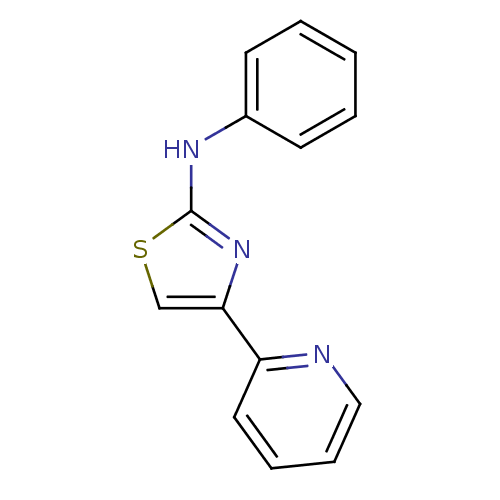
Report error Found 26 Enz. Inhib. hit(s) with all data for entry = 50038682
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 0.0640nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by electrophysiology assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 0.168nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Displacement of [125I]apamin from Kca2.3 channel expressed in HEK293 cells by scintillation proximity assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 1(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of Kca2.1 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 2(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of Kca2.2 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 25nMAssay Description:Displacement of [125I]apamin from Kca2.3 channel expressed in HEK293 cells by scintillation proximity assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 1(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 43nMAssay Description:Inhibition of Kca2.1 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 56nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by electrophysiology assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 2(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 59nMAssay Description:Displacement of [125I]apamin from Kca2.2 channel expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 59nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 2(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 123nMAssay Description:Inhibition of Kca2.2 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 290nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by electrophysiology assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 543nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 2(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 543nMAssay Description:Displacement of [125I]apamin from Kca2.2 channel expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 1.30E+3nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by electrophysiology assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair