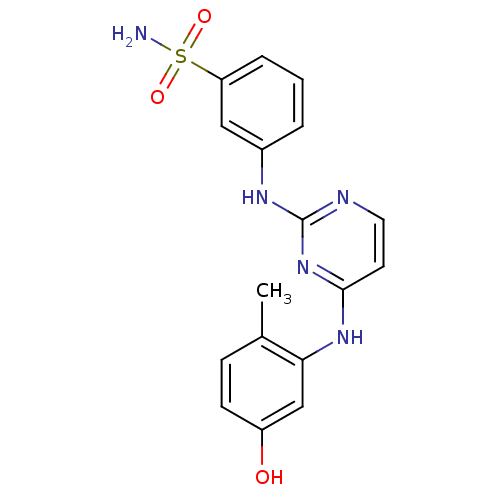
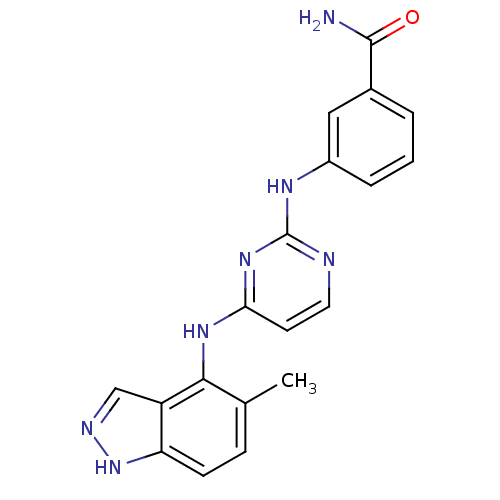
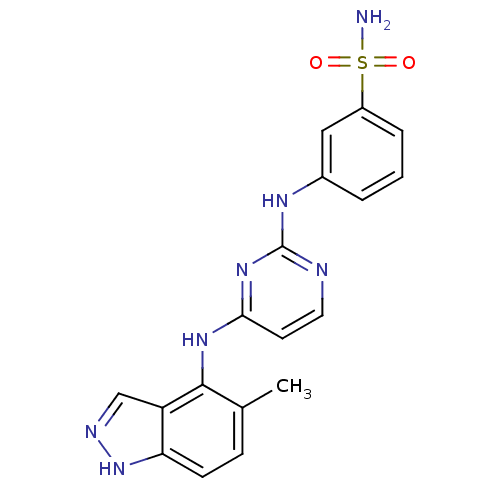
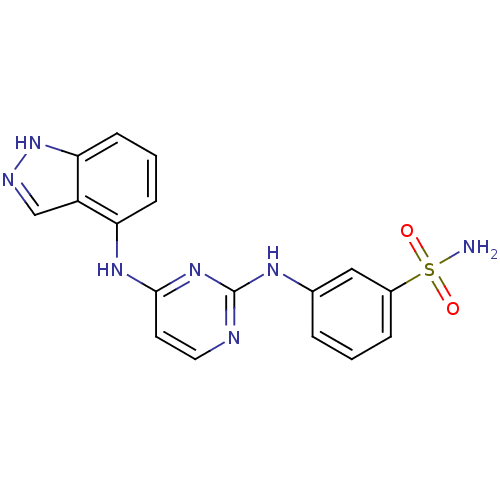
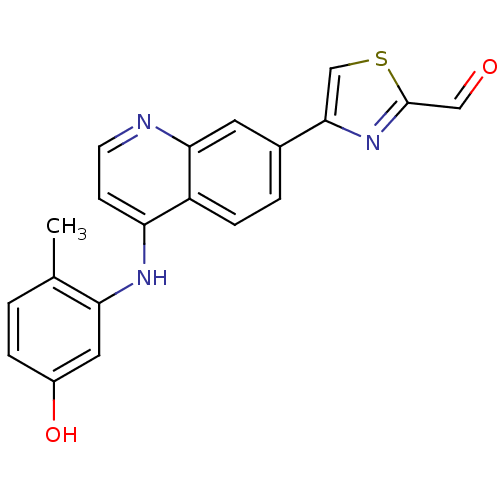
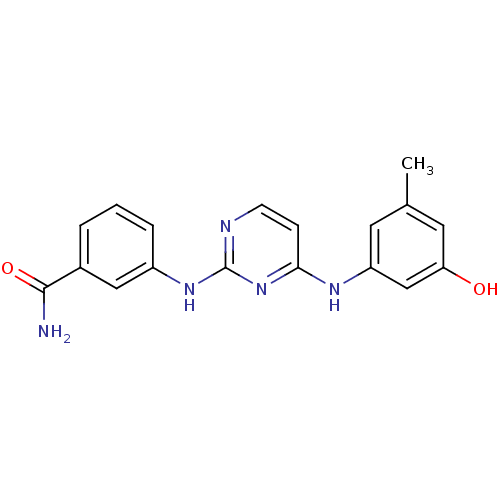
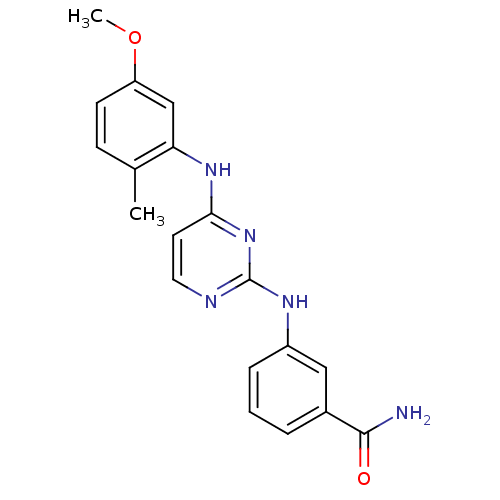
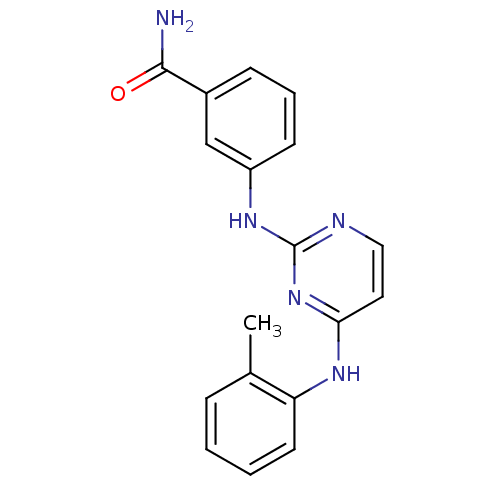
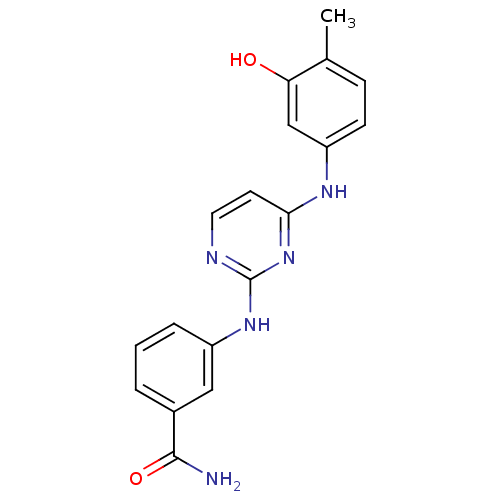
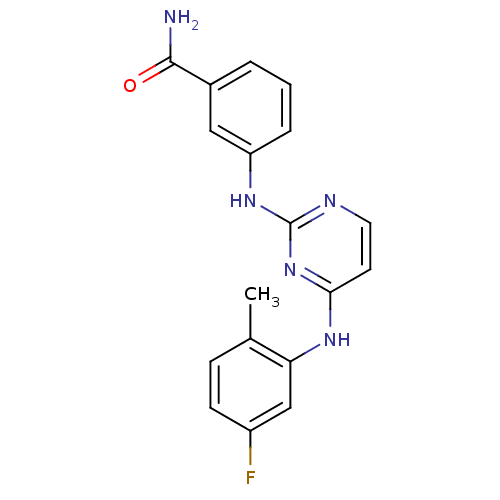
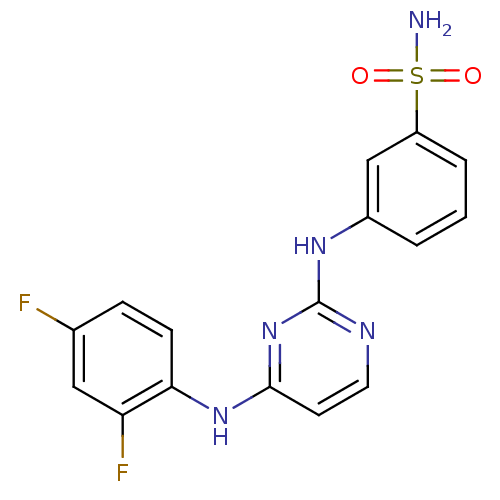
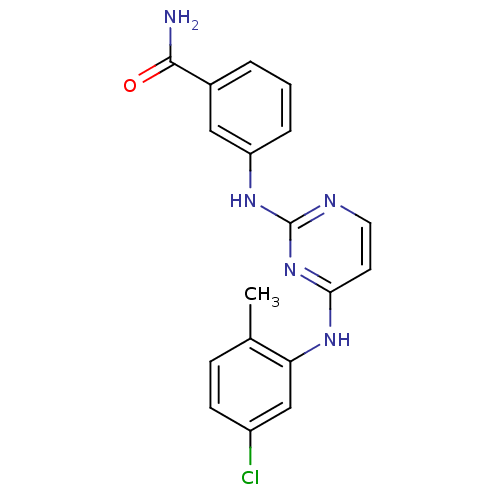
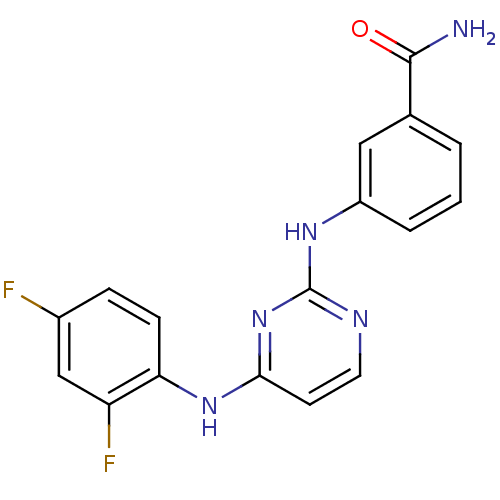
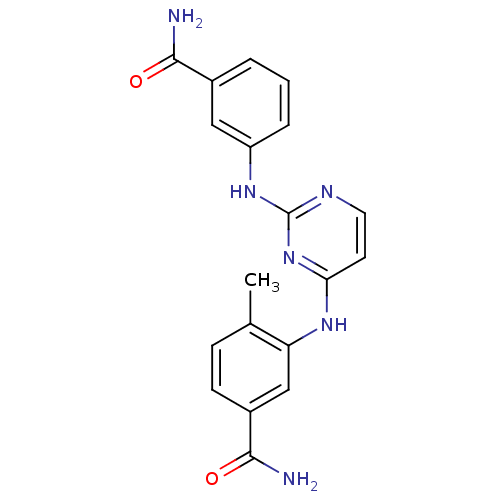
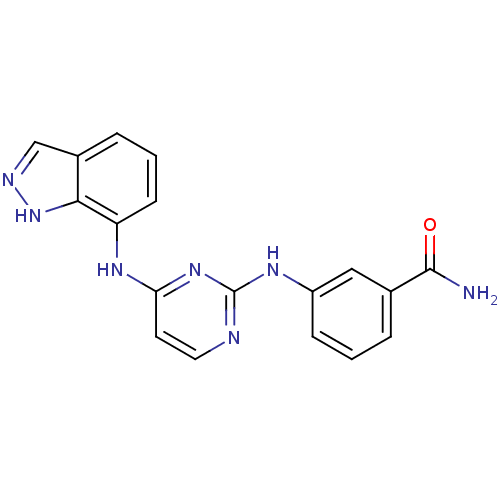
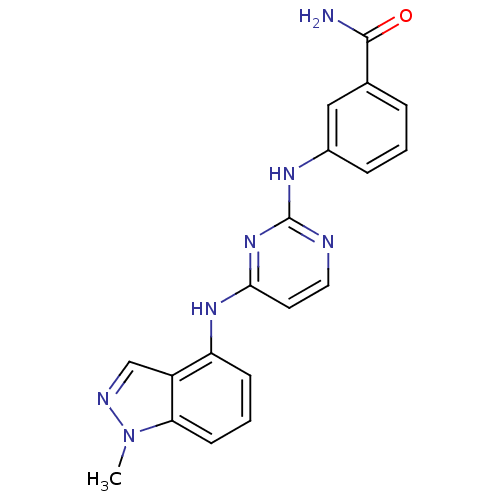
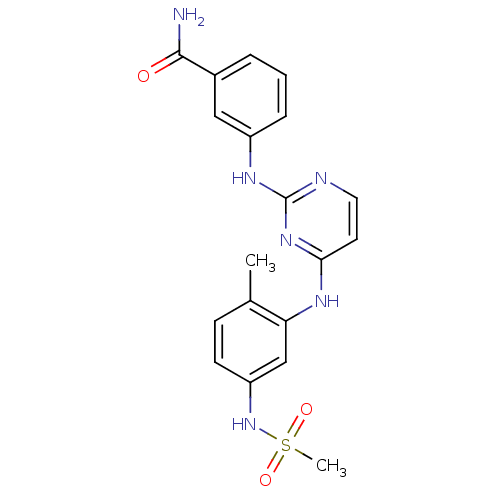
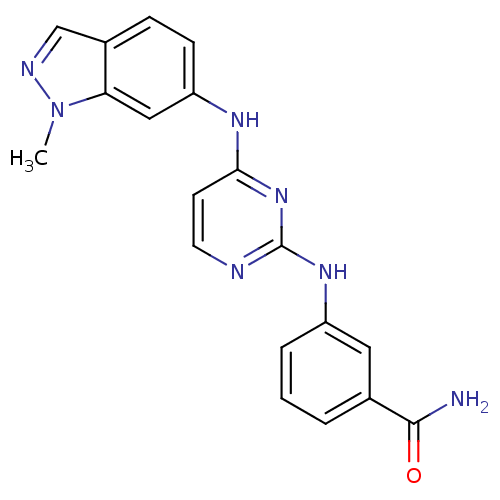
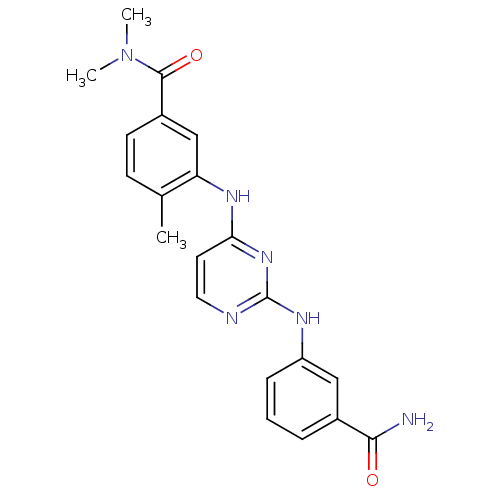
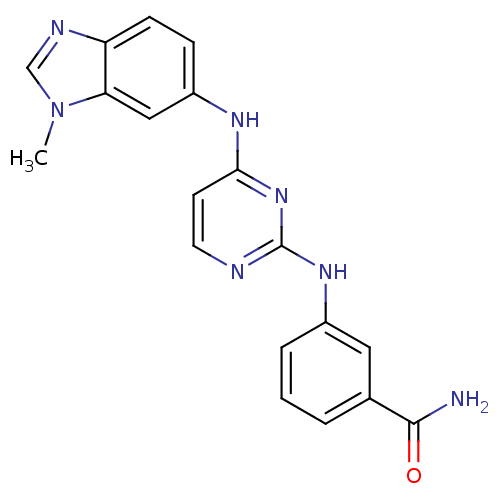
Report error Found 43 Enz. Inhib. hit(s) with all data for entry = 2928
Affinity DataIC50: 8.5nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 33nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 41nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 54nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 55nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 59nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 73nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 86nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 89nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 97nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 190nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 270nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 350nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 380nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 490nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 520nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 560nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 880nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+3nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+3nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+3nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 3.40E+3nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 4.40E+3nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 5.60E+3nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 7.70E+3nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 8.20E+3nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 9.30E+3nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+4nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair