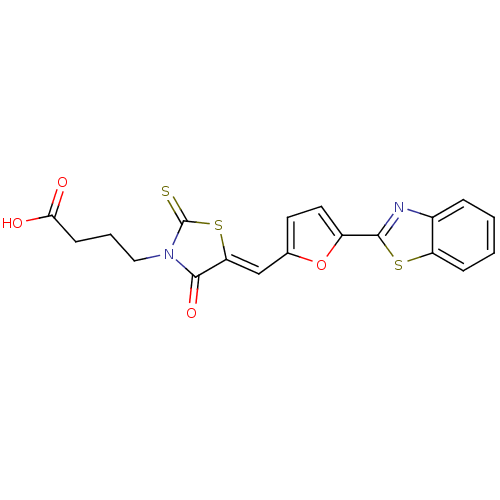
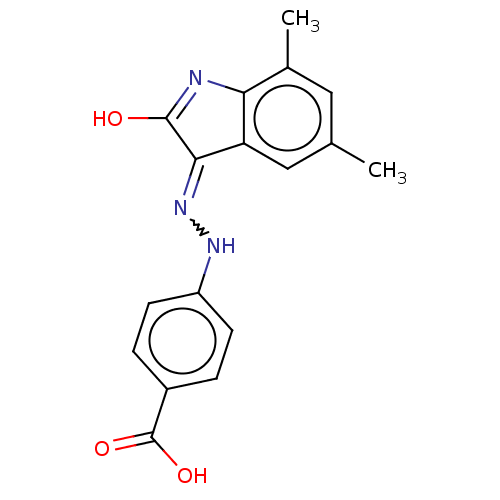
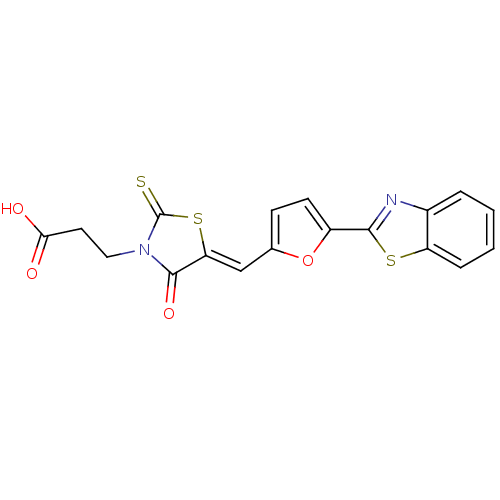
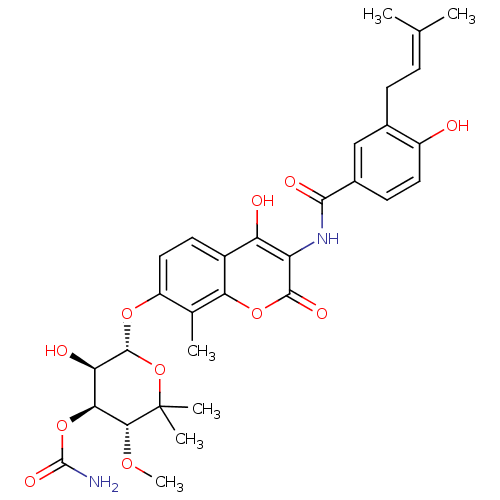
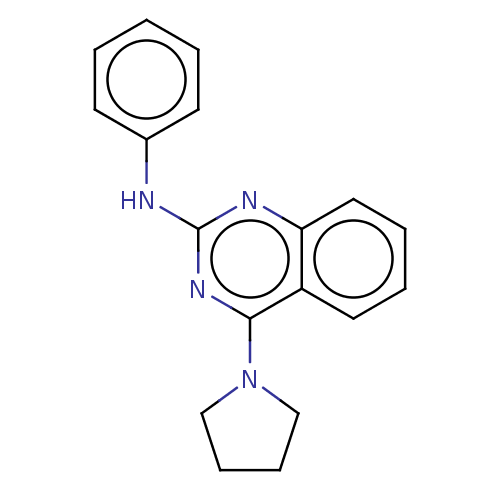
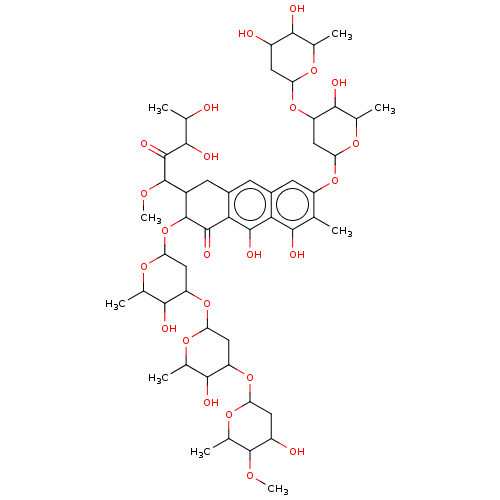
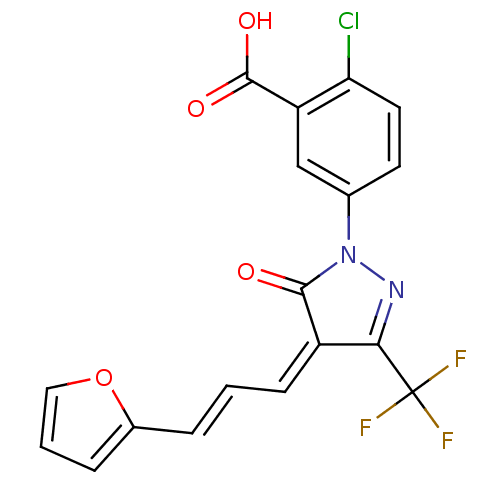
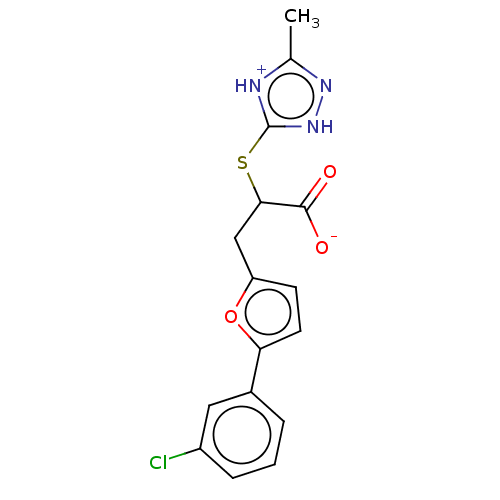
Report error Found 103 Enz. Inhib. hit(s) with all data for entry = 11635
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
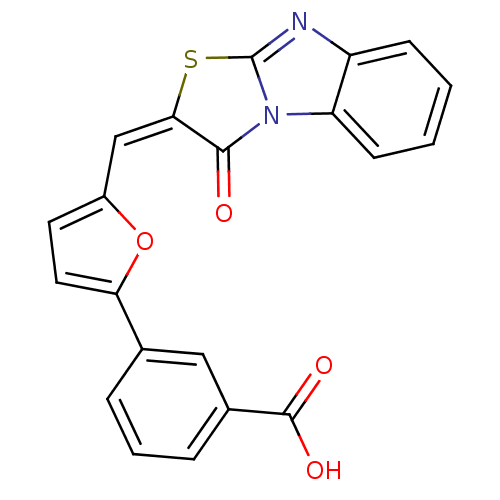
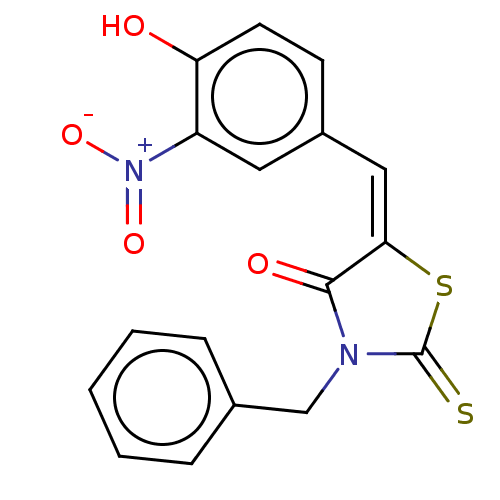
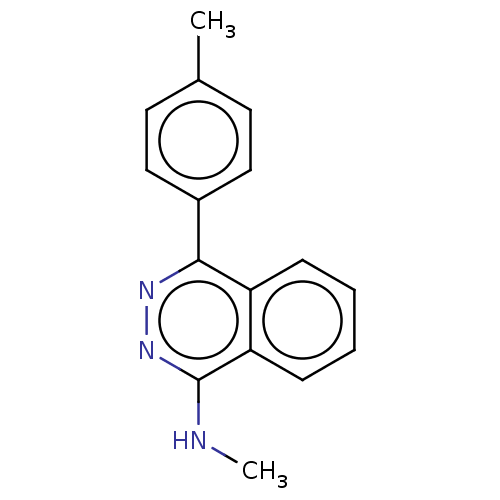
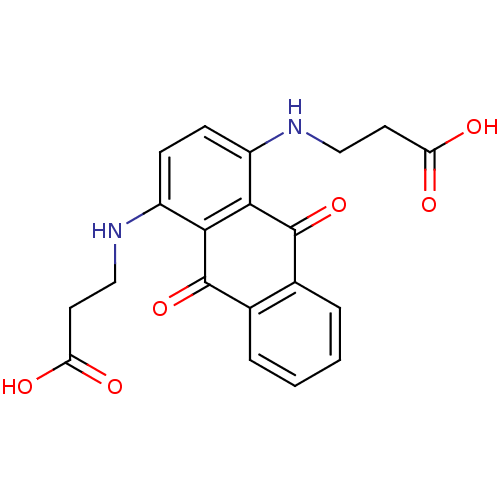
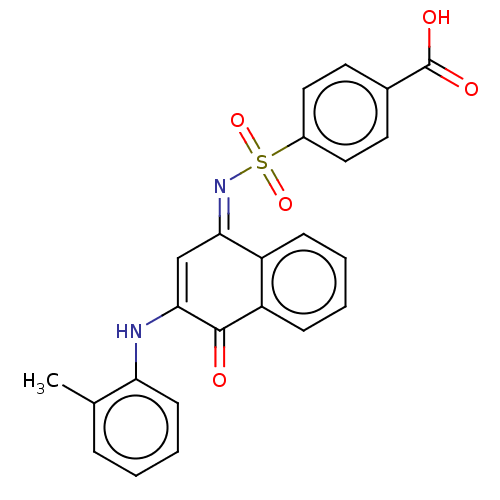
Affinity DataIC50: 1.48E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
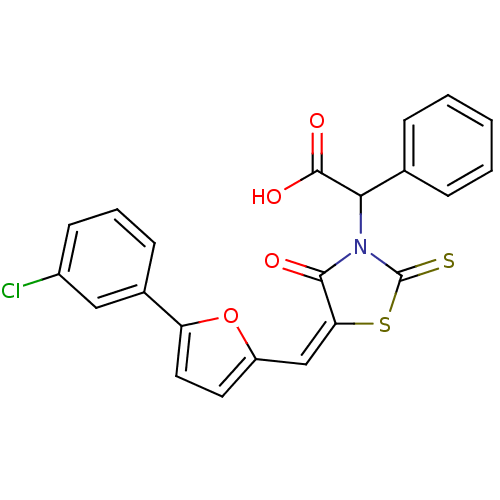
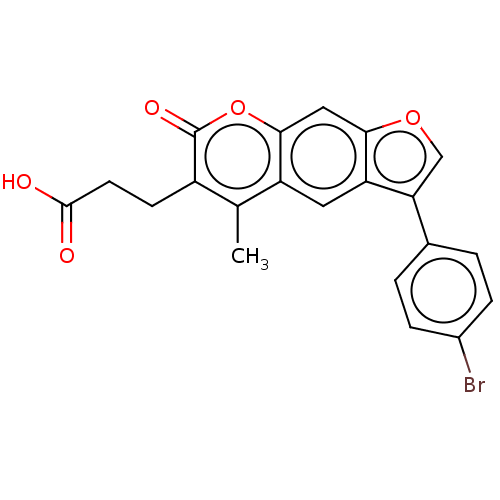
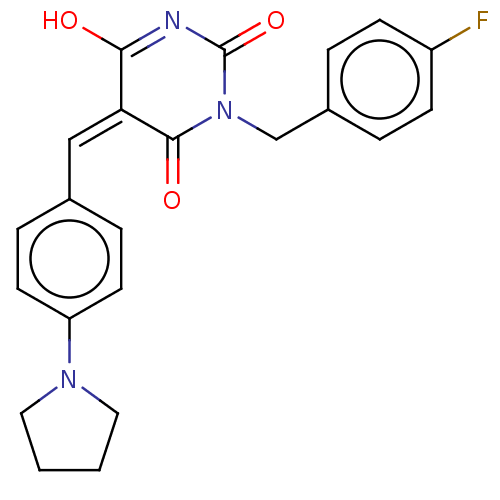
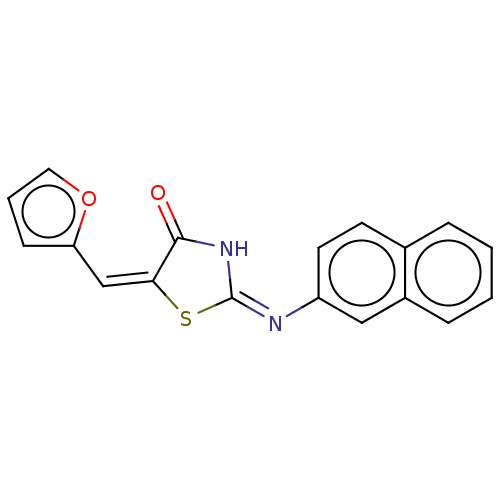
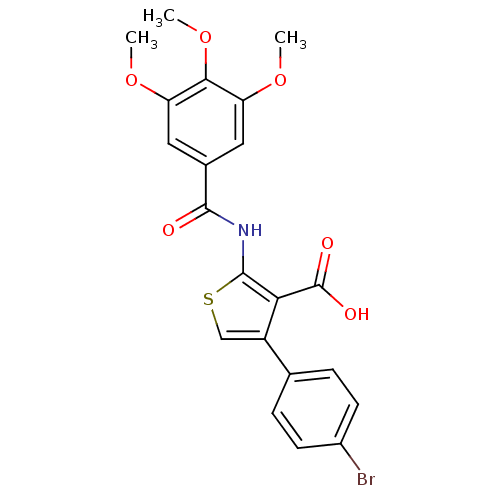
Affinity DataIC50: 1.56E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
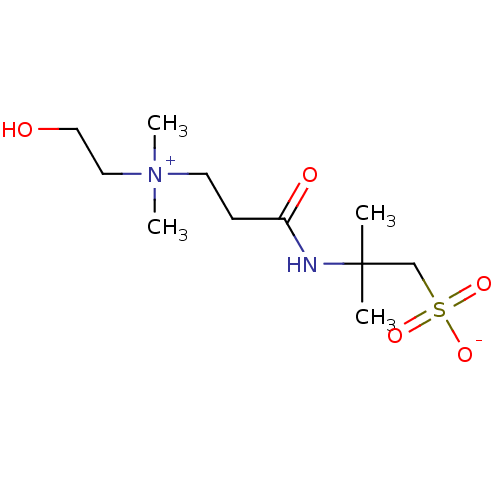
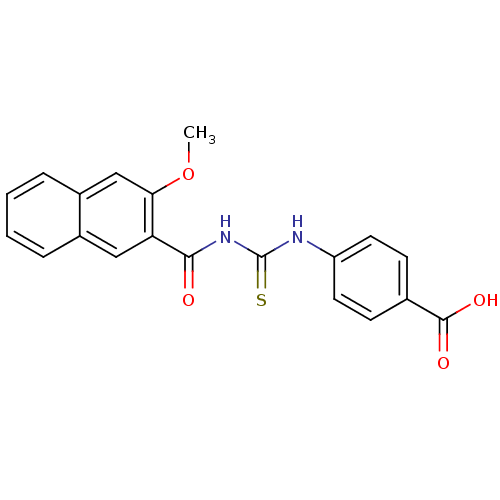
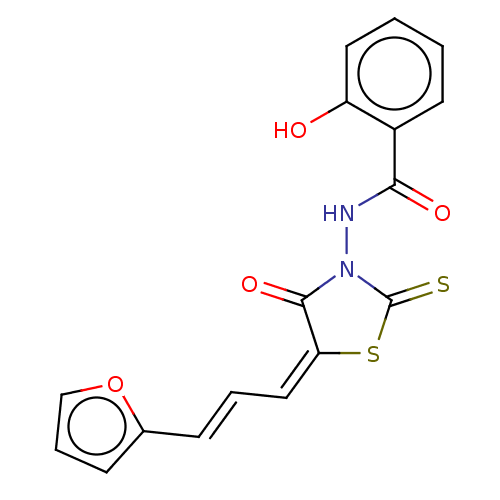
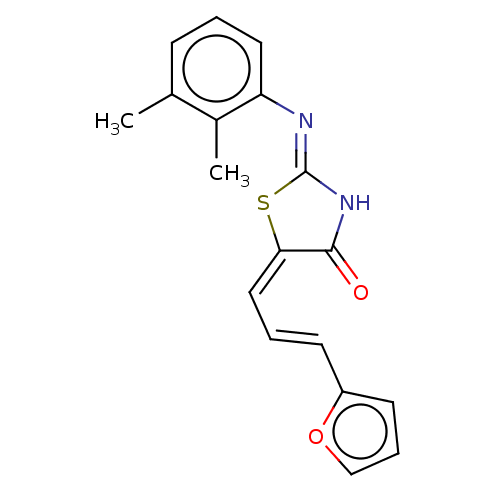
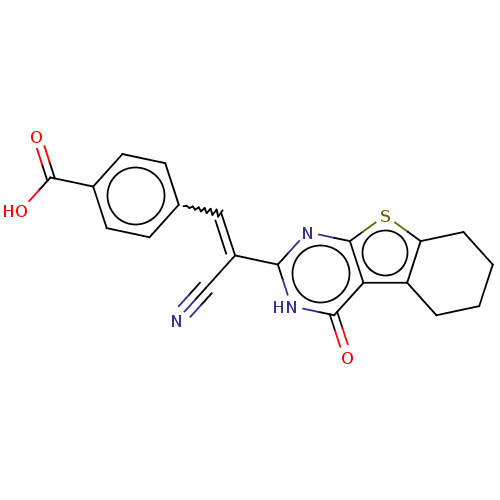
Affinity DataIC50: 1.69E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
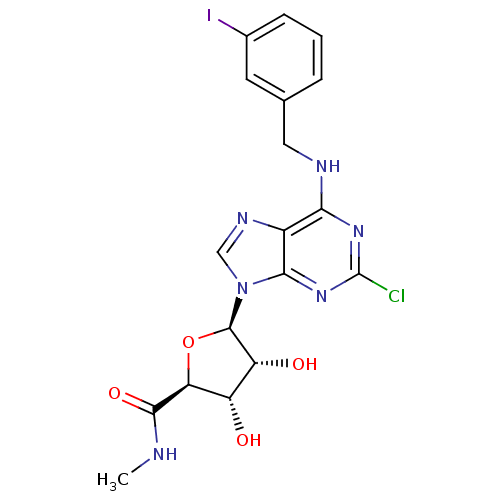
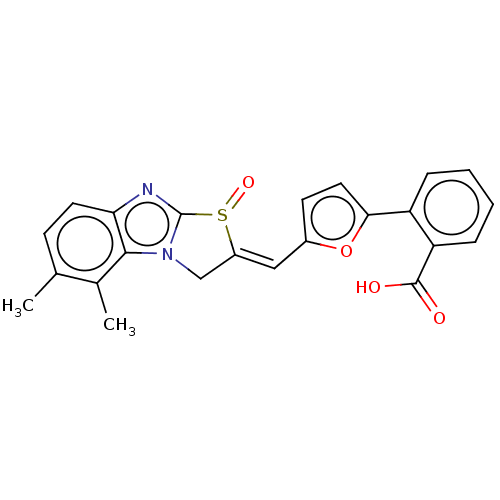
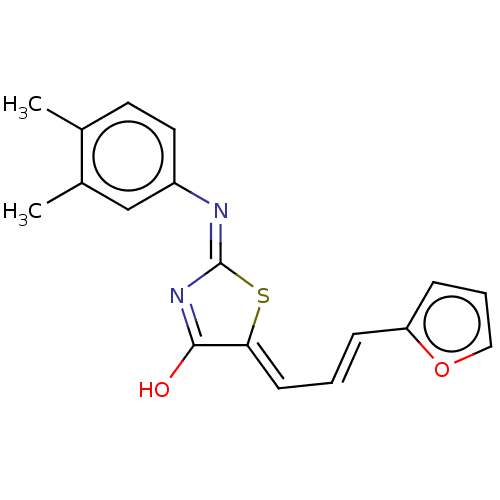
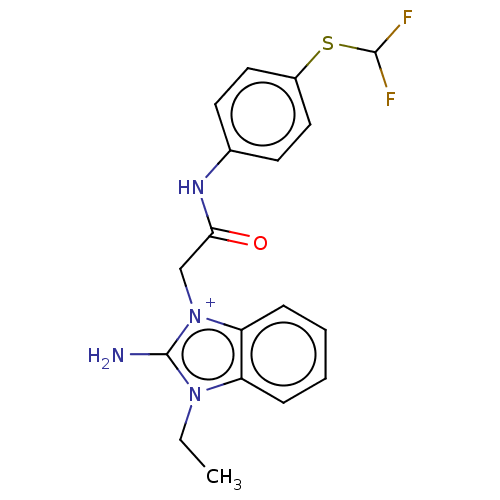
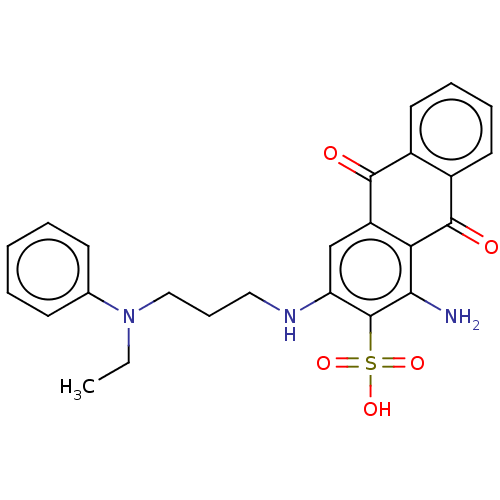
Affinity DataIC50: 1.74E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 1.85E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 1.87E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 2.00E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 2.18E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 2.53E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.13E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.13E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.55E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 4.28E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 5.28E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 6.25E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 6.25E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 7.42E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 7.86E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 8.00E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 1.00E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 1.06E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 1.12E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 1.24E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 2.00E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 2.00E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 2.00E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 2.00E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 2.00E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 2.00E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 2.00E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 2.00E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 2.00E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 2.00E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 2.00E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.75E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.75E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.75E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.75E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.75E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.75E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.75E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.75E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.75E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.75E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.75E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.75E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.75E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.75E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.75E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
TargetDNA gyrase inhibitor(Escherichia coli (strain K12))
The Florida International University Board of Trustees
US Patent
The Florida International University Board of Trustees
US Patent
Affinity DataIC50: 3.75E+4nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 µL of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair