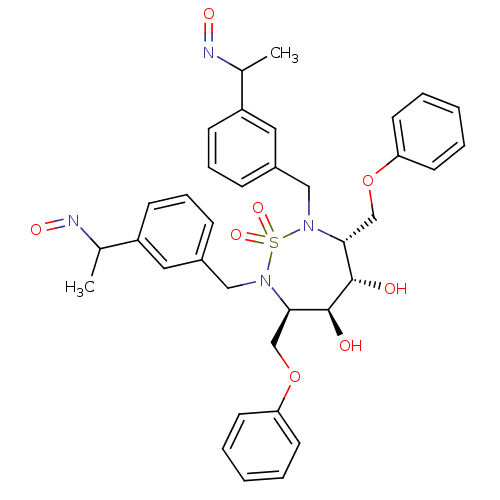
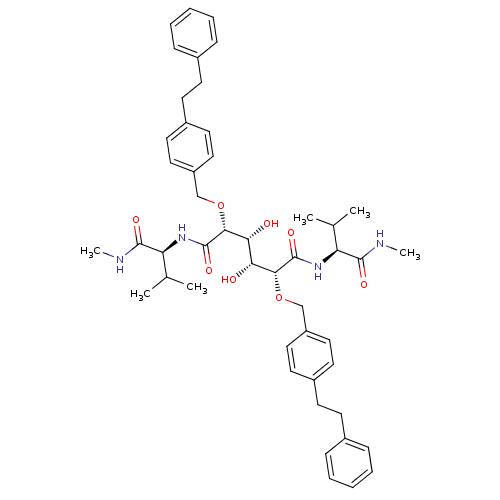
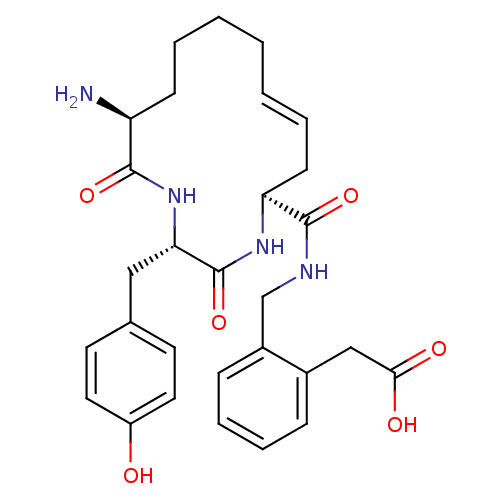
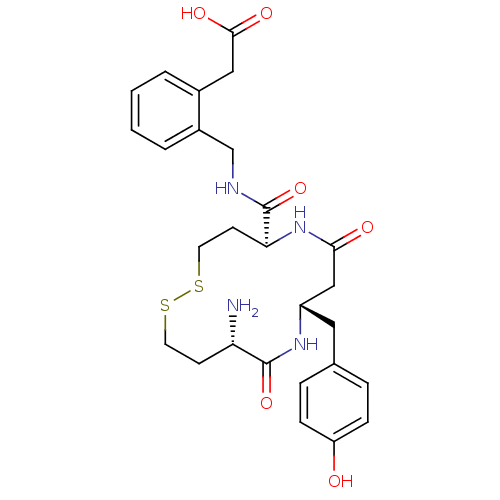
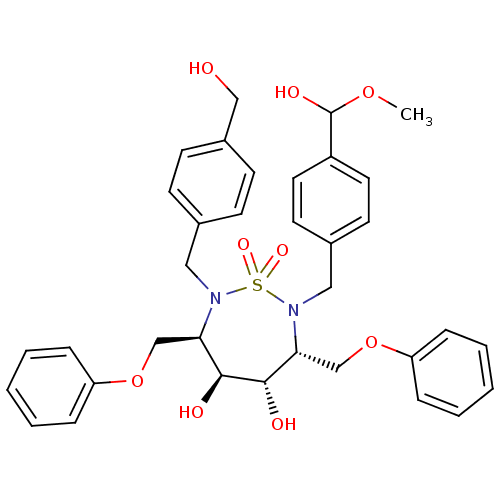
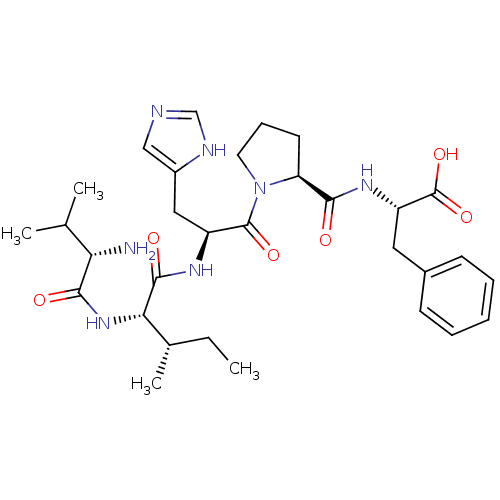
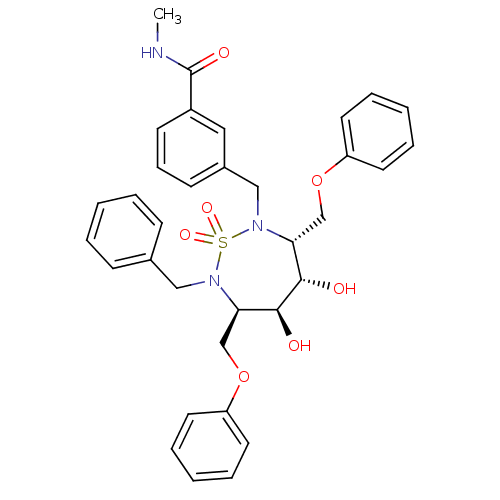
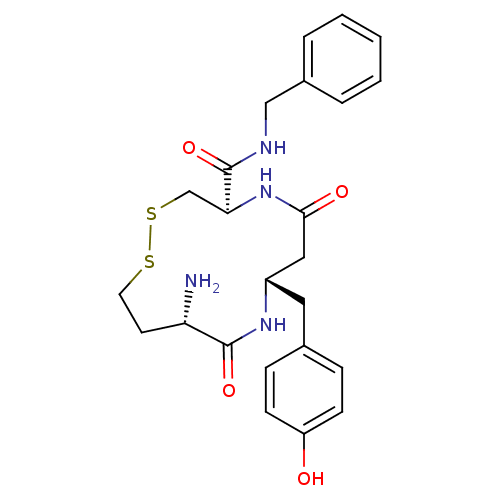
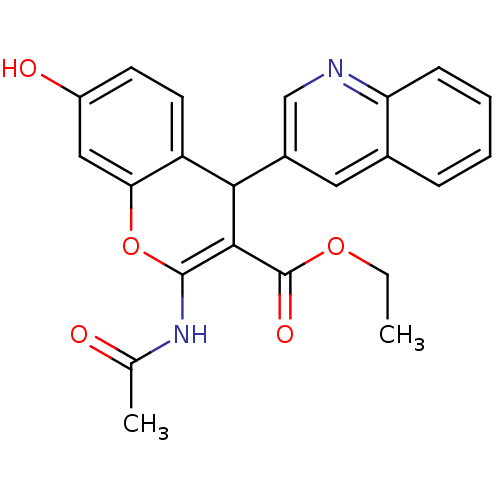
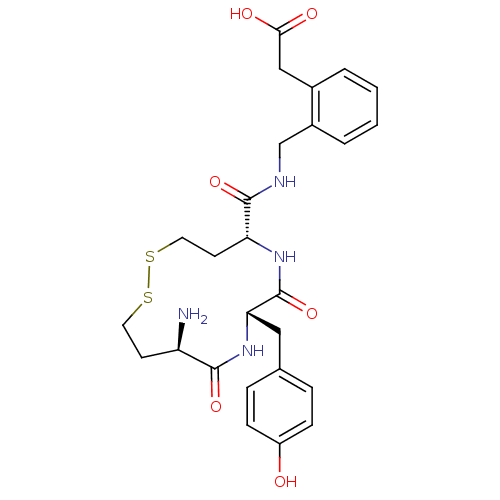
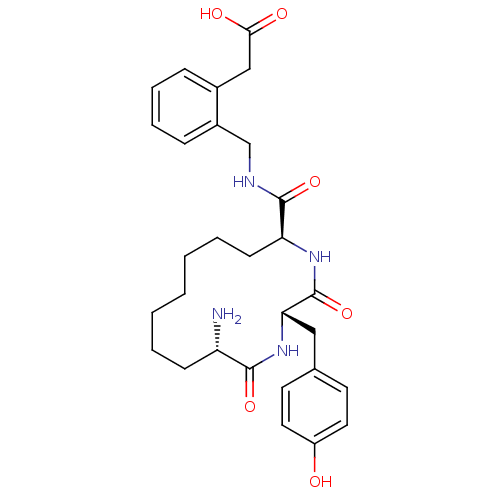
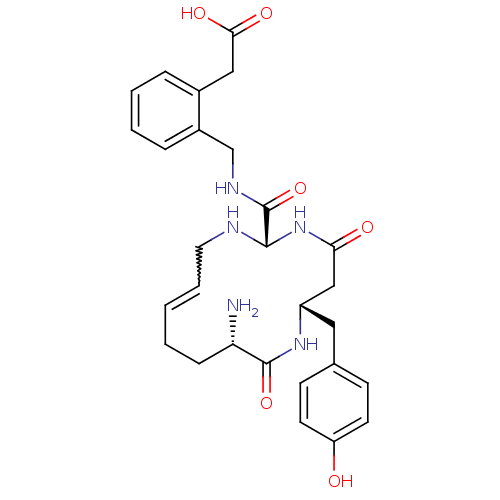
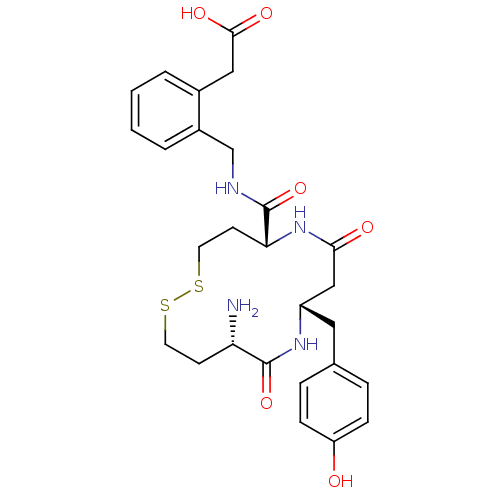
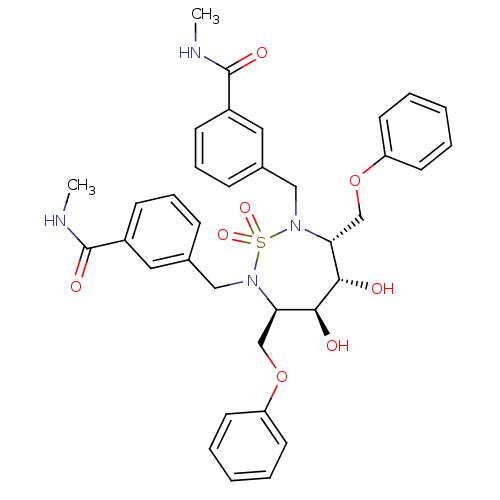
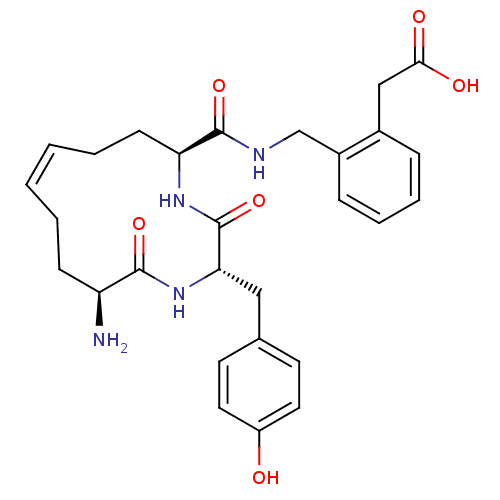
Affinity DataKi: 0.0900nM ΔG°: -58.3kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nM ΔG°: -58.0kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nM ΔG°: -58.0kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 0.200nM ΔG°: -56.3kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nM ΔG°: -55.3kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nM ΔG°: -55.3kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nM ΔG°: -55.3kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
Affinity DataKi: 0.400nM ΔG°: -54.5kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
Affinity DataKi: 0.400nM ΔG°: -54.5kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 0.600nM ΔG°: -53.5kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
Affinity DataKi: 0.600nM ΔG°: -53.5kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
Affinity DataKi: 0.700nM ΔG°: -53.1kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
Affinity DataKi: 0.900nM ΔG°: -52.5kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 0.920nM ΔG°: -52.4kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 1.20nM ΔG°: -51.8kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
Affinity DataKi: 1.20nM ΔG°: -51.8kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
Affinity DataKi: 1.20nM ΔG°: -51.8kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 1.40nM ΔG°: -51.4kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 1.40nM ΔG°: -51.4kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:Inhibition of catalytic activity of recombinant human IRAP transfected in human HEK-293 cells assessed as clevage of substrate L-leucine-p-nitroanili...More data for this Ligand-Target Pair
Affinity DataKi: 3.10nM ΔG°: -49.4kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 3.10nMAssay Description:Inhibitory activity against HIV-1 Protease expressed in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKi: 3.30nMAssay Description:Inhibition of catalytic activity of human recombinant IRAP expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 3.40nM ΔG°: -49.1kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 3.40nMAssay Description:Inhibitory activity against HIV-1 Protease expressed in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKi: 3.80nM ΔG°: -48.9kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
Affinity DataKi: 4.10nMAssay Description:Inhibition of catalytic activity of recombinant human IRAP transfected in human HEK-293 cells assessed as clevage of substrate L-leucine-p-nitroanili...More data for this Ligand-Target Pair
Affinity DataKi: 4.40nM ΔG°: -48.5kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 5.20nMAssay Description:Inhibition of catalytic activity of human recombinant IRAP expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Displacement of [125I]Angiotensin 4 from human recombinant IRAP expressed in CHOK1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 7.30nM ΔG°: -47.2kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 9.30nMAssay Description:Displacement of [3H]AL-11 from human IRAP expressed in CHO-K1 cells after 30 mins by liquid scintillation counting in presence of 30 mM EDTA/600 uM 1...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Displacement of [3H]AL-11 from human IRAP expressed in CHO-K1 cells after 30 mins by liquid scintillation counting in presence of 30 mM EDTA/600 uM 1...More data for this Ligand-Target Pair
Affinity DataKi: 11nM ΔG°: -46.2kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 19nMAssay Description:Inhibitory activity against HIV-1 Protease expressed in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:Inhibition of catalytic activity of human recombinant IRAP expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Inhibition of catalytic activity of recombinant human IRAP transfected in human HEK-293 cells assessed as clevage of substrate L-leucine-p-nitroanili...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Inhibition of catalytic activity of human recombinant IRAP expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Displacement of [3H]AL-11 from human IRAP expressed in CHO-K1 cells after 30 mins by liquid scintillation counting in presence of 30 mM EDTA/600 uM 1...More data for this Ligand-Target Pair
Affinity DataKi: 23nM ΔG°: -44.3kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Inhibition of catalytic activity of human recombinant IRAP expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:Displacement of [3H]AL-11 from human IRAP expressed in CHO-K1 cells after 30 mins by liquid scintillation counting in presence of 30 mM EDTA/600 uM 1...More data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:Inhibition of catalytic activity of recombinant human IRAP transfected in human HEK-293 cells assessed as clevage of substrate L-leucine-p-nitroanili...More data for this Ligand-Target Pair
Affinity DataKi: 30.4nMAssay Description:Inhibition of catalytic activity of recombinant human IRAP transfected in human HEK-293 cells assessed as clevage of substrate L-leucine-p-nitroanili...More data for this Ligand-Target Pair
Affinity DataKi: 35nMAssay Description:Displacement of [3H]AL-11 from human IRAP expressed in CHO-K1 cells after 30 mins by liquid scintillation counting in presence of 30 mM EDTA/600 uM 1...More data for this Ligand-Target Pair
Affinity DataKi: 35nMAssay Description:Displacement of [3H]AL-11 from human IRAP expressed in CHO-K1 cells after 30 mins by liquid scintillation counting in presence of 30 mM EDTA/600 uM 1...More data for this Ligand-Target Pair
Affinity DataKi: 39nM ΔG°: -43.0kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 41.1nMAssay Description:Inhibition of catalytic activity of recombinant human IRAP transfected in human HEK-293 cells assessed as clevage of substrate L-leucine-p-nitroanili...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 43nMAssay Description:Inhibitory activity against HIV-1 Protease expressed in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKi: 43nM ΔG°: -42.8kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair

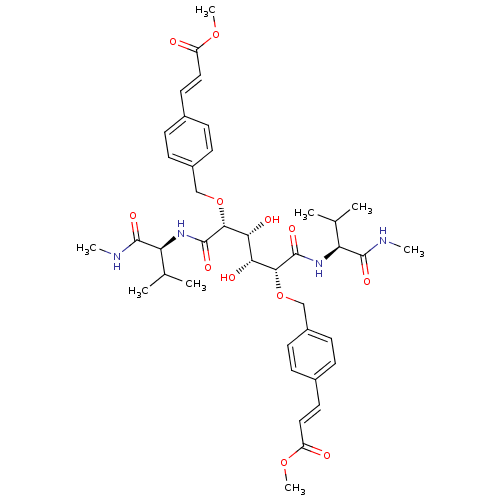
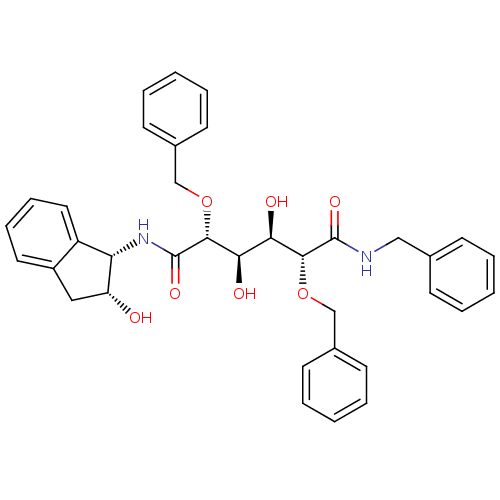
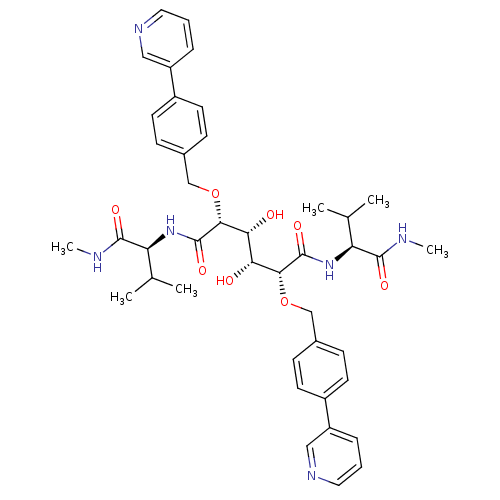
 3D Structure (crystal)
3D Structure (crystal)