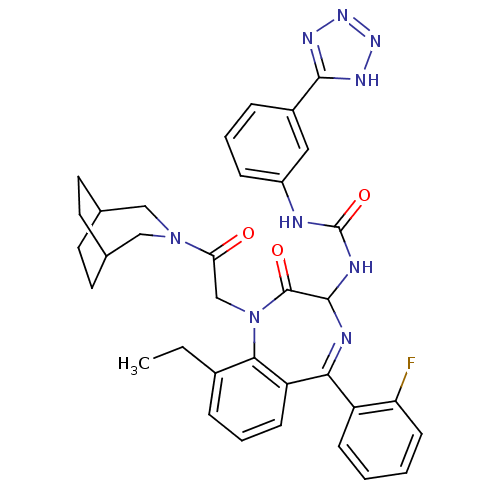
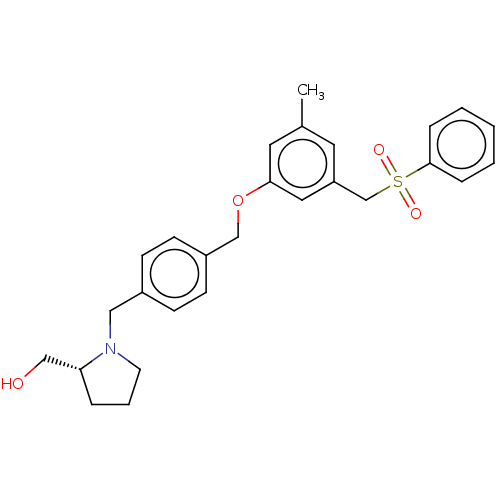
Affinity DataKi: 0.0330nM ΔG°: -59.2kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 4.90nM ΔG°: -47.0kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 7.5nM ΔG°: -45.9kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 7.70nM ΔG°: -45.8kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 7.70nM ΔG°: -45.8kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 7.70nM ΔG°: -45.8kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 9.80nM ΔG°: -45.3kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 11nM ΔG°: -45.0kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 11nM ΔG°: -45.0kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 12nM ΔG°: -44.8kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 13nM ΔG°: -44.6kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 13nM ΔG°: -44.6kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 13nM ΔG°: -44.6kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 16nM ΔG°: -44.0kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 17nM ΔG°: -43.9kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 17nM ΔG°: -43.9kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 24nM ΔG°: -43.1kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 26nM ΔG°: -42.9kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 30nM ΔG°: -42.5kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 34nM ΔG°: -42.2kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 37nM ΔG°: -42.0kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 38nM ΔG°: -41.9kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 40nMAssay Description:Inhibition of recombinant human SphK1 expressed in baculovirus infected sf9 cells using D-erythro-sphingosine as substrate in presence of [gamma-33P]...More data for this Ligand-Target Pair
Affinity DataKi: 55nM ΔG°: -41.0kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 57nM ΔG°: -40.9kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 91nM ΔG°: -39.8kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 190nMAssay Description:Inhibition of recombinant human maltase-glucoamylase using p-nitrophenyl-alpha-D-glucopyranoside as substrate incubated for 35 mins by microtiter pla...More data for this Ligand-Target Pair
Affinity DataKi: 240nM ΔG°: -37.4kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 680nM ΔG°: -34.8kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 1.20E+3nM ΔG°: -33.5kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 1.20E+3nMAssay Description:Inhibition of recombinant human SphK2 expressed in baculovirus infected sf9 cells using D-erythro-sphingosine as substrate after 20 mins in presence ...More data for this Ligand-Target Pair
Affinity DataKi: 1.30E+3nM ΔG°: -33.3kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 1.90E+3nM ΔG°: -32.3kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 5.90E+3nM ΔG°: -29.5kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 5.90E+3nM ΔG°: -29.5kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 6.00E+3nMAssay Description:Inhibition of recombinant human maltase-glucoamylase using p-nitrophenyl-alpha-D-glucopyranoside as substrate incubated for 35 mins by microtiter pla...More data for this Ligand-Target Pair
Affinity DataKi: 6.20E+3nM ΔG°: -29.4kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Inhibition of recombinant human SphK1 expressed in baculovirus infected sf9 cells using D-erythro-sphingosine as substrate after 20 mins in presence ...More data for this Ligand-Target Pair
Affinity DataKi: 1.41E+4nMAssay Description:Inhibition of recombinant mouse SphK2 expressed in baculovirus infected sf9 cells using D-erythro-sphingosine as substrate in presence of [gamma-33P]...More data for this Ligand-Target Pair
Affinity DataKi: 3.20E+4nMAssay Description:Inhibition of recombinant human maltase-glucoamylase using p-nitrophenyl-alpha-D-glucopyranoside as substrate incubated for 35 mins by microtiter pla...More data for this Ligand-Target Pair
Affinity DataKi: 5.10E+4nMAssay Description:Inhibition of recombinant human maltase-glucoamylase using p-nitrophenyl-alpha-D-glucopyranoside as substrate incubated for 35 mins by microtiter pla...More data for this Ligand-Target Pair
Affinity DataKi: 5.40E+4nM ΔG°: -24.1kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nM ΔG°: >-22.6kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nM ΔG°: >-22.6kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nM ΔG°: >-22.6kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
TargetGastrin/cholecystokinin type B receptor(Homo sapiens (Human))
Fujisawa Pharmaceutical
Curated by ChEMBL
Fujisawa Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 0.0870nMAssay Description:Compound was evaluated for the inhibition of 124 I-CCK-8 binding at Cholecystokinin type B receptor on guinea pig cerebral cortical membranesMore data for this Ligand-Target Pair
TargetGastrin/cholecystokinin type B receptor(Homo sapiens (Human))
Fujisawa Pharmaceutical
Curated by ChEMBL
Fujisawa Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 0.660nMAssay Description:Compound was evaluated for the inhibition of 124 I-CCK-8 binding at CCK-B receptorCholecystokinin type B receptorcerebral cortical membranesMore data for this Ligand-Target Pair
TargetGastrin/cholecystokinin type B receptor(Homo sapiens (Human))
Fujisawa Pharmaceutical
Curated by ChEMBL
Fujisawa Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 0.680nMAssay Description:Compound was evaluated for the inhibition of 124 I-CCK-8 binding at Cholecystokinin type B receptor on guinea pig cerebral cortical membranesMore data for this Ligand-Target Pair
TargetGastrin/cholecystokinin type B receptor(Homo sapiens (Human))
Fujisawa Pharmaceutical
Curated by ChEMBL
Fujisawa Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 0.810nMAssay Description:Compound was evaluated for the inhibition of 124 I-CCK-8 binding at Cholecystokinin type B receptor on guinea pig cerebral cortical membranesMore data for this Ligand-Target Pair
Affinity DataIC50: 2.70nMAssay Description:Inhibition of recombinant C-terminal His6-tagged human SphK1 expressed in baculovirus infected sf21 cells using FITC-sphingosine as substrate after 1...More data for this Ligand-Target Pair

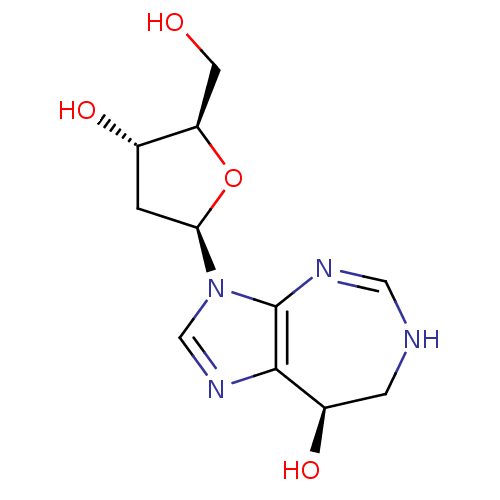
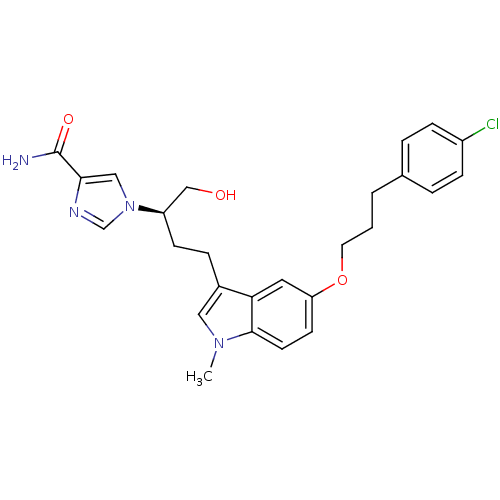
 3D Structure (crystal)
3D Structure (crystal)