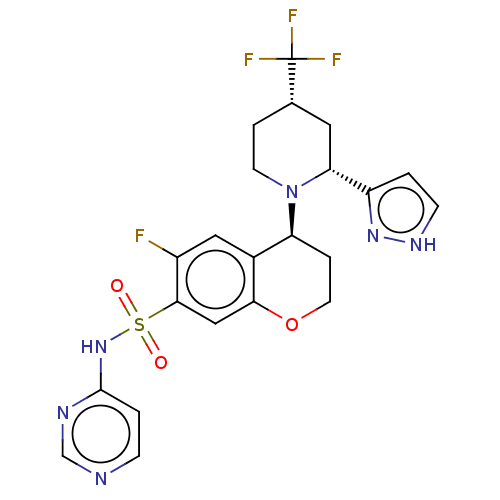
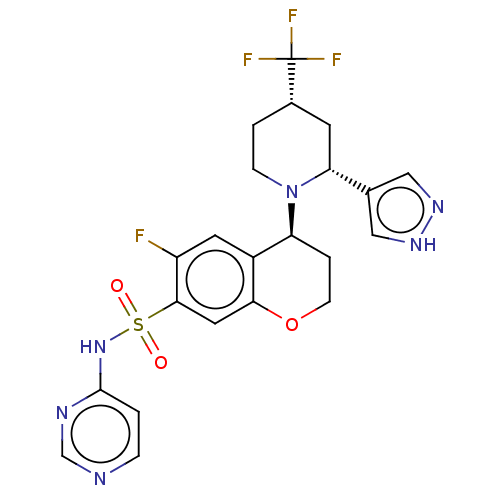
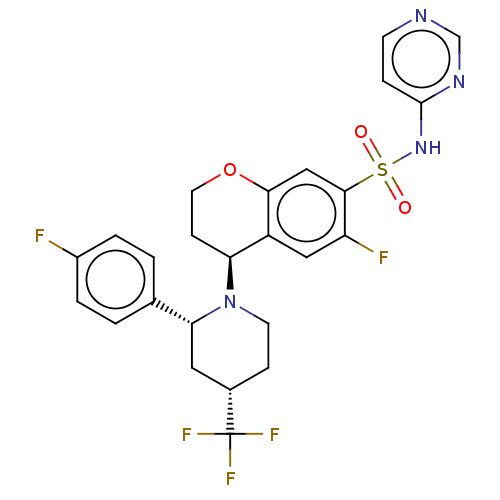
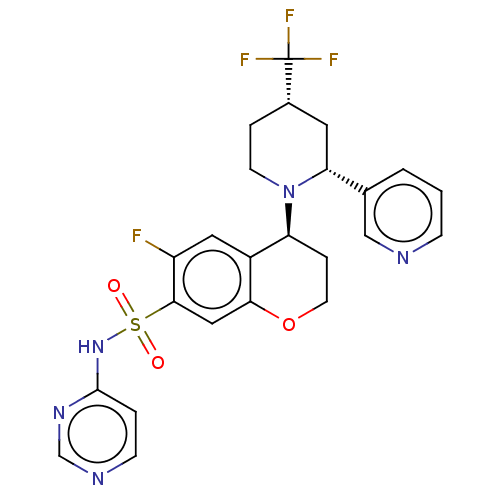
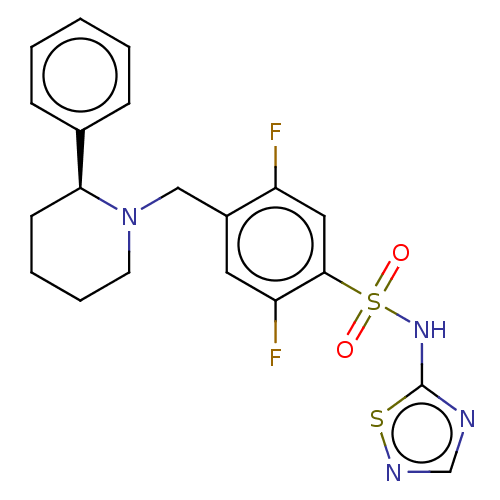
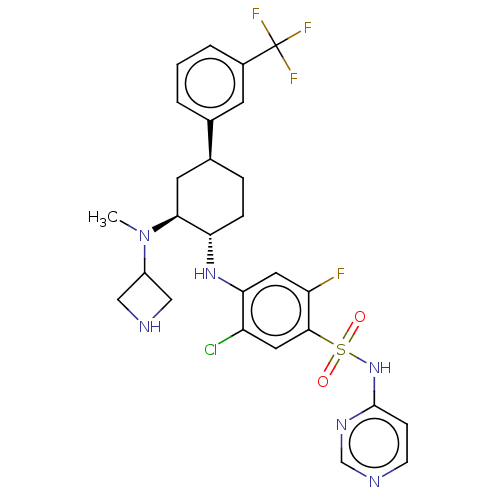
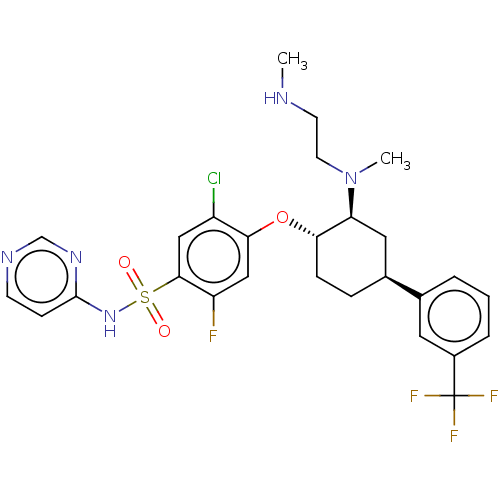
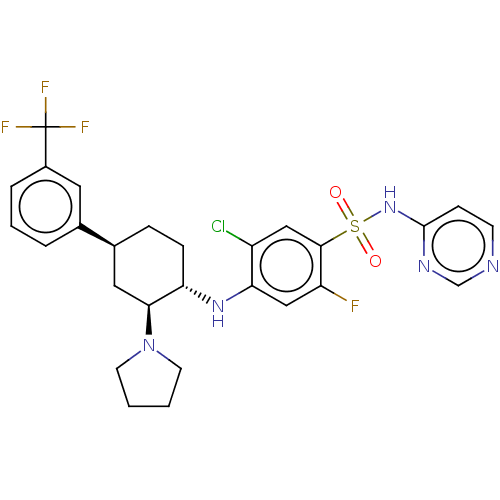
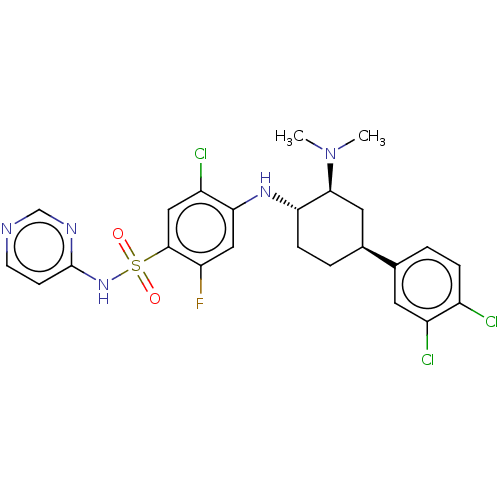
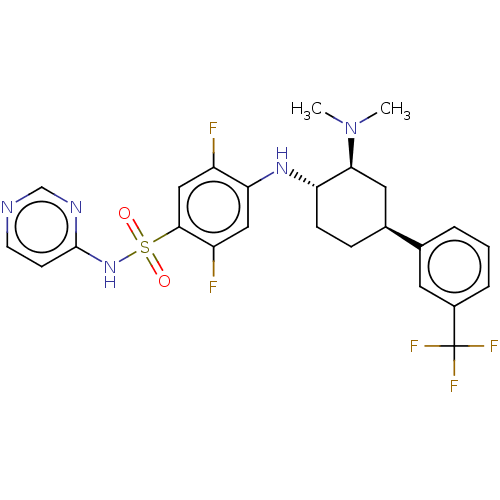
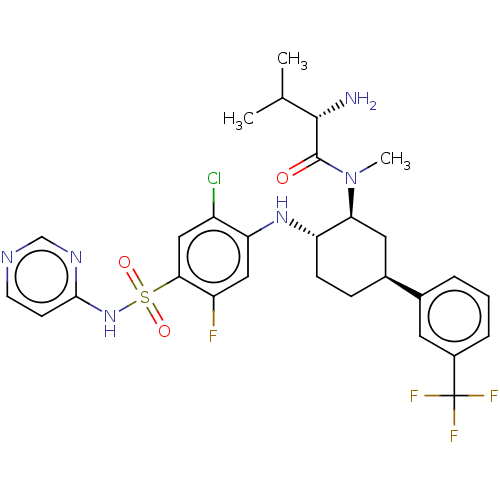
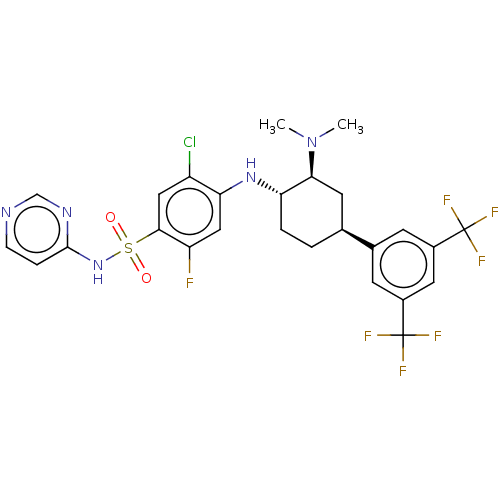
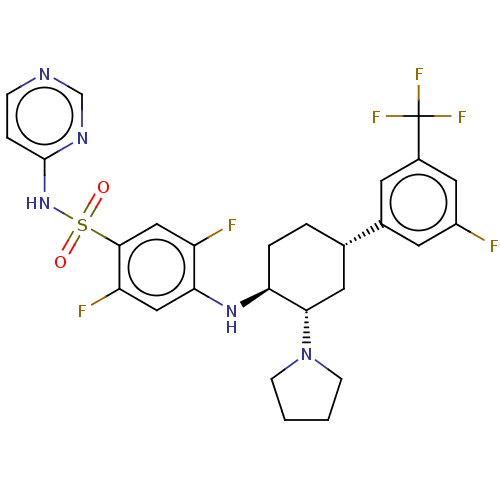
Affinity DataKi: 0.320nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
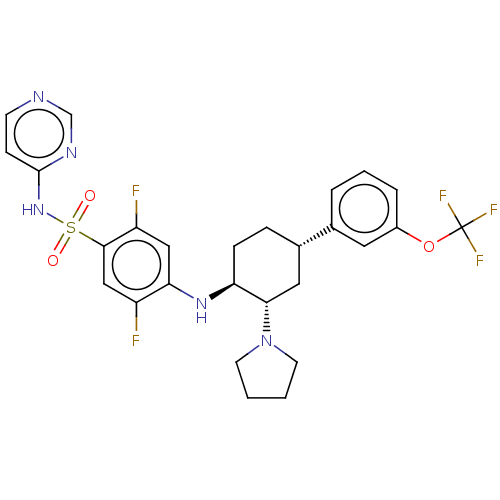
Affinity DataKi: 0.510nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
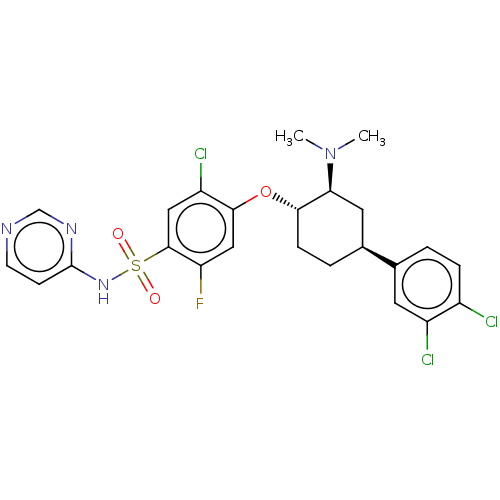
Affinity DataKi: 0.530nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
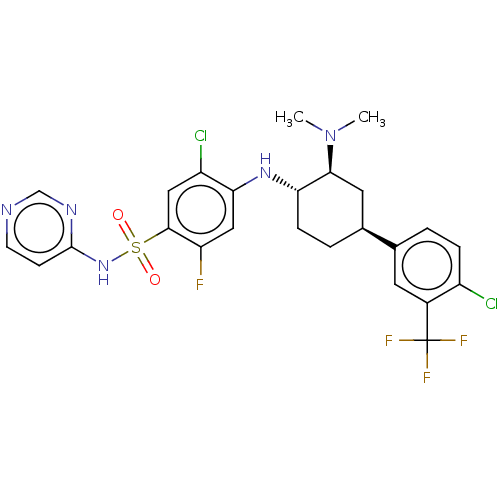
Affinity DataKi: 0.680nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 0.760nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 0.790nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 0.850nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 0.930nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 0.970nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 1.70nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 2.90nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 2.90nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 3.90nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 72nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 509nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: >700nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: >700nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataIC50: 0.141nMAssay Description:The following voltage clamp electrophysiology studies are performed on representative compounds using cells heterologously expressing Nav1.7 or Nav1....More data for this Ligand-Target Pair
Affinity DataIC50: 0.173nMAssay Description:The following voltage clamp electrophysiology studies are performed on representative compounds using cells heterologously expressing Nav1.7 or Nav1....More data for this Ligand-Target Pair
Affinity DataIC50: 0.182nMAssay Description:Patch voltage clamp electrophysiology allows for the direct measurement and quantification of block of voltage-gated sodium channels (NaV's), and...More data for this Ligand-Target Pair
Affinity DataIC50: 0.241nMAssay Description:The following voltage clamp electrophysiology studies are performed on representative compounds using cells heterologously expressing Nav1.7 or Nav1....More data for this Ligand-Target Pair
Affinity DataIC50: 0.255nMAssay Description:Preparation of membranes containing recombinantly expressed sodium channels: Following the procedure described in Example 243A, and making modificati...More data for this Ligand-Target Pair
Affinity DataIC50: 0.263nMAssay Description:The following voltage clamp electrophysiology studies are performed on representative compounds using cells heterologously expressing Nav1.7 or Nav1....More data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:Inhibition of human Nav1.7 expressed in HEK293 cells at holding potential of -120 mV by manual electrophysiology methodMore data for this Ligand-Target Pair
Affinity DataIC50: 0.305nMAssay Description:The following voltage clamp electrophysiology studies are performed on representative compounds using cells heterologously expressing Nav1.7 or Nav1....More data for this Ligand-Target Pair
Affinity DataIC50: 0.317nMAssay Description:The following voltage clamp electrophysiology studies are performed on representative compounds using cells heterologously expressing Nav1.7 or Nav1....More data for this Ligand-Target Pair
Affinity DataIC50: 0.322nMAssay Description:The following voltage clamp electrophysiology studies are performed on representative compounds using cells heterologously expressing Nav1.7 or Nav1....More data for this Ligand-Target Pair
Affinity DataIC50: 0.325nMAssay Description:The following voltage clamp electrophysiology studies are performed on representative compounds using cells heterologously expressing Nav1.7 or Nav1....More data for this Ligand-Target Pair
Affinity DataIC50: 0.327nMAssay Description:The following voltage clamp electrophysiology studies are performed on representative compounds using cells heterologously expressing Nav1.7 or Nav1....More data for this Ligand-Target Pair
Affinity DataIC50: 0.334nMAssay Description:The following voltage clamp electrophysiology studies are performed on representative compounds using cells heterologously expressing Nav1.7 or Nav1....More data for this Ligand-Target Pair
Affinity DataIC50: 0.355nMAssay Description:The following voltage clamp electrophysiology studies are performed on representative compounds using cells heterologously expressing Nav1.7 or Nav1....More data for this Ligand-Target Pair
Affinity DataIC50: 0.372nMAssay Description:Patch voltage clamp electrophysiology allows for the direct measurement and quantification of block of voltage-gated sodium channels (NaV's), and...More data for this Ligand-Target Pair
Affinity DataIC50: 0.383nMAssay Description:The following voltage clamp electrophysiology studies are performed on representative compounds using cells heterologously expressing Nav1.7 or Nav1....More data for this Ligand-Target Pair
Affinity DataIC50: 0.390nMAssay Description:Saturation experiments. A competitive NaV1.7 inhibitor having a methyl group is tritiated. Three tritiums are incorporated in place of methyl hydroge...More data for this Ligand-Target Pair
Affinity DataIC50: 0.399nMAssay Description:Saturation experiments. A competitive NaV1.7 inhibitor having a methyl group is tritiated. Three tritiums are incorporated in place of methyl hydroge...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inhibition of full length human Nav1.7 expressed in HEK cells by whole cell voltage clamp analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inhibition of Nav1.7 (unknown origin) by electrophysiology assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.418nMAssay Description:Patch voltage clamp electrophysiology allows for the direct measurement and quantification of block of voltage-gated sodium channels (NaV's), and...More data for this Ligand-Target Pair
Affinity DataIC50: 0.428nMAssay Description:Preparation of membranes containing recombinantly expressed sodium channels: Following the procedure described in Example 243A, and making modificati...More data for this Ligand-Target Pair
Affinity DataIC50: 0.462nMAssay Description:The following voltage clamp electrophysiology studies are performed on representative compounds using cells heterologously expressing Nav1.7 or Nav1....More data for this Ligand-Target Pair
Affinity DataIC50: 0.473nMAssay Description:Saturation experiments. A competitive NaV1.7 inhibitor having a methyl group is tritiated. Three tritiums are incorporated in place of methyl hydroge...More data for this Ligand-Target Pair
Affinity DataIC50: 0.474nMAssay Description:Saturation experiments. A competitive NaV1.7 inhibitor having a methyl group is tritiated. Three tritiums are incorporated in place of methyl hydroge...More data for this Ligand-Target Pair
Affinity DataIC50: 0.494nMAssay Description:The following voltage clamp electrophysiology studies are performed on representative compounds using cells heterologously expressing Nav1.7 or Nav1....More data for this Ligand-Target Pair
Affinity DataIC50: 0.5nMAssay Description:Binding affinity to recombinant human Nav1.7 after 18 hrs by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.519nMAssay Description:The following voltage clamp electrophysiology studies are performed on representative compounds using cells heterologously expressing Nav1.7 or Nav1....More data for this Ligand-Target Pair
Affinity DataIC50: 0.520nMAssay Description:Saturation experiments. A competitive NaV1.7 inhibitor having a methyl group is tritiated. Three tritiums are incorporated in place of methyl hydroge...More data for this Ligand-Target Pair