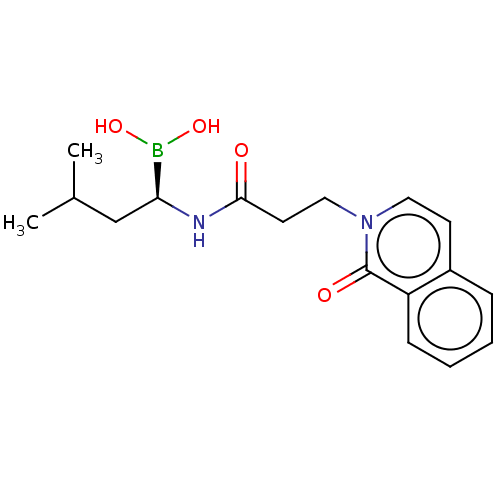
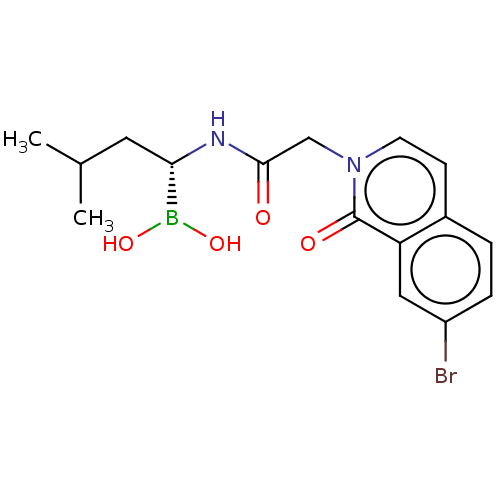
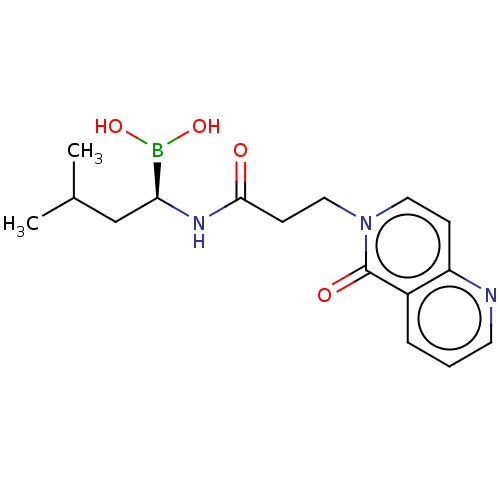
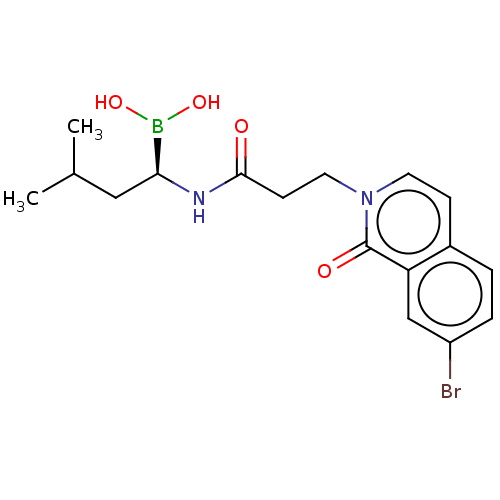
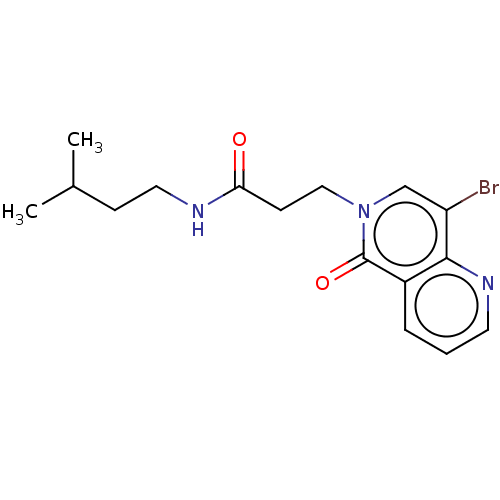
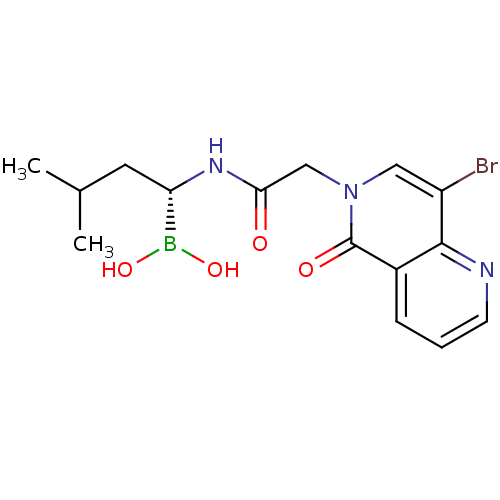
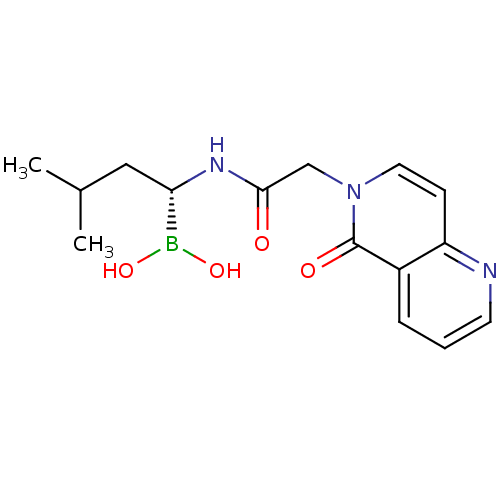
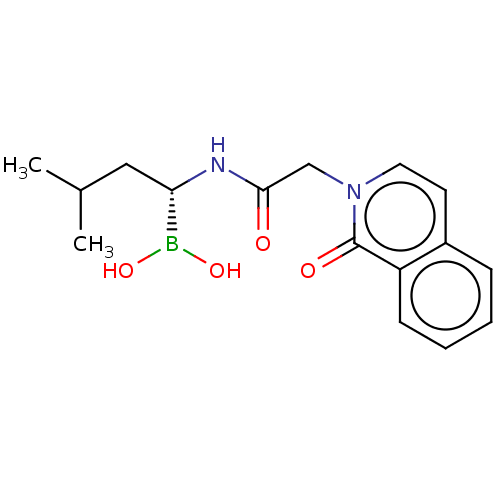
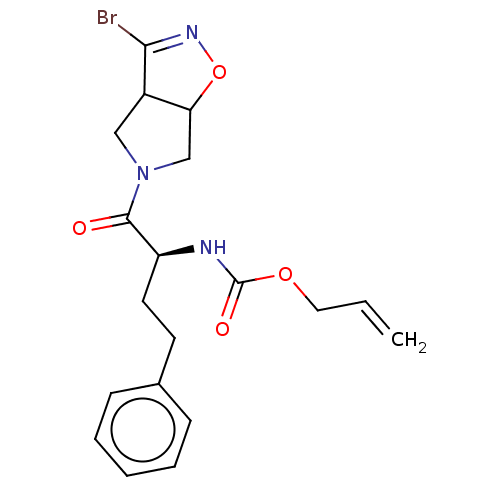
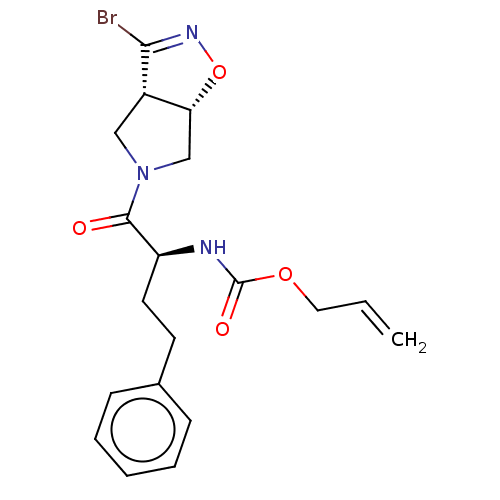
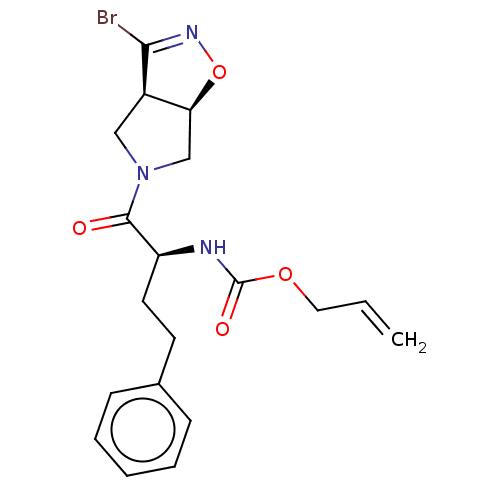
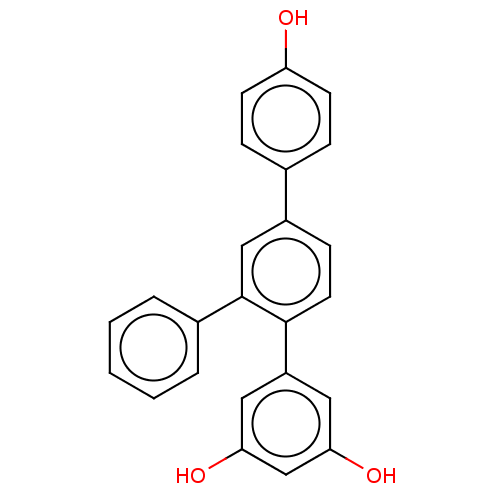
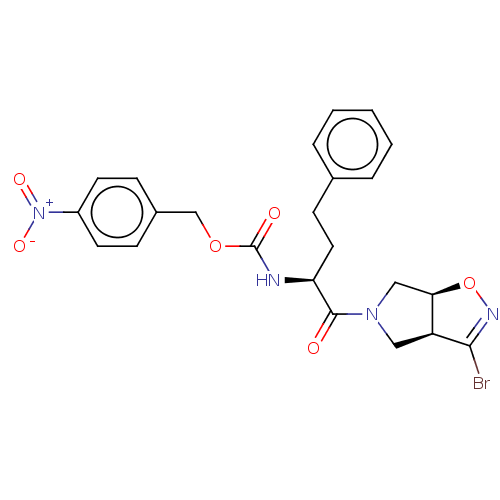
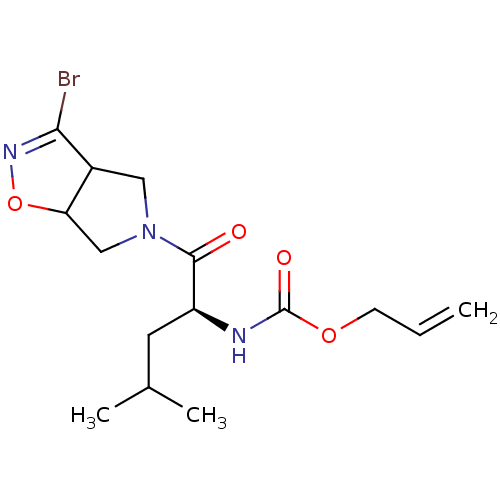
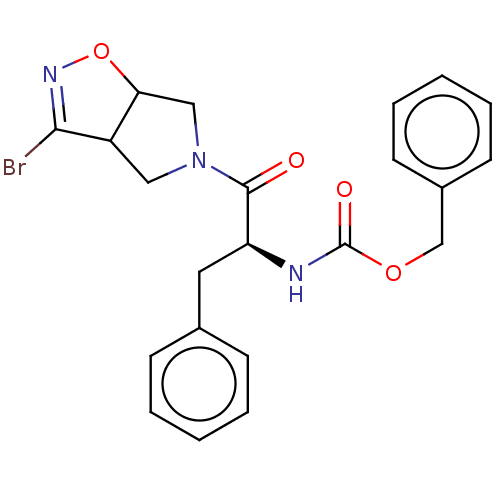
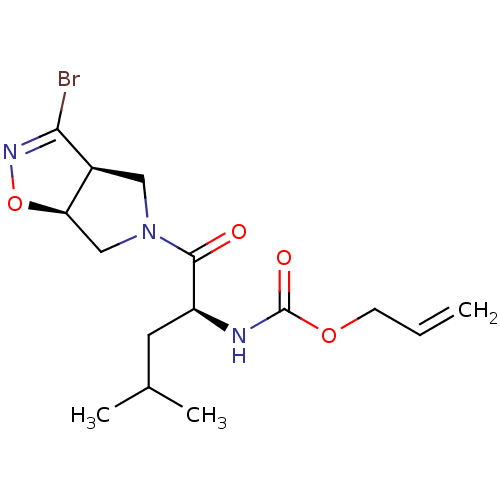
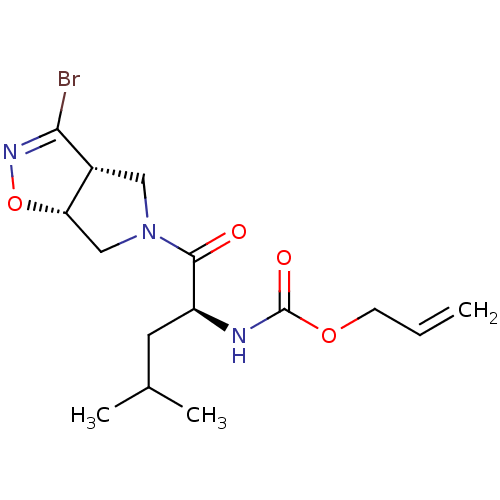
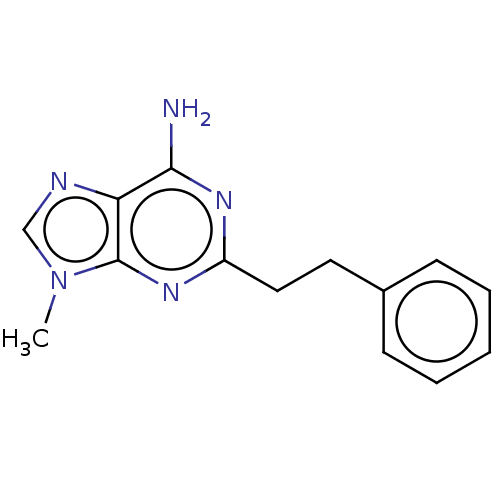
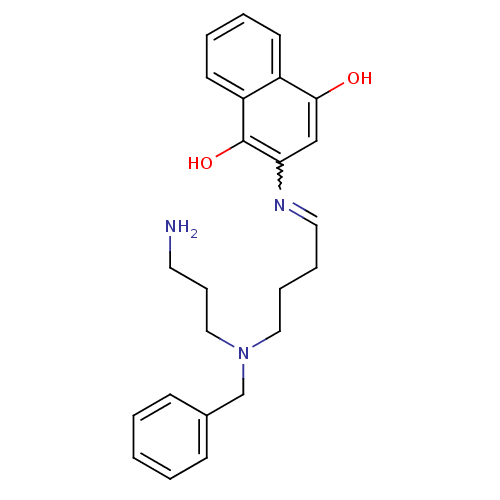
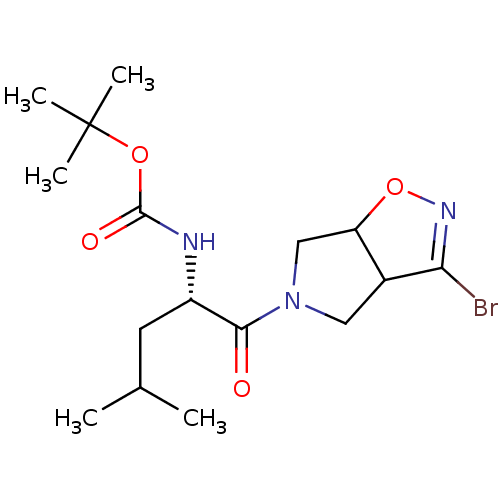
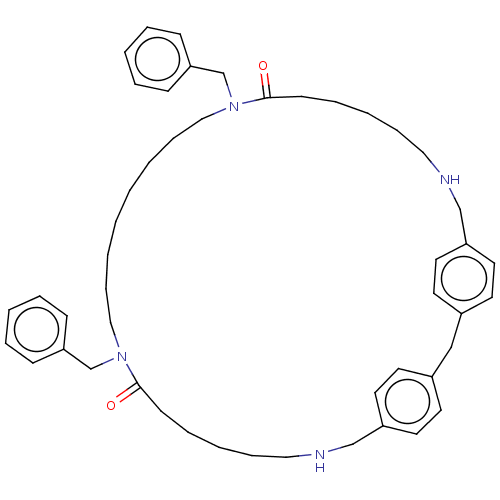
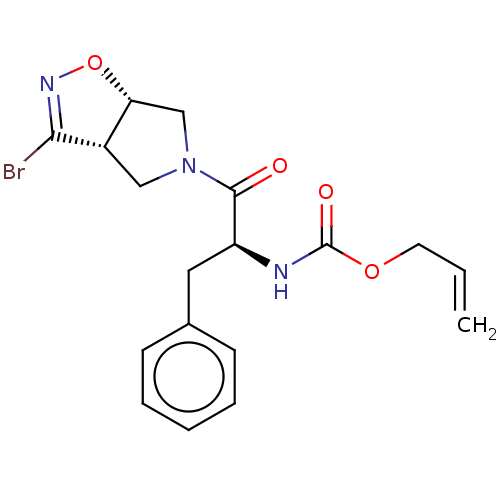
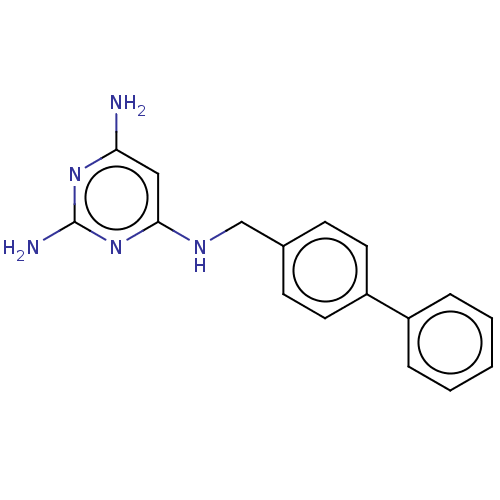
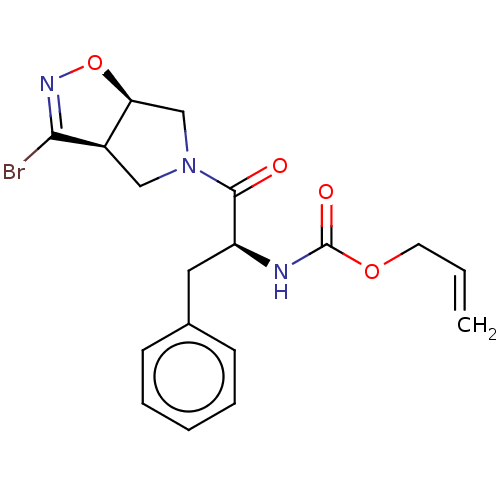
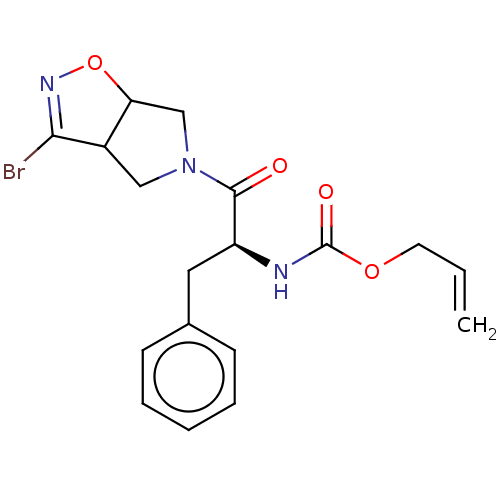
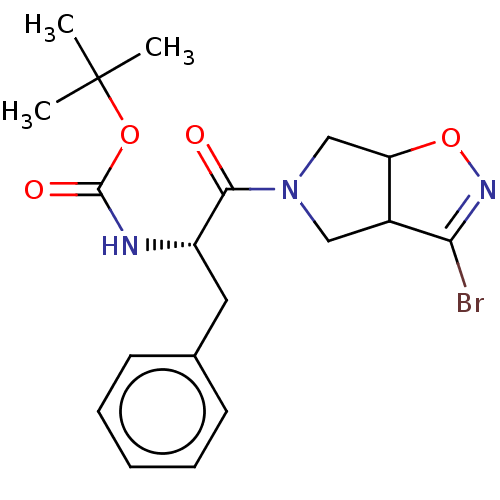
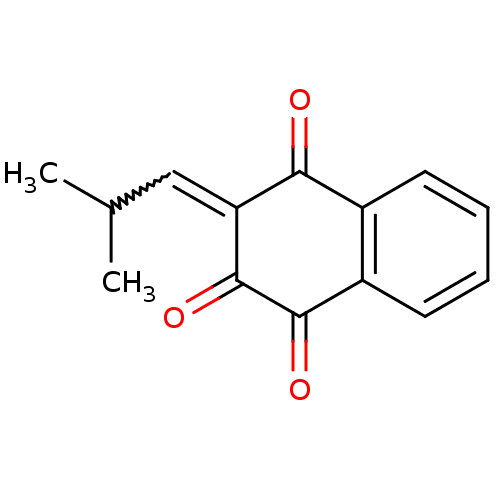
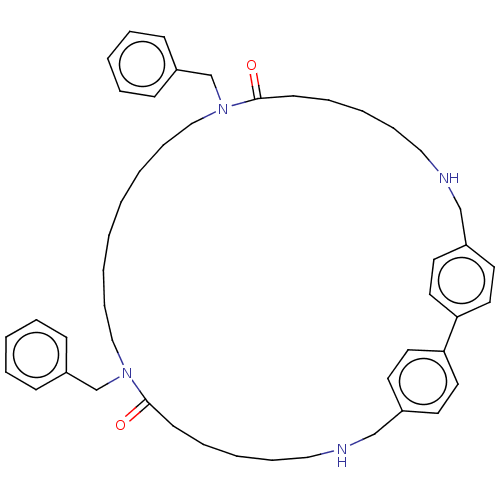
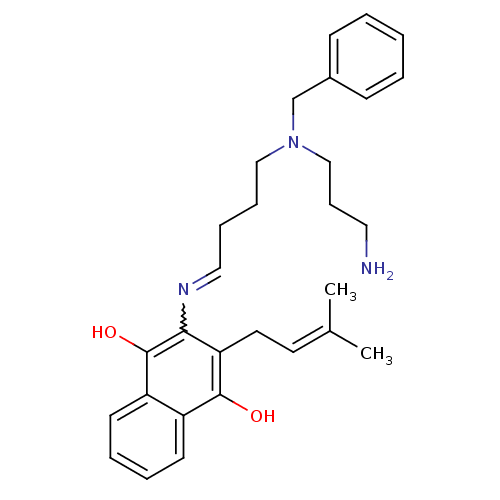
Affinity DataKi: 9.80nMAssay Description:Inhibition of chymotrypsin-like activity of 20S proteasome in human erythrocytes using Suc-Leu-Leu-Val-Tyr-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
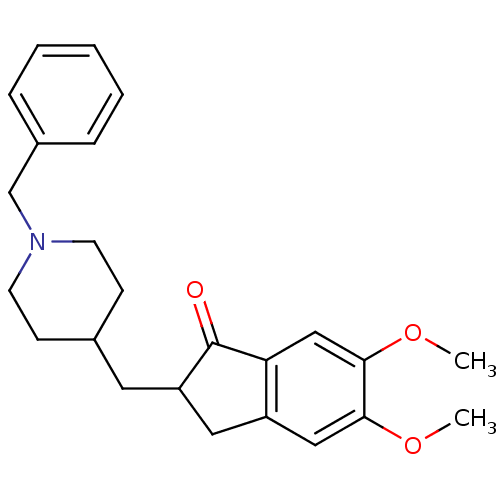
Affinity DataKi: 13nMAssay Description:Mixed type inhibition of human AChE using acetylthiocholineiodide as substrate measured for every 30 sec for 5 mins by Ellman's methodMore data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:Mixed type inhibition of human AChE using acetylthiocholineiodide as substrate measured for every 30 sec for 5 mins by Ellman's methodMore data for this Ligand-Target Pair
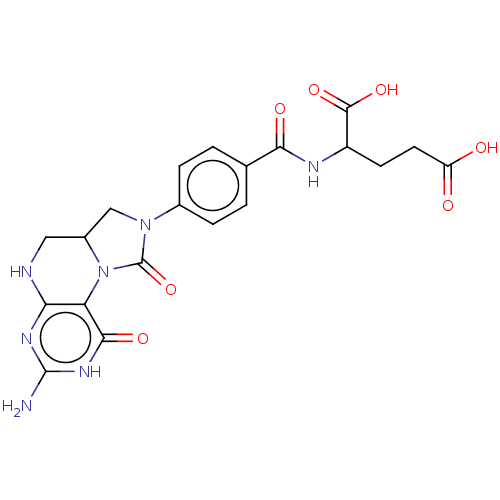
TargetBifunctional methylenetetrahydrofolate dehydrogenase/cyclohydrolase, mitochondrial(Homo sapiens (Human))
University Of Dundee
Curated by ChEMBL
University Of Dundee
Curated by ChEMBL
Affinity DataKi: 29nMAssay Description:Competitive inhibition of human FolD dehydrogenase activityMore data for this Ligand-Target Pair
Affinity DataKi: 51nMAssay Description:Inhibition of chymotrypsin-like activity of 20S proteasome in human erythrocytes using Suc-Leu-Leu-Val-Tyr-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 53nMAssay Description:Inhibition of chymotrypsin-like activity of 20S proteasome in human erythrocytes using Suc-Leu-Leu-Val-Tyr-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 57nMAssay Description:Inhibition of chymotrypsin-like activity of 20S proteasome in human erythrocytes using Suc-Leu-Leu-Val-Tyr-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 60nMAssay Description:Inhibition of chymotrypsin-like activity of 20S proteasome in human erythrocytes using Suc-Leu-Leu-Val-Tyr-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 80nMAssay Description:Inhibition of chymotrypsin-like activity of 20S proteasome in human erythrocytes using Suc-Leu-Leu-Val-Tyr-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 85nMAssay Description:Inhibition of chymotrypsin-like activity of 20S proteasome in human erythrocytes using Suc-Leu-Leu-Val-Tyr-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 93nMAssay Description:Mixed type inhibition of human AChE using acetylthiocholineiodide as substrate measured for every 30 sec for 5 mins by Ellman's methodMore data for this Ligand-Target Pair
Affinity DataKi: 93.4nMAssay Description:Mixed type inhibition of human AChE using acetylthiocholineiodide as substrate measured for every 30 sec for 5 mins by Ellman's methodMore data for this Ligand-Target Pair
Affinity DataKi: 170nMAssay Description:Inhibition of chymotrypsin-like activity of 20S proteasome in human erythrocytes using Suc-Leu-Leu-Val-Tyr-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 240nMAssay Description:Inhibition of PGPH-like activity of 20S proteasome in human erythrocytes using Z-Leu-Leu-Glu-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 290nMAssay Description:Inhibition of bovine pancreatic alpha-chymotrypsin using Suc-Leu-Leu-Val-Tyr-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 310nMAssay Description:Inhibition of PGPH-like activity of 20S proteasome in human erythrocytes using Z-Leu-Leu-Glu-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 440nMAssay Description:Inhibition of chymotrypsin-like activity of 20S proteasome in human erythrocytes using Suc-Leu-Leu-Val-Tyr-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 490nMAssay Description:Inhibition of chymotrypsin-like activity of 20S proteasome in human erythrocytes using Suc-Leu-Leu-Val-Tyr-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 690nMAssay Description:Inhibition of PGPH-like activity of 20S proteasome in human erythrocytes using Z-Leu-Leu-Glu-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 990nMAssay Description:Inhibition of PGPH-like activity of 20S proteasome in human erythrocytes using Z-Leu-Leu-Glu-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.25E+3nMAssay Description:Inhibition of PGPH-like activity of 20S proteasome in human erythrocytes using Z-Leu-Leu-Glu-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.55E+3nMAssay Description:Inhibition of PGPH-like activity of 20S proteasome in human erythrocytes using Z-Leu-Leu-Glu-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.04E+3nMAssay Description:Inhibition of PGPH-like activity of 20S proteasome in human erythrocytes using Z-Leu-Leu-Glu-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.34E+3nMAssay Description:Inhibition of chymotrypsin-like activity of 20S proteasome in human erythrocytes using Suc-Leu-Leu-Val-Tyr-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.40E+3nMAssay Description:Inhibition of bovine pancreatic alpha-chymotrypsin using Suc-Leu-Leu-Val-Tyr-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.62E+3nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-7-amino-4-methylcoumarin assessed as substrate hydrolysis measured by fluore...More data for this Ligand-Target Pair
Affinity DataKi: 2.73E+3nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-7-amino-4-methylcoumarin assessed as substrate hydrolysis measured by fluore...More data for this Ligand-Target Pair
Affinity DataKi: 3.35E+3nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-7-amino-4-methylcoumarin assessed as substrate hydrolysis measured by fluore...More data for this Ligand-Target Pair
Affinity DataKi: 4.57E+3nMAssay Description:Inhibition of chymotrypsin-like activity of 20S proteasome in human erythrocytes using Suc-Leu-Leu-Val-Tyr-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
TargetPteridine reductase 1(Leishmania major)
University Of Modena And Reggio Emilia
Curated by ChEMBL
University Of Modena And Reggio Emilia
Curated by ChEMBL
Affinity DataKi: 5.70E+3nMAssay Description:Inhibition of Leishmania major PTR1 using H2B as substrate preincubated for 10 mins followed by NADPH addition and measured up to 50 mins relative to...More data for this Ligand-Target Pair
Affinity DataKi: 6.81E+3nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-7-amino-4-methylcoumarin assessed as substrate hydrolysis measured by fluore...More data for this Ligand-Target Pair
Affinity DataKi: 8.36E+3nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-7-amino-4-methylcoumarin assessed as substrate hydrolysis measured by fluore...More data for this Ligand-Target Pair
Affinity DataKi: 8.41E+3nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-7-amino-4-methylcoumarin assessed as substrate hydrolysis measured by fluore...More data for this Ligand-Target Pair
Affinity DataKi: 8.52E+3nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-7-amino-4-methylcoumarin assessed as substrate hydrolysis measured by fluore...More data for this Ligand-Target Pair
Affinity DataKi: 9.05E+3nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-7-amino-4-methylcoumarin assessed as substrate hydrolysis measured by fluore...More data for this Ligand-Target Pair
Affinity DataKi: 9.11E+3nMAssay Description:Inhibition of bovine pancreatic alpha-chymotrypsin using Suc-Leu-Leu-Val-Tyr-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
TargetPteridine reductase 1(Leishmania major)
University Of Modena And Reggio Emilia
Curated by ChEMBL
University Of Modena And Reggio Emilia
Curated by ChEMBL
Affinity DataKi: 1.20E+4nMAssay Description:Inhibition of Leishmania major PTR1 using H2B as substrate preincubated for 10 mins followed by NADPH addition and measured up to 50 mins relative to...More data for this Ligand-Target Pair
Affinity DataKi: 1.20E+4nM ΔG°: -29.2kJ/molepH: 7.4 T: 2°CAssay Description:The reaction mixture (final volume 200 µL) contained 5 µg/mL MAO-A or MAO-B in a 50 mM sodium phosphate buffer pH 7.4, 1 mM p-tyramine (sub...More data for this Ligand-Target Pair
Affinity DataKi: 1.36E+4nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-7-amino-4-methylcoumarin assessed as substrate hydrolysis measured by fluore...More data for this Ligand-Target Pair
TargetPteridine reductase 1(Leishmania major)
University Of Modena And Reggio Emilia
Curated by ChEMBL
University Of Modena And Reggio Emilia
Curated by ChEMBL
Affinity DataKi: 1.48E+4nMAssay Description:Inhibition of Leishmania major PTR1 using H2B as substrate preincubated for 10 mins followed by NADPH addition and measured up to 50 mins relative to...More data for this Ligand-Target Pair
Affinity DataKi: 1.51E+4nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-7-amino-4-methylcoumarin assessed as substrate hydrolysis measured by fluore...More data for this Ligand-Target Pair
TargetPteridine reductase 1(Leishmania major)
University Of Modena And Reggio Emilia
Curated by ChEMBL
University Of Modena And Reggio Emilia
Curated by ChEMBL
Affinity DataKi: 1.53E+4nMAssay Description:Inhibition of Leishmania major PTR1 using H2B as substrate preincubated for 10 mins followed by NADPH addition and measured up to 50 mins relative to...More data for this Ligand-Target Pair
Affinity DataKi: 1.66E+4nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-7-amino-4-methylcoumarin assessed as substrate hydrolysis measured by fluore...More data for this Ligand-Target Pair
Affinity DataKi: 1.68E+4nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-7-amino-4-methylcoumarin assessed as substrate hydrolysis measured by fluore...More data for this Ligand-Target Pair
Affinity DataKi: 1.76E+4nMAssay Description:Inhibition of PGPH-like activity of 20S proteasome in human erythrocytes using Z-Leu-Leu-Glu-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.77E+4nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-7-amino-4-methylcoumarin assessed as substrate hydrolysis measured by fluore...More data for this Ligand-Target Pair
Affinity DataKi: 1.80E+4nM ΔG°: -28.2kJ/molepH: 7.4 T: 2°CAssay Description:The reaction mixture (final volume 200 µL) contained 5 µg/mL MAO-A or MAO-B in a 50 mM sodium phosphate buffer pH 7.4, 1 mM p-tyramine (sub...More data for this Ligand-Target Pair
TargetPteridine reductase 1(Leishmania major)
University Of Modena And Reggio Emilia
Curated by ChEMBL
University Of Modena And Reggio Emilia
Curated by ChEMBL
Affinity DataKi: 1.92E+4nMAssay Description:Inhibition of Leishmania major PTR1 using H2B as substrate preincubated for 10 mins followed by NADPH addition and measured up to 50 mins relative to...More data for this Ligand-Target Pair
Affinity DataKi: 2.20E+4nM ΔG°: -27.7kJ/molepH: 7.4 T: 2°CAssay Description:The reaction mixture (final volume 200 µL) contained 5 µg/mL MAO-A or MAO-B in a 50 mM sodium phosphate buffer pH 7.4, 1 mM p-tyramine (sub...More data for this Ligand-Target Pair
Affinity DataKi: 2.37E+4nMAssay Description:Inhibition of bovine pancreatic alpha-chymotrypsin using Suc-Leu-Leu-Val-Tyr-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair

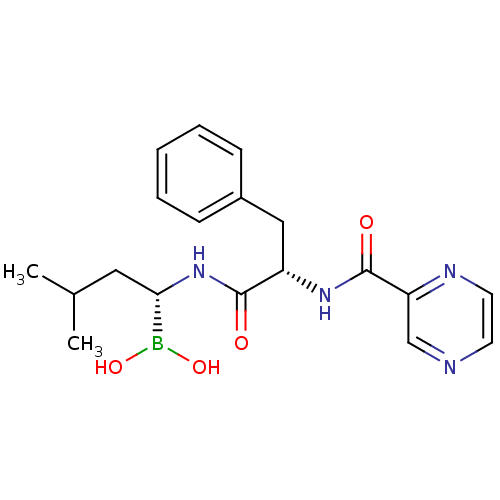
 3D Structure (crystal)
3D Structure (crystal)