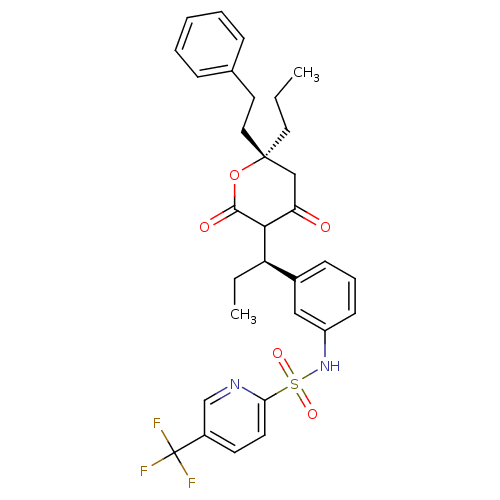
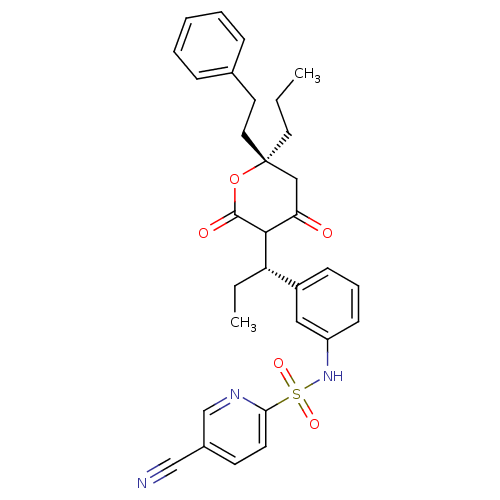
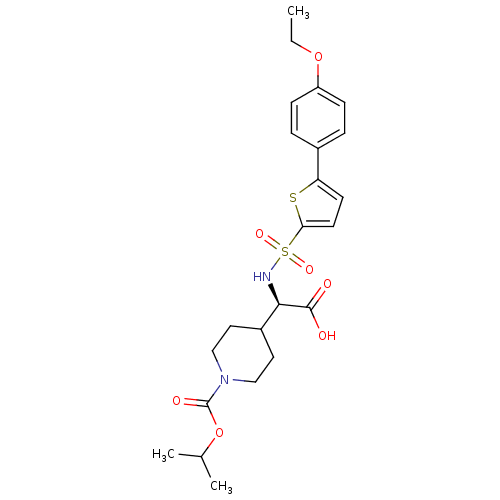
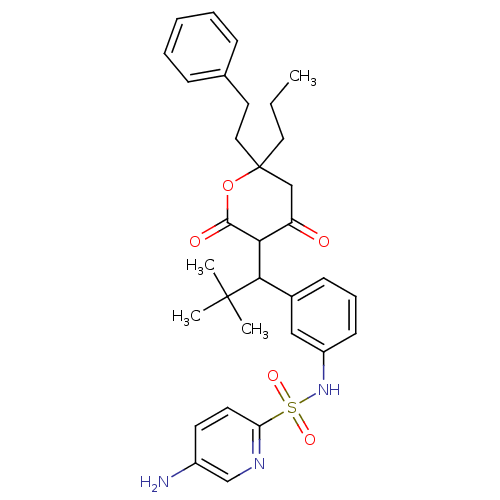
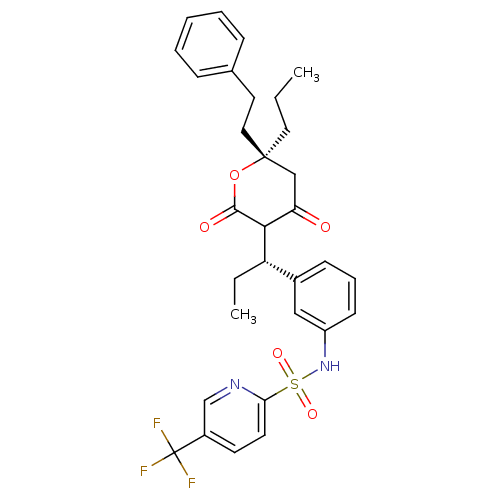
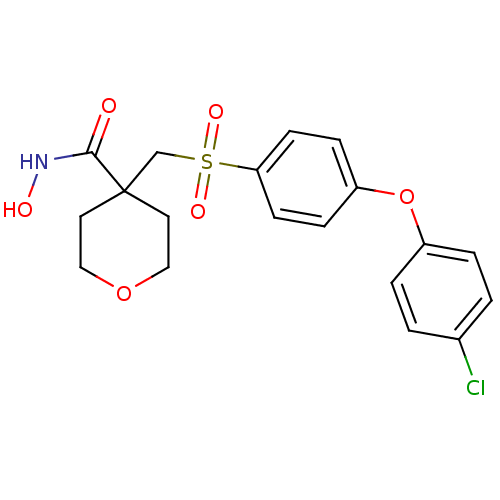
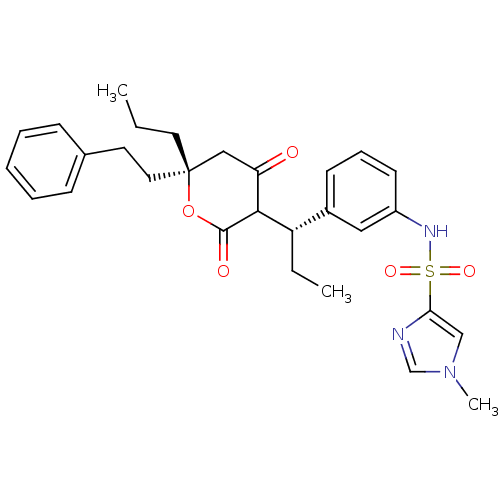
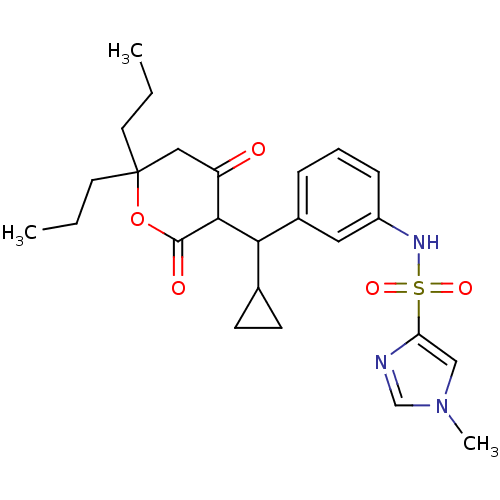
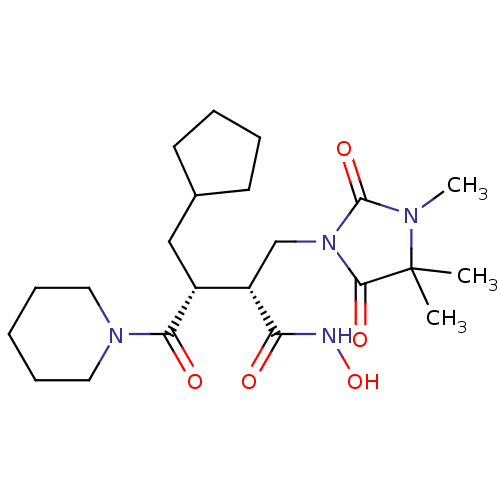
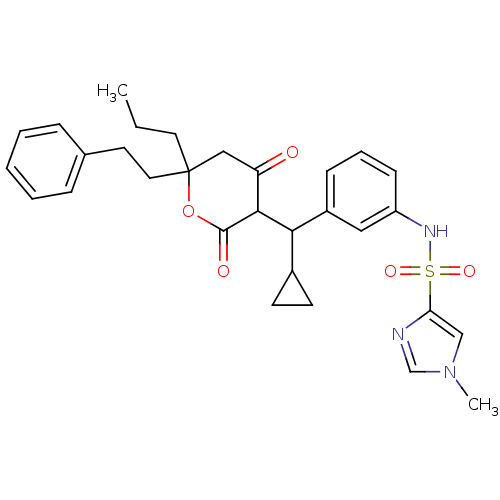
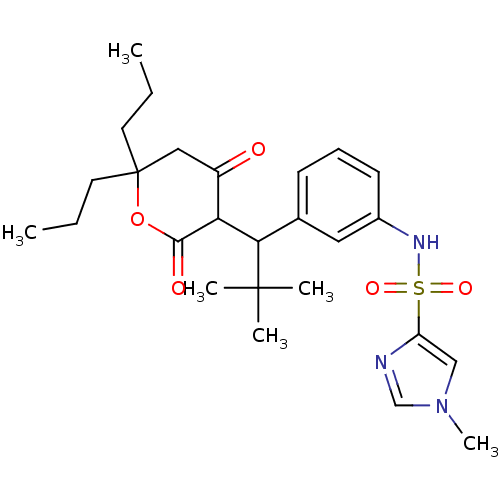
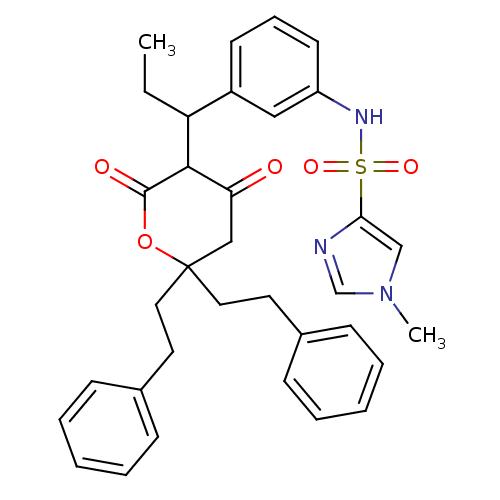
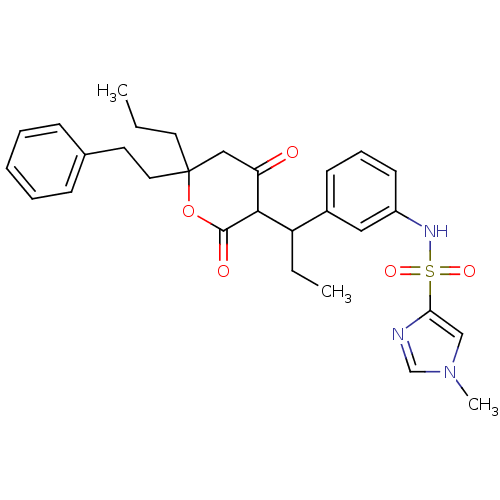
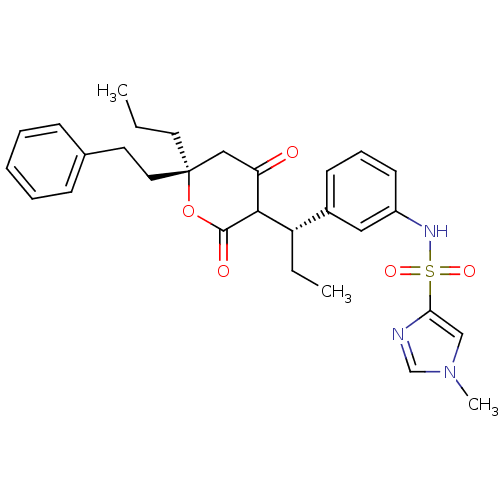
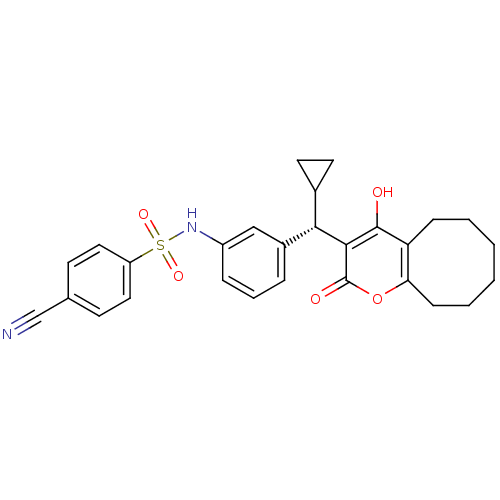
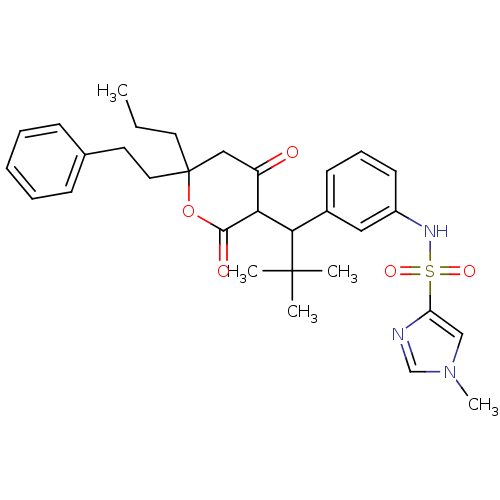
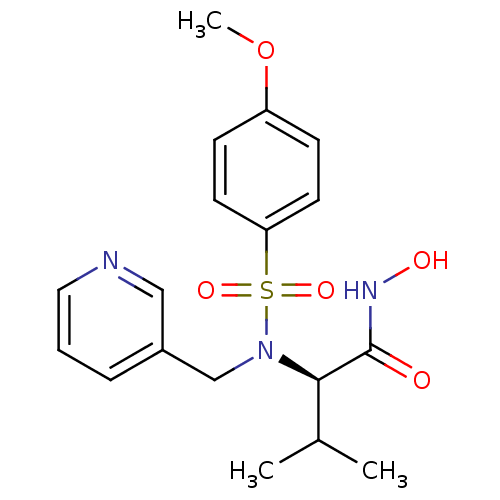
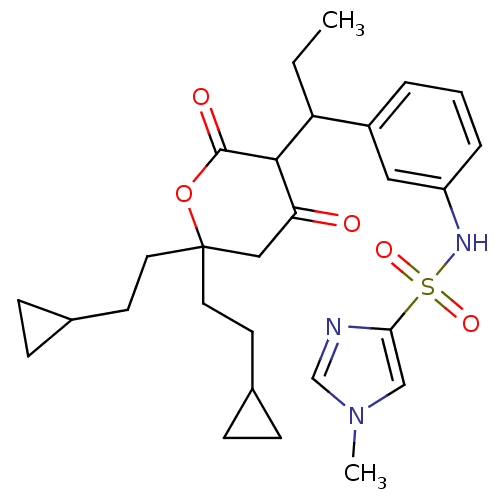
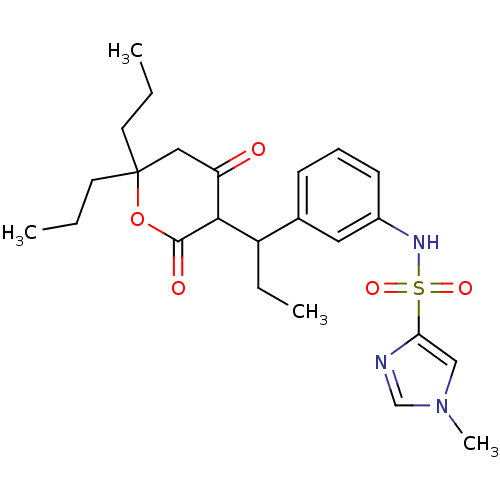
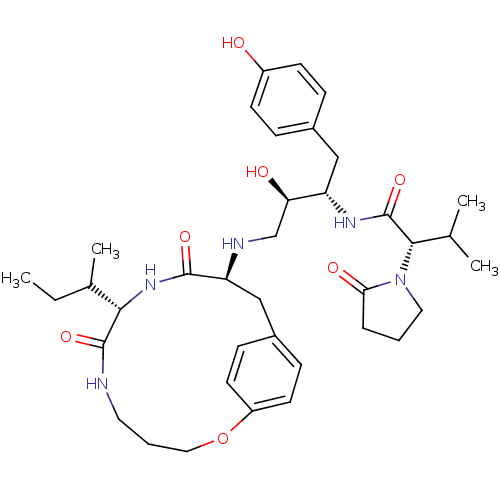
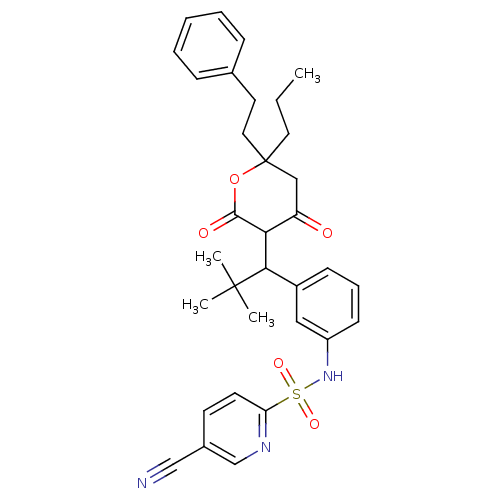
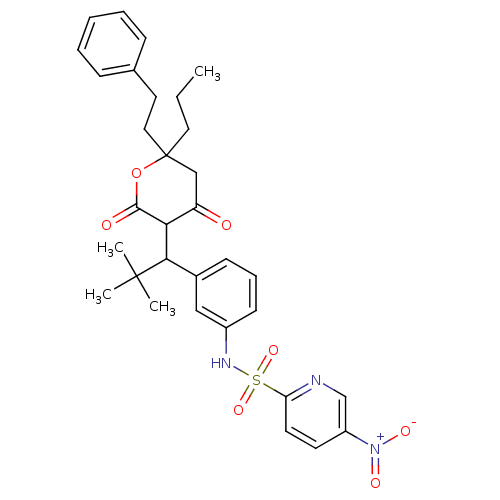
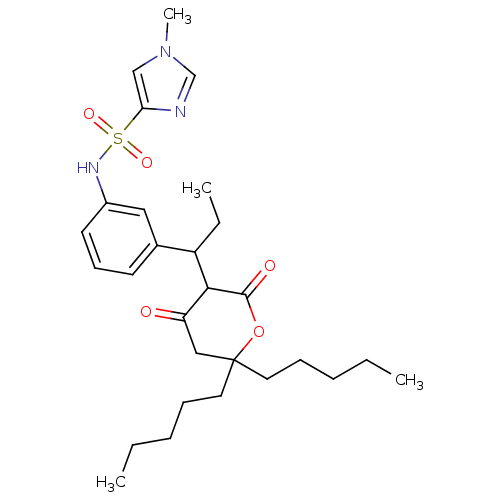
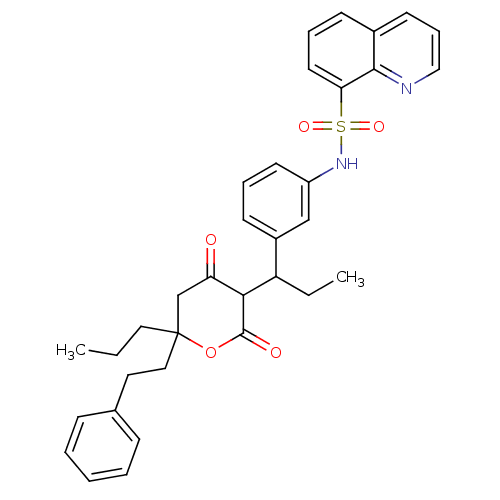
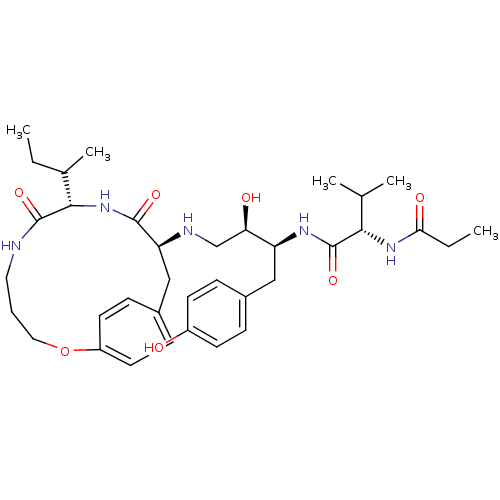
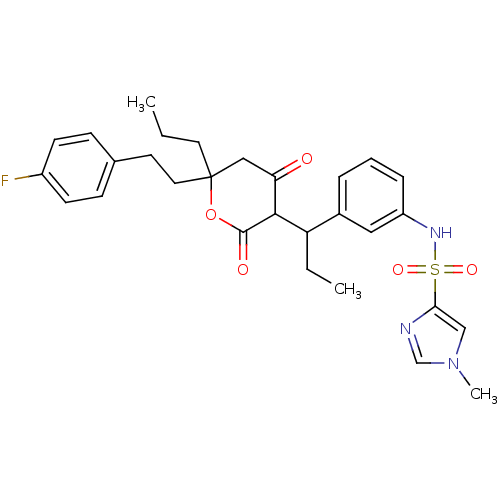
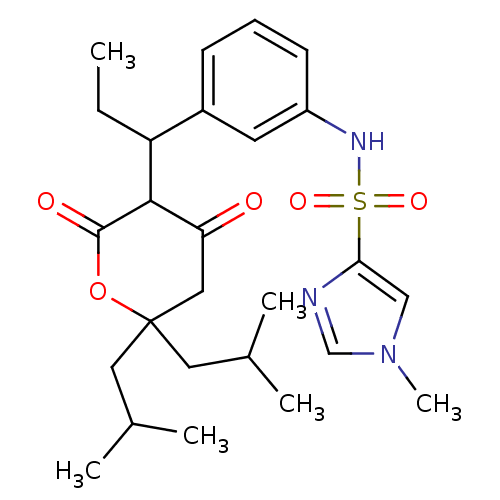
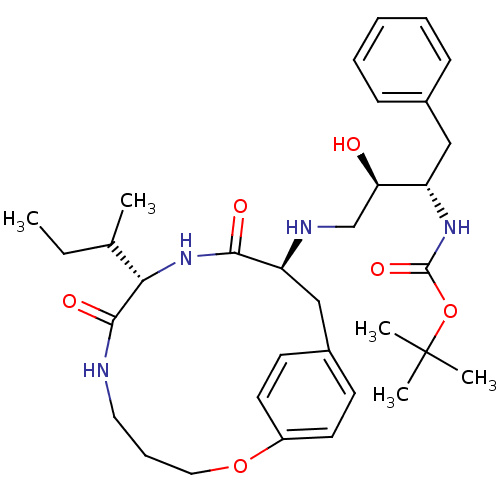
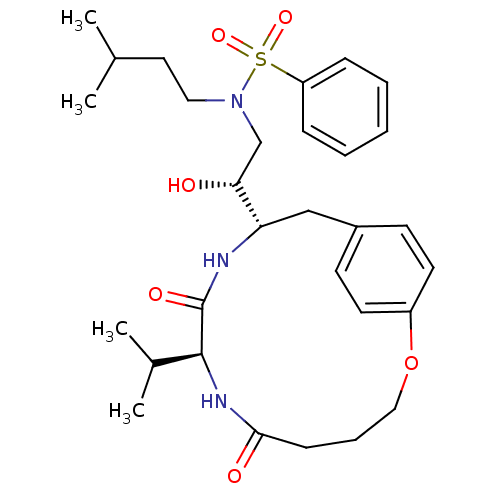
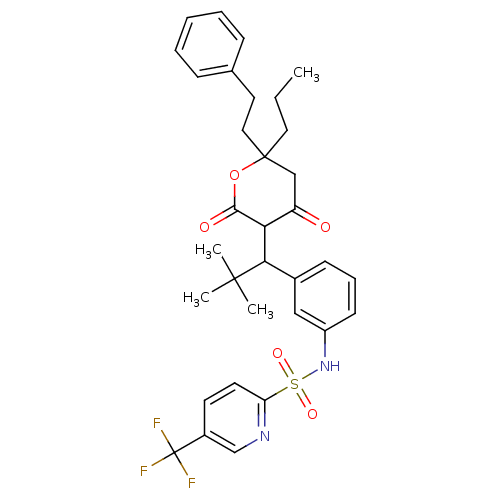
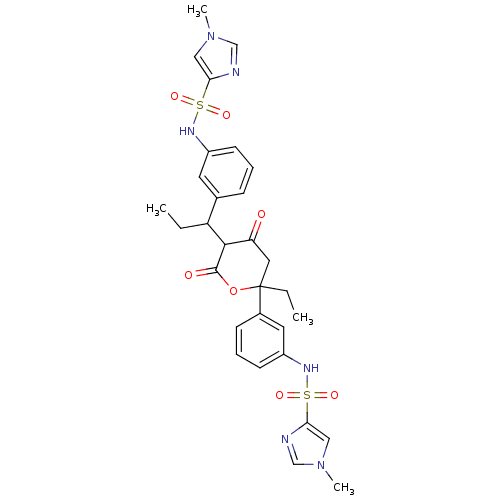
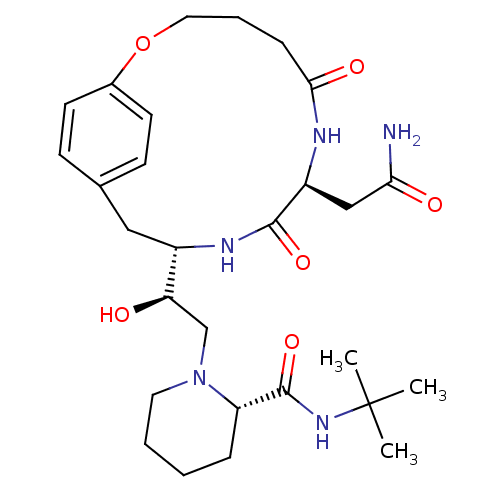
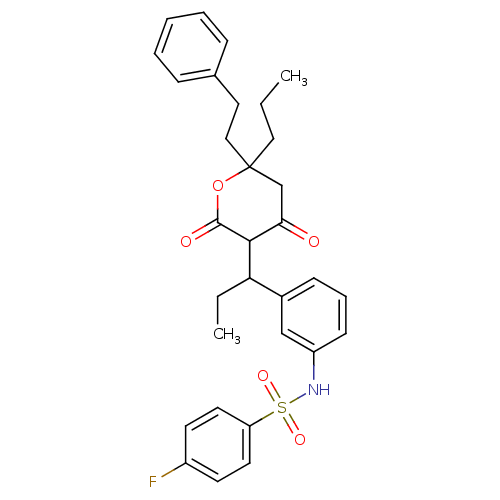
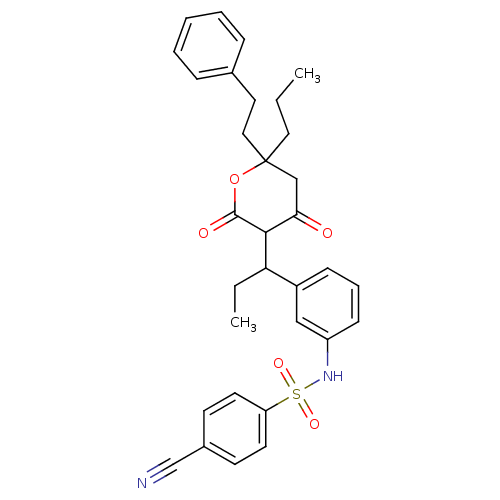
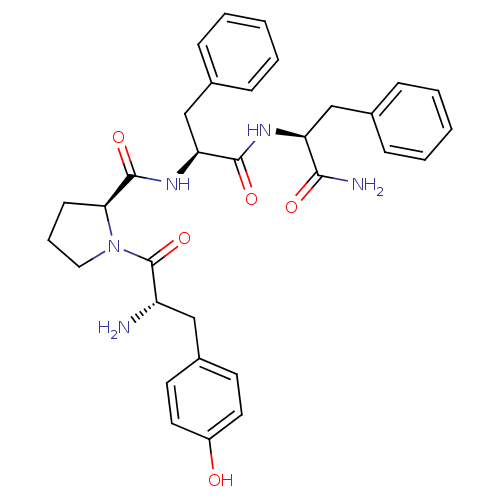
Affinity DataKi: 0.00700nM ΔG°: -63.0kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
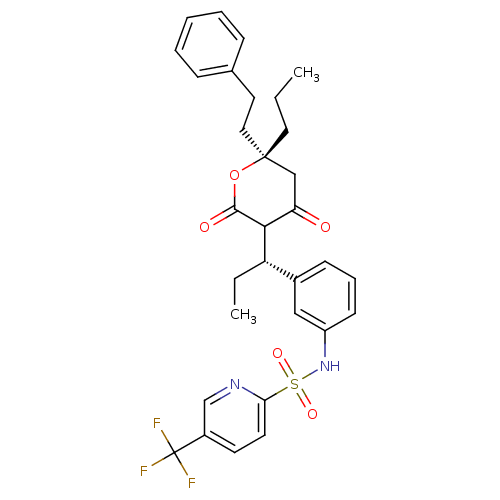
Affinity DataKi: 0.00800nM ΔG°: -62.7kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
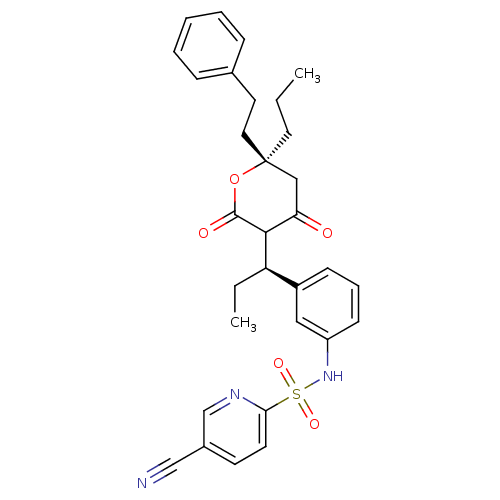
Affinity DataKi: 0.0180nM ΔG°: -60.7kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
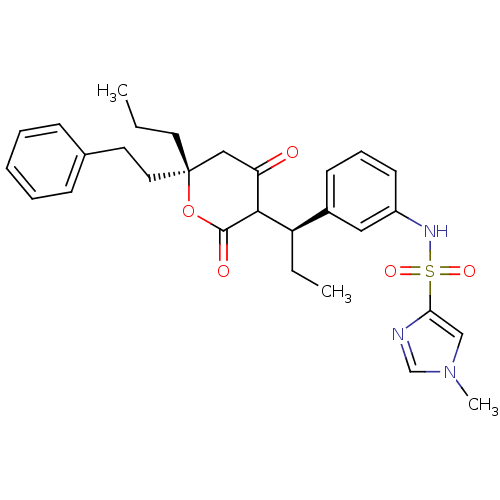
Affinity DataKi: 0.0320nM ΔG°: -59.3kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.0400nM ΔG°: -58.8kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.0600nM ΔG°: -57.8kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nM ΔG°: -56.5kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Pharmacia & Upjohn
Curated by ChEMBL
Pharmacia & Upjohn
Curated by ChEMBL
Affinity DataKi: 0.100nMAssay Description:Compound was evaluated for in vitro inhibition of human immunodeficiency virus type 1 (HIV-1) ProteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Pharmacia & Upjohn
Curated by ChEMBL
Pharmacia & Upjohn
Curated by ChEMBL
Affinity DataKi: 0.100nMAssay Description:Compound was evaluated for in vitro inhibition of human immunodeficiency virus type 1 (HIV-1) ProteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.100nM ΔG°: -56.5kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.120nM ΔG°: -56.1kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.120nM ΔG°: -56.1kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.190nM ΔG°: -54.9kJ/mole IC50: 0.5nMpH: 7.5 T: 2°CAssay Description:Test compounds were serially diluted in the assay buffer. In each well of a 96-well microtiter plate (Immunofluor B, Dynatech), the inhibitor solutio...More data for this Ligand-Target Pair
Affinity DataKi: 0.200nM ΔG°: -54.8kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.220nM ΔG°: -54.6kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.280nM ΔG°: -54.0kJ/mole IC50: 0.300nMpH: 7.5 T: 2°CAssay Description:Test compounds were serially diluted in the assay buffer. In each well of a 96-well microtiter plate (Immunofluor B, Dynatech), the inhibitor solutio...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nM ΔG°: -53.8kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.520nM ΔG°: -52.5kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.530nM ΔG°: -52.4kJ/mole IC50: 3.5nMpH: 7.5 T: 2°CAssay Description:Test compounds were serially diluted in the assay buffer. In each well of a 96-well microtiter plate (Immunofluor B, Dynatech), the inhibitor solutio...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
University of Queensland
University of Queensland
Affinity DataKi: 0.600nM ΔG°: -54.8kJ/molepH: 6.5 T: 2°CAssay Description:he Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in fl...More data for this Ligand-Target Pair
Affinity DataKi: 0.710nM ΔG°: -51.7kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.870nM ΔG°: -51.2kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -50.9kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -50.9kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -50.9kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Pharmacia & Upjohn
Curated by ChEMBL
Pharmacia & Upjohn
Curated by ChEMBL
Affinity DataKi: <1nMAssay Description:Compound was evaluated for in vitro inhibition of human immunodeficiency virus type 1 (HIV-1) ProteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Pharmacia & Upjohn
Curated by ChEMBL
Pharmacia & Upjohn
Curated by ChEMBL
Affinity DataKi: 1nMAssay Description:Compound was evaluated for in vitro inhibition of human immunodeficiency virus type 1 (HIV-1) ProteaseMore data for this Ligand-Target Pair
Affinity DataKi: 1.20nM ΔG°: -50.4kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 1.27nM ΔG°: -50.3kJ/mole IC50: 1.90nMpH: 7.5 T: 2°CAssay Description:Test compounds were serially diluted in the assay buffer. In each well of a 96-well microtiter plate (Immunofluor B, Dynatech), the inhibitor solutio...More data for this Ligand-Target Pair
Affinity DataKi: 1.70nM ΔG°: -49.6kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 1.80nM ΔG°: -49.4kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
University of Queensland
University of Queensland
Affinity DataKi: 1.80nM ΔG°: -51.9kJ/molepH: 6.5 T: 2°CAssay Description:he Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in fl...More data for this Ligand-Target Pair
Affinity DataKi: 1.90nM ΔG°: -49.3kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 2.30nM ΔG°: -48.8kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 2.60nM ΔG°: -48.5kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -48.2kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
University of Queensland
University of Queensland
Affinity DataKi: 3nM ΔG°: -50.6kJ/molepH: 6.5 T: 2°CAssay Description:he Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in fl...More data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -48.2kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -48.2kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Pharmacia & Upjohn
Curated by ChEMBL
Pharmacia & Upjohn
Curated by ChEMBL
Affinity DataKi: 3nMAssay Description:Compound was evaluated for in vitro inhibition of human immunodeficiency virus type 1 (HIV-1) ProteaseMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
University of Queensland
University of Queensland
Affinity DataKi: 4nM ΔG°: -49.9kJ/molepH: 6.5 T: 2°CAssay Description:he Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in fl...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
University of Queensland
University of Queensland
Affinity DataKi: 4nM ΔG°: -49.9kJ/molepH: 6.5 T: 2°CAssay Description:he Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in fl...More data for this Ligand-Target Pair
Affinity DataKi: 4.30nM ΔG°: -47.3kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Pharmacia & Upjohn
Curated by ChEMBL
Pharmacia & Upjohn
Curated by ChEMBL
Affinity DataKi: 4.70nMAssay Description:Compound was evaluated for in vitro inhibition of human immunodeficiency virus type 1 (HIV-1) ProteaseMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
University of Queensland
University of Queensland
Affinity DataKi: 5nM ΔG°: -49.3kJ/molepH: 6.5 T: 2°CAssay Description:he Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in fl...More data for this Ligand-Target Pair
Affinity DataKi: 6nM ΔG°: -46.5kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Pharmacia & Upjohn
Curated by ChEMBL
Pharmacia & Upjohn
Curated by ChEMBL
Affinity DataKi: 8nMAssay Description:Compound was evaluated for in vitro inhibition of human immunodeficiency virus type 1 (HIV-1) ProteaseMore data for this Ligand-Target Pair
Affinity DataKi: 8nM ΔG°: -45.7kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 8.20nM ΔG°: -45.7kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 8.23nMAssay Description:Displacement of [3H]DAMGO from mu opioid receptor in rat brain membrane after 3 hrs by liquid scintillation countingMore data for this Ligand-Target Pair

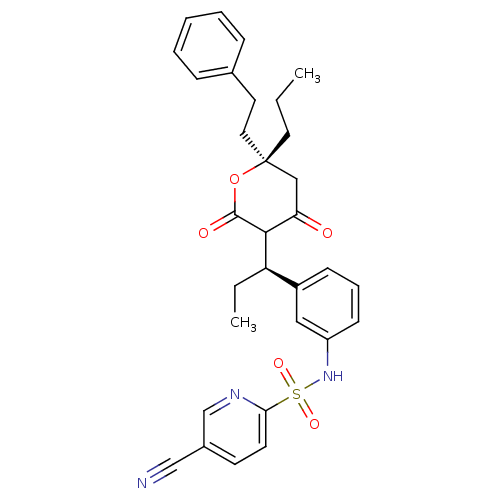
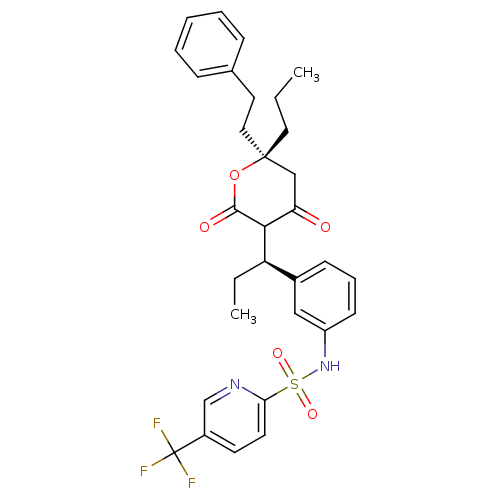
 3D Structure (crystal)
3D Structure (crystal)