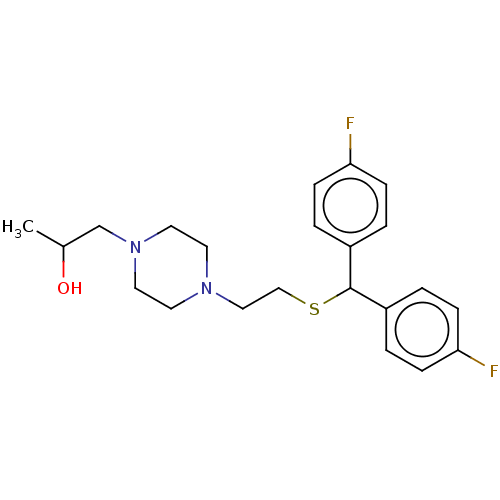
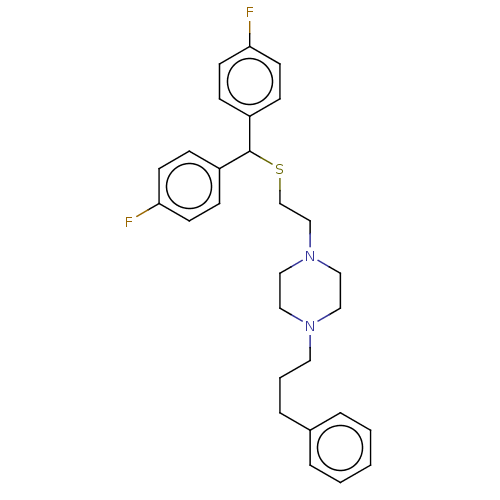
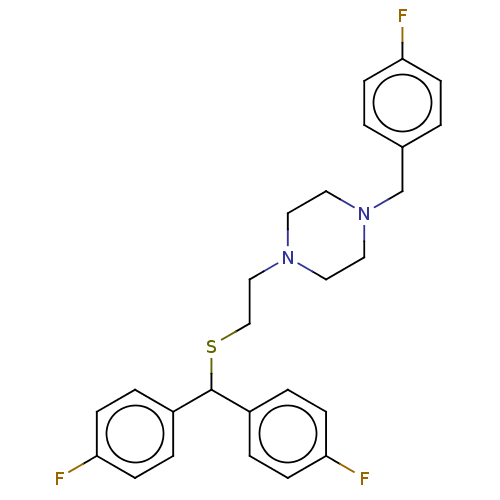
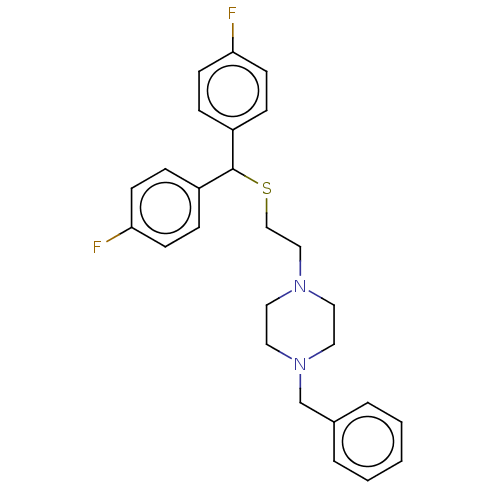
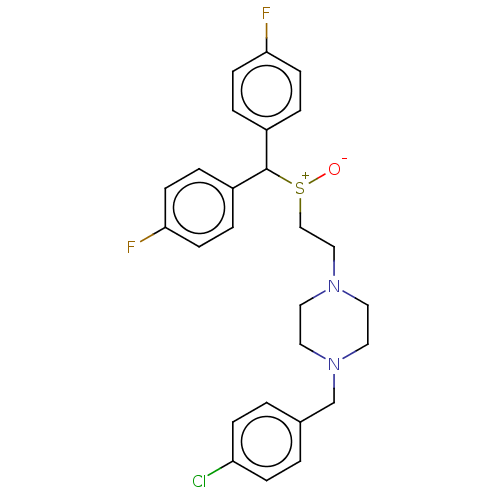
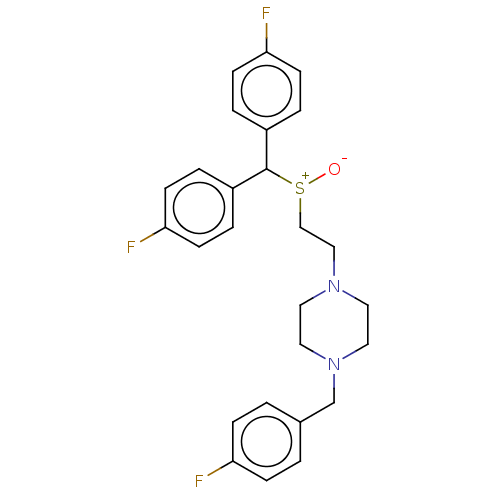
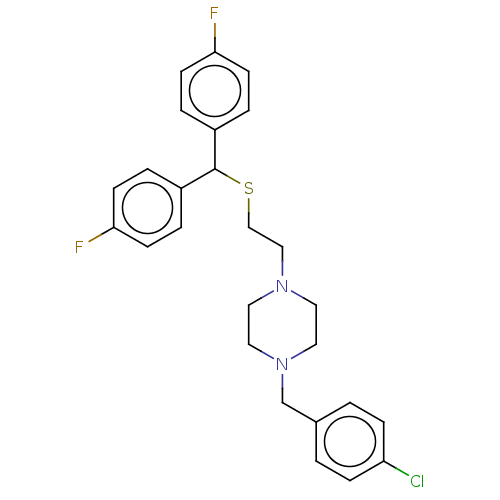
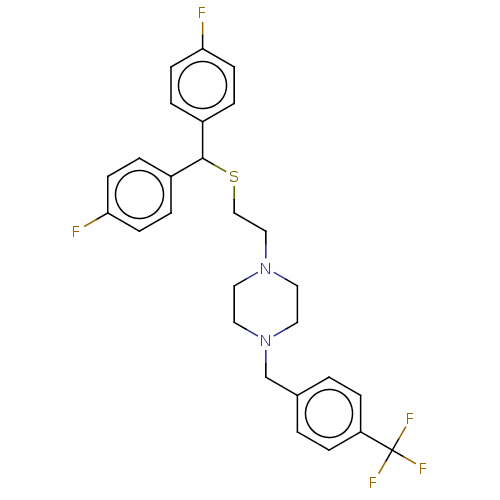
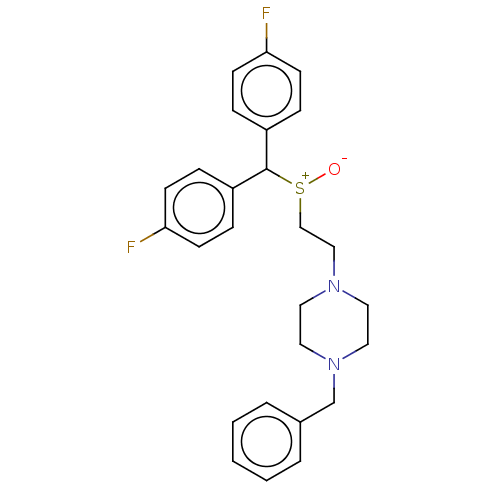
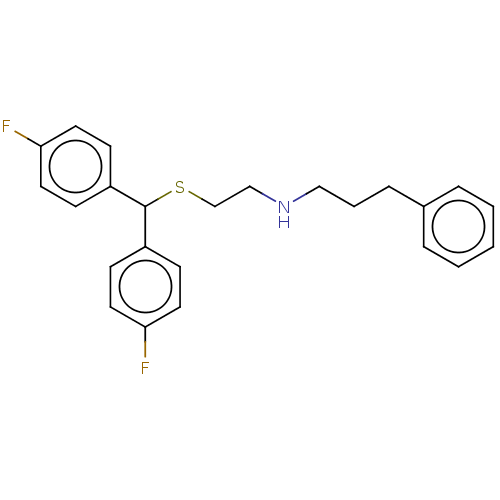
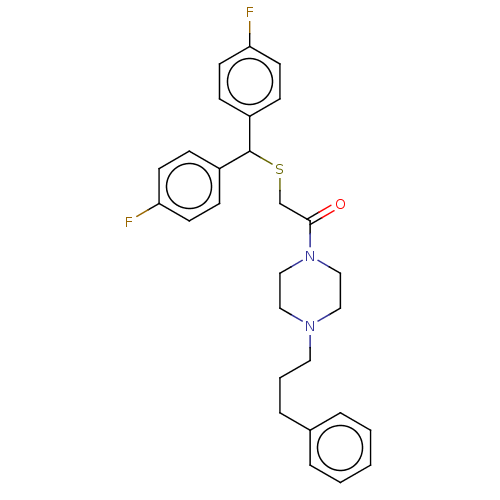
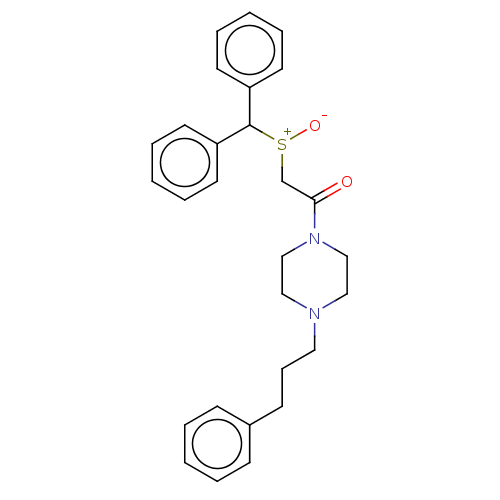
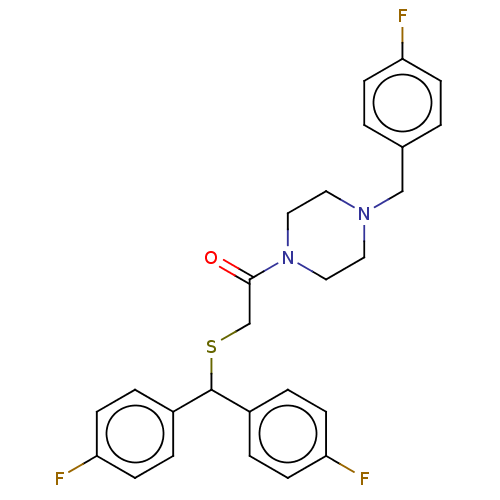
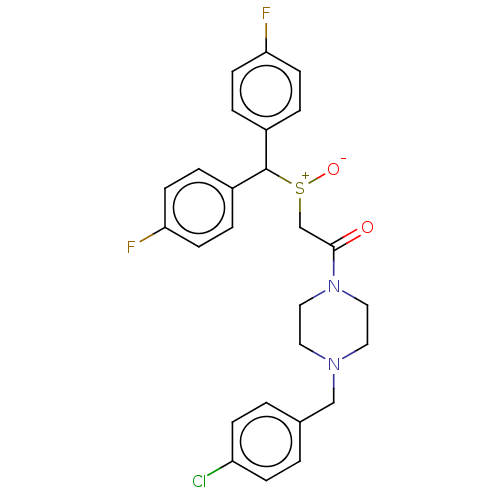
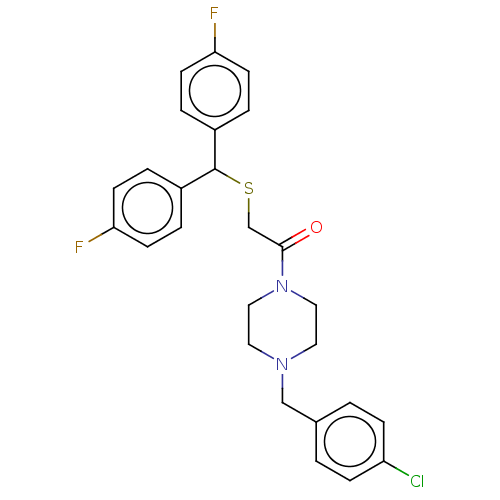
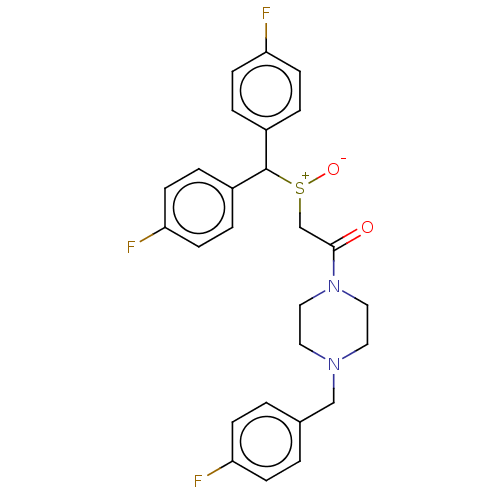
Report error Found 122 Enz. Inhib. hit(s) with all data for entry = 50047981
Affinity DataKi: 0.110nMAssay Description:Displacement of [3H]N-methylspiperone from human D3R expressed in HEK293 cell membranes incubated for 1 hr by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.260nMAssay Description:Displacement of [3H]N-methylspiperone from human D2R expressed in HEK293 cell membranes incubated for 1 hr by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.10nMAssay Description:Antagonist activity at DRD2 (unknown origin) expressed in HEK293T cells transfected with Galphai1-RLuc8 and gamma2-GFP10 assessed as inhibition of qu...More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Guinea pig)
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
Affinity DataKi: 2.40nMAssay Description:Displacement of [3H](+)-pentazocine from sigma1 receptor in whole-guinea pig brain minus cerebellum homogenates incubated for 120 mins by radioligand...More data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.90nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.20nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.40nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.60nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Guinea pig)
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
Affinity DataKi: 4nMAssay Description:Displacement of [3H](+)-pentazocine from sigma1 receptor in whole-guinea pig brain minus cerebellum homogenates incubated for 120 mins by radioligand...More data for this Ligand-Target Pair
Affinity DataKi: 4.5nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 6.5nMAssay Description:Displacement of [3H]WIN35,428 from wild type human DAT expressed in COS7 cell membranes incubated for >90 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 6.70nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 7.40nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 7.60nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 9.70nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Inhibition of DAT (unknown origin) expressed in COS7 cells assessed as reduction in [3H]DA uptake incubated for 5 mins by beta-scintillation counting...More data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 27nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 27nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 28nMAssay Description:Displacement of [3H]N-methylspiperone from human D4R expressed in HEK293 cell membranes incubated for 1 hr by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 29nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 33nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 34nMAssay Description:Displacement of [3H]N-methylspiperone from human D3R expressed in HEK293 cell membranes incubated for 1 hr by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 38nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 39nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 47nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 47nMAssay Description:Displacement of [3H]WIN35,428 from wild type human DAT expressed in COS7 cell membranes incubated for >90 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 50nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 54nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 56nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 59nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 66nMAssay Description:Displacement of [3H]N-methylspiperone from human D3R expressed in HEK293 cell membranes incubated for 1 hr by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 68nMAssay Description:Displacement of [3H]N-methylspiperone from human D2R expressed in HEK293 cell membranes incubated for 1 hr by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 70nMAssay Description:Displacement of [3H]N-methylspiperone from human D4R expressed in HEK293 cell membranes incubated for 1 hr by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 74nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 79nMAssay Description:Displacement of [3H]N-methylspiperone from human D2R expressed in HEK293 cell membranes incubated for 1 hr by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 101nMAssay Description:Displacement of [3H]WIN35,428 from wild type human DAT expressed in COS7 cell membranes incubated for >90 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 116nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 143nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Guinea pig)
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
Affinity DataKi: 159nMAssay Description:Displacement of [3H](+)-pentazocine from sigma1 receptor in whole-guinea pig brain minus cerebellum homogenates incubated for 120 mins by radioligand...More data for this Ligand-Target Pair
Affinity DataKi: 200nMAssay Description:Inhibition of DAT (unknown origin) expressed in COS7 cells assessed as reduction in [3H]DA uptake incubated for 5 mins by beta-scintillation counting...More data for this Ligand-Target Pair
Affinity DataKi: 200nMAssay Description:Inhibition of DAT (unknown origin) expressed in COS7 cells assessed as reduction in [3H]DA uptake incubated for 5 mins by beta-scintillation counting...More data for this Ligand-Target Pair
Affinity DataKi: 213nMAssay Description:Displacement of [3H]citalopram from SERT in Sprague-Dawley rat brain stem membranes incubated for 60 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 214nMAssay Description:Displacement of [3H]WIN35,428 from DAT in Sprague-Dawley rat brain membranes incubated for 120 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 223nMAssay Description:Displacement of [3H]WIN35,428 from wild type human DAT expressed in COS7 cell membranes incubated for >90 mins by radioligand binding assayMore data for this Ligand-Target Pair