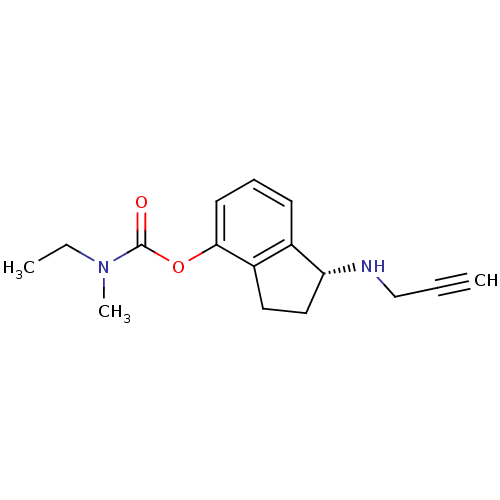
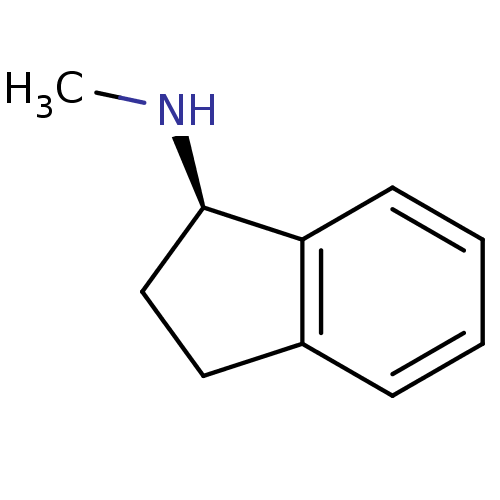
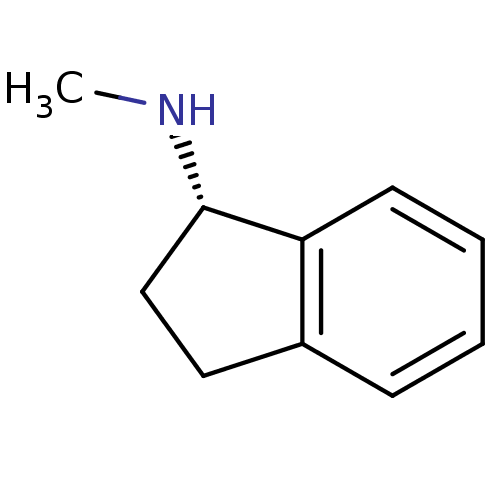
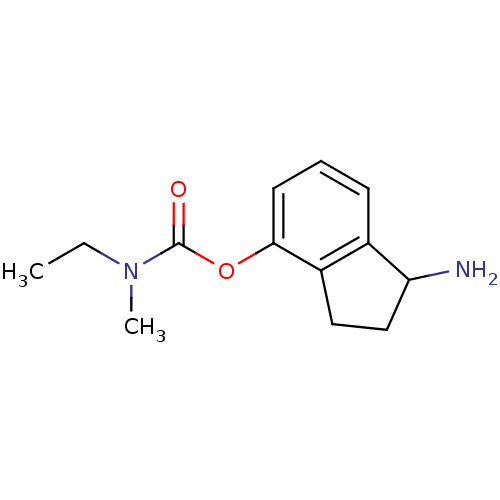
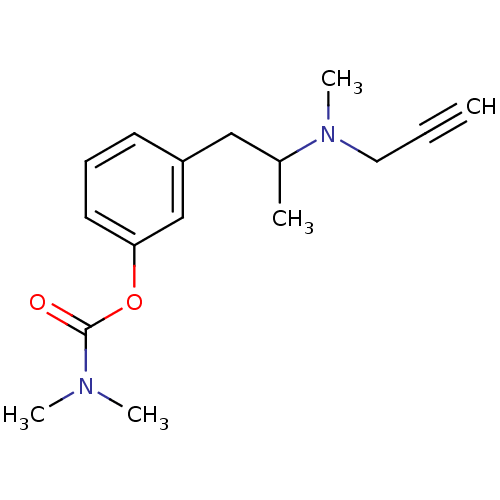
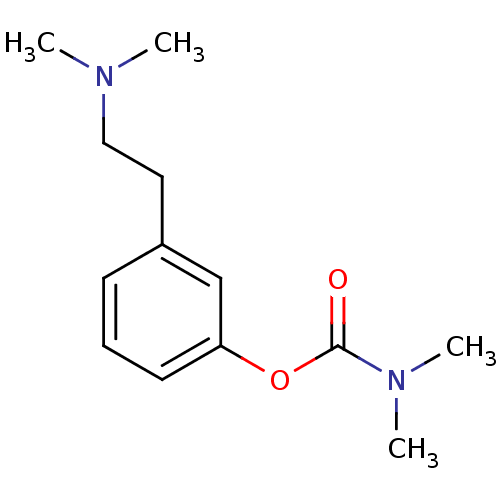
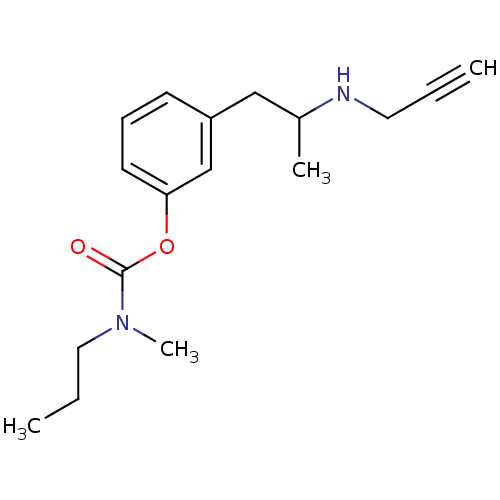
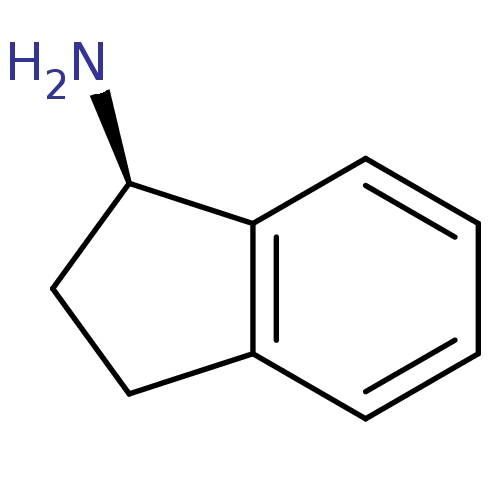Affinity DataKi: 600nM ΔG°: -35.5kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
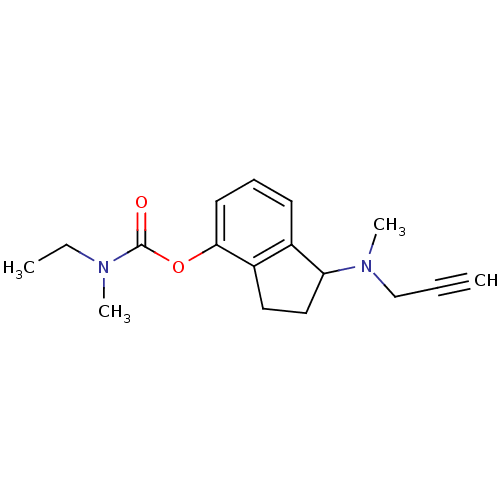
Affinity DataKi: 700nM ΔG°: -35.1kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
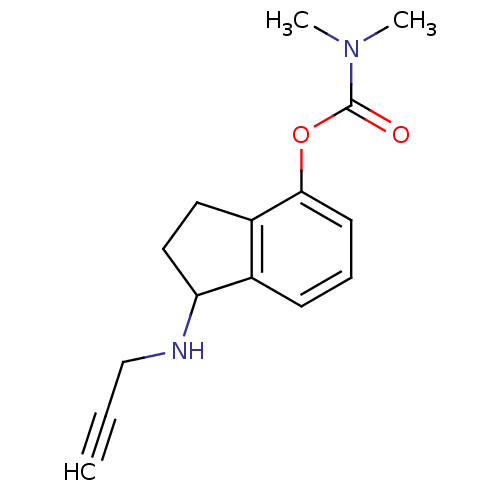
Affinity DataKi: 700nM ΔG°: -35.1kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
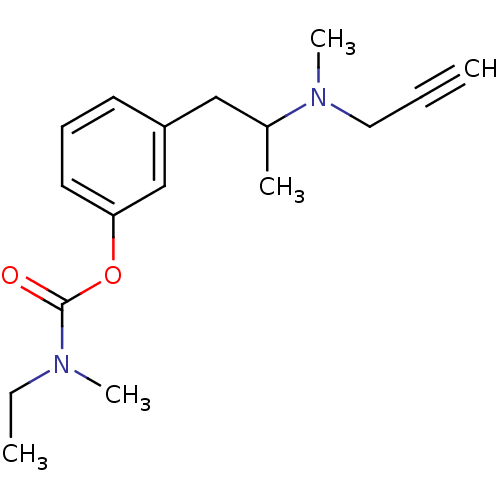
Affinity DataKi: 1.90E+3nM ΔG°: -32.7kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+3nM ΔG°: -32.5kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 4.90E+3nM ΔG°: -30.3kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 6.00E+3nM ΔG°: -29.8kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 7.00E+3nM ΔG°: -29.4kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 9.70E+3nM ΔG°: -28.6kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 9.70E+3nM ΔG°: -28.6kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 1.70E+4nM ΔG°: -27.2kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 1.70E+4nM ΔG°: -27.2kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 3.10E+4nM ΔG°: -25.7kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 3.20E+4nM ΔG°: -25.7kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 5.00E+4nM ΔG°: -24.5kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 5.60E+4nM ΔG°: -24.3kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 6.70E+4nM ΔG°: -23.8kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 1.12E+5nM ΔG°: -22.5kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 1.27E+5nM ΔG°: -22.2kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 1.30E+5nM ΔG°: -22.2kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 1.37E+5nM ΔG°: -22.1kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 1.54E+5nM ΔG°: -21.8kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 1.60E+5nM ΔG°: -21.7kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 2.13E+5nM ΔG°: -21.0kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 2.27E+5nM ΔG°: -20.8kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 2.70E+5nM ΔG°: -20.4kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 2.95E+5nM ΔG°: -20.1kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 1.08E+6nM ΔG°: -16.9kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 27nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 30nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 53nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 54nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 70nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 70nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair

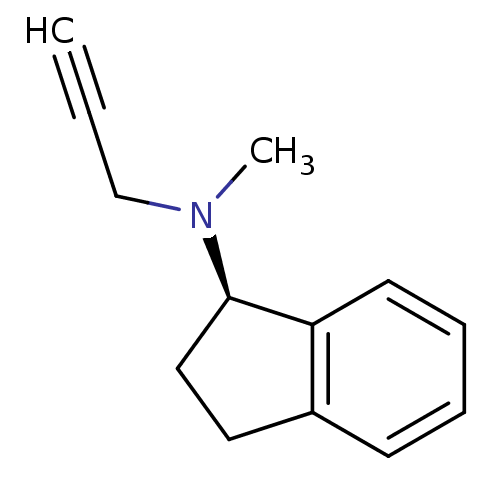
 3D Structure (crystal)
3D Structure (crystal)