Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Glutamate receptor ionotropic, kainate 1
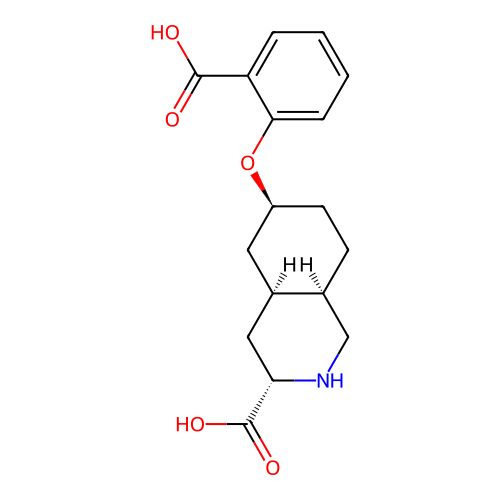
Ligand
BDBM50494060
Substrate
n/a
Meas. Tech.
ChEMBL_990076 (CHEMBL2443753)
Ki
2300±n/a nM
Citation
 Martinez-Perez, JA; Iyengar, S; Shannon, HE; Bleakman, D; Alt, A; Arnold, BM; Bell, MG; Bleisch, TJ; Casta�o, AM; Del Prado, M; Dominguez, E; Escribano, AM; Filla, SA; Ho, KH; Hudziak, KJ; Jones, CK; Mateo, A; Mathes, BM; Mattiuz, EL; Ogden, AM; Simmons, RM; Stack, DR; Stratford, RE; Winter, MA; Wu, Z; Ornstein, PL GluK1 antagonists from 6-(carboxy)phenyl decahydroisoquinoline derivatives. SAR and evaluation of a prodrug strategy for oral efficacy in pain models. Bioorg Med Chem Lett 23:6459-62 (2013) [PubMed] Article
Martinez-Perez, JA; Iyengar, S; Shannon, HE; Bleakman, D; Alt, A; Arnold, BM; Bell, MG; Bleisch, TJ; Casta�o, AM; Del Prado, M; Dominguez, E; Escribano, AM; Filla, SA; Ho, KH; Hudziak, KJ; Jones, CK; Mateo, A; Mathes, BM; Mattiuz, EL; Ogden, AM; Simmons, RM; Stack, DR; Stratford, RE; Winter, MA; Wu, Z; Ornstein, PL GluK1 antagonists from 6-(carboxy)phenyl decahydroisoquinoline derivatives. SAR and evaluation of a prodrug strategy for oral efficacy in pain models. Bioorg Med Chem Lett 23:6459-62 (2013) [PubMed] Article More Info.:
Target
Name:
Glutamate receptor ionotropic, kainate 1
Synonyms:
GLUR5 | GRIK1 | GRIK1_HUMAN | Glutamate Receptor | Glutamate kainate | Glutamate receptor ionotropic kainate | Glutamate receptor ionotropic kainate 1 | Glutamate receptor, ionotropic kainate 1 | Glutamate-Kainate | Glutamate-Kainate, GluR5 | Grik1 protein | hmglur5 flipr
Type:
Enzyme Catalytic Domain
Mol. Mass.:
103984.96
Organism:
Human
Description:
P39086
Residue:
918
Sequence:
MEHGTLLAQPGLWTRDTSWALLYFLCYILPQTAPQVLRIGGIFETVENEPVNVEELAFKFAVTSINRNRTLMPNTTLTYDIQRINLFDSFEASRRACDQLALGVAALFGPSHSSSVSAVQSICNALEVPHIQTRWKHPSVDNKDLFYINLYPDYAAISRAILDLVLYYNWKTVTVVYEDSTGLIRLQELIKAPSRYNIKIKIRQLPSGNKDAKPLLKEMKKGKEFYVIFDCSHETAAEILKQILFMGMMTEYYHYFFTTLDLFALDLELYRYSGVNMTGFRLLNIDNPHVSSIIEKWSMERLQAPPRPETGLLDGMMTTEAALMYDAVYMVAIASHRASQLTVSSLQCHRHKPWRLGPRFMNLIKEARWDGLTGHITFNKTNGLRKDFDLDIISLKEEGTEKAAGEVSKHLYKVWKKIGIWNSNSGLNMTDSNKDKSSNITDSLANRTLIVTTILEEPYVMYRKSDKPLYGNDRFEGYCLDLLKELSNILGFIYDVKLVPDGKYGAQNDKGEWNGMVKELIDHRADLAVAPLTITYVREKVIDFSKPFMTLGISILYRKPNGTNPGVFSFLNPLSPDIWMYVLLACLGVSCVLFVIARFTPYEWYNPHPCNPDSDVVENNFTLLNSFWFGVGALMQQGSELMPKALSTRIVGGIWWFFTLIIISSYTANLAAFLTVERMESPIDSADDLAKQTKIEYGAVRDGSTMTFFKKSKISTYEKMWAFMSSRQQTALVRNSDEGIQRVLTTDYALLMESTSIEYVTQRNCNLTQIGGLIDSKGYGVGTPIGSPYRDKITIAILQLQEEGKLHMMKEKWWRGNGCPEEDNKEASALGVENIGGIFIVLAAGLVLSVFVAIGEFIYKSRKNNDIEQAFCFFYGLQCKQTHPTNSTSGTTLSTDLECGKLIREERGIRKQSSVHTV