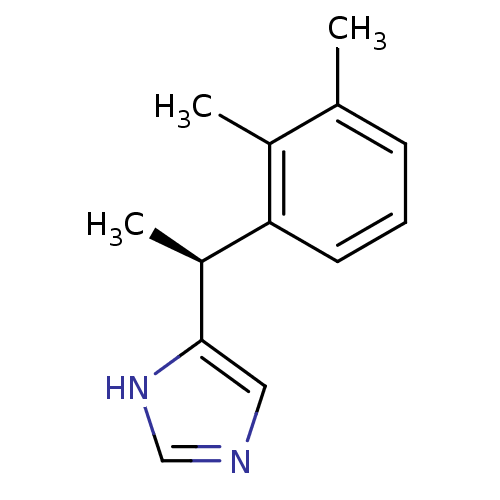
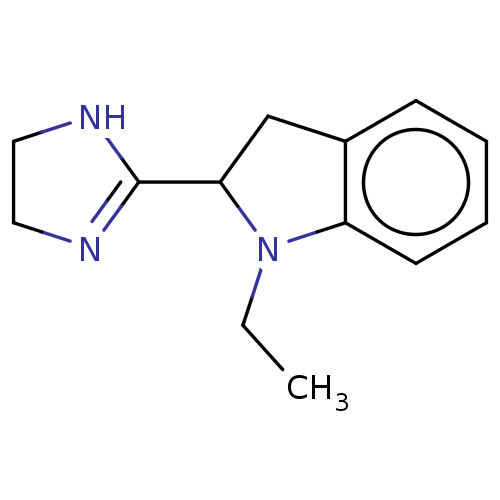
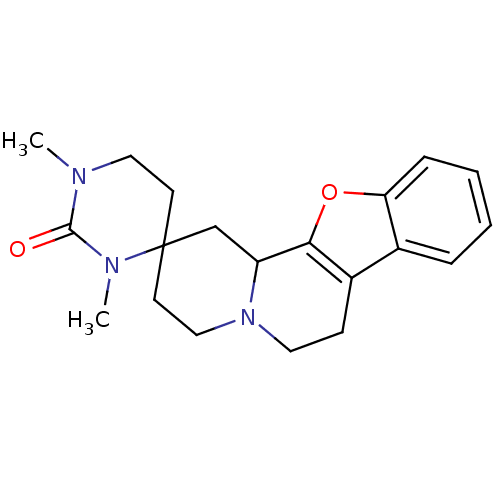
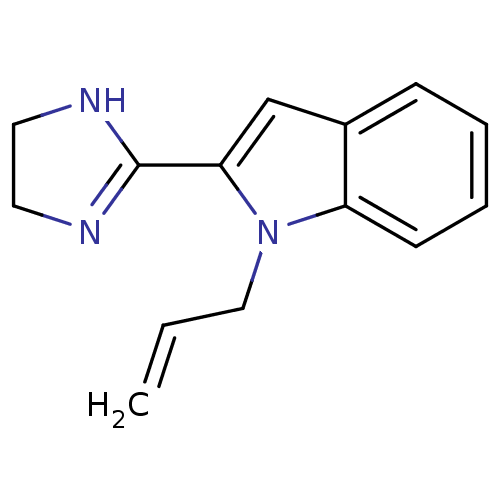
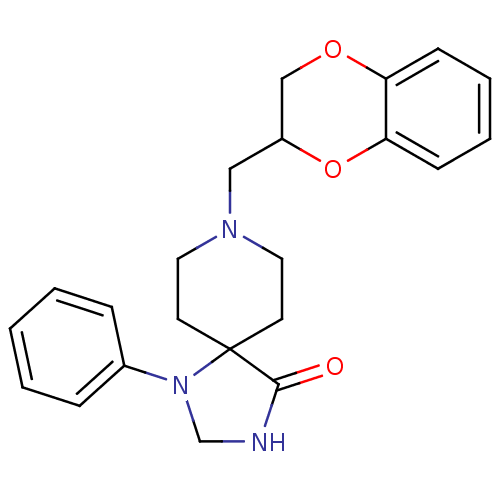
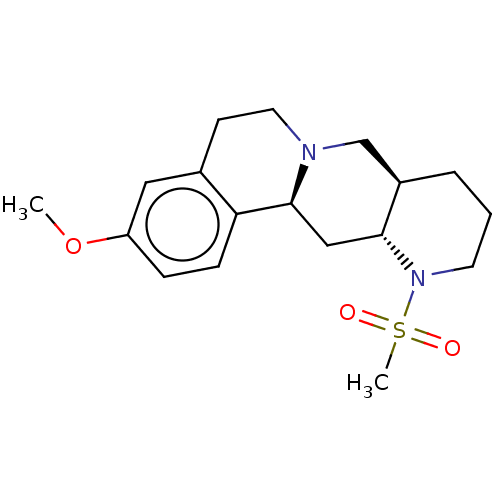
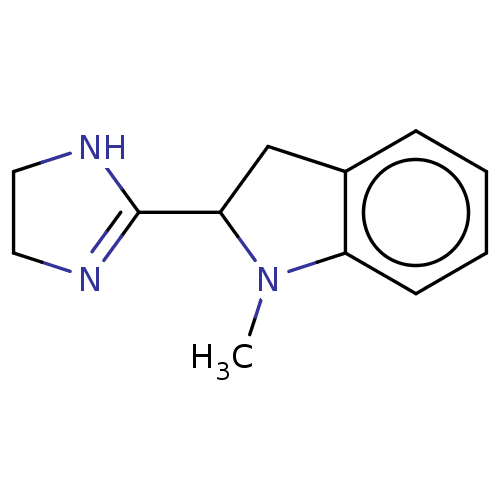
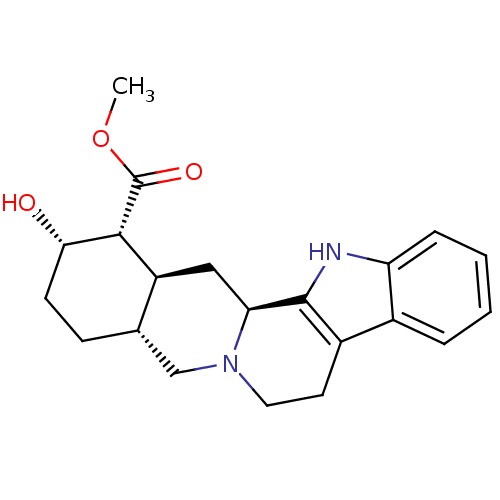
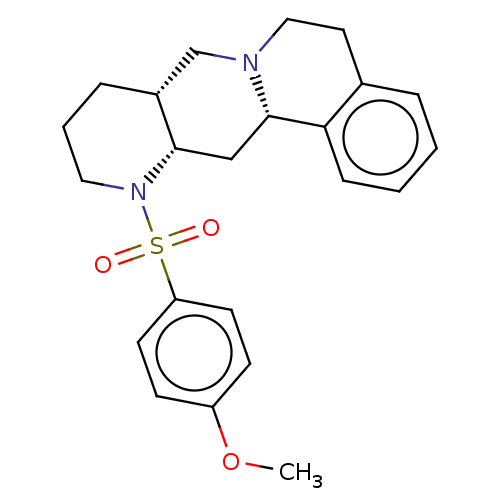
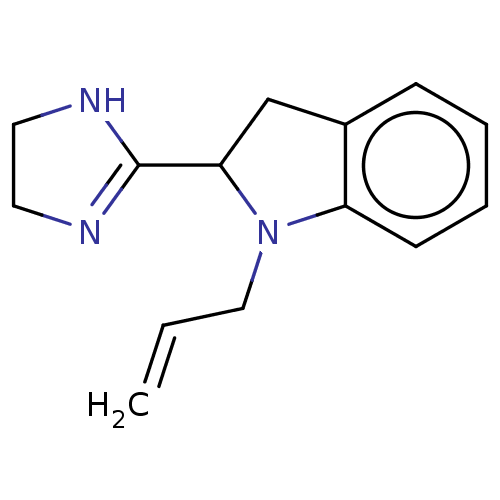
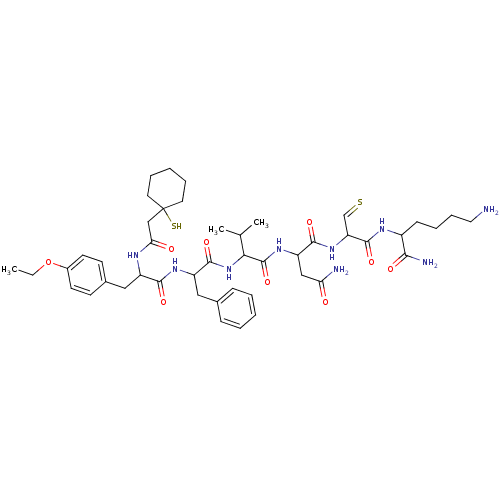
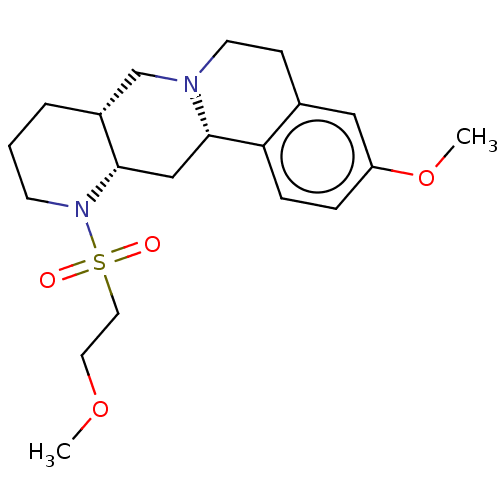
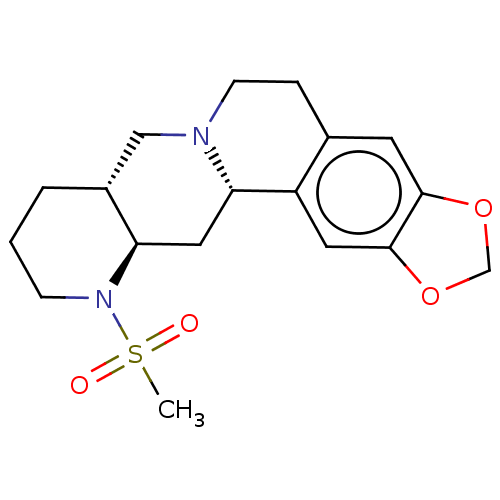
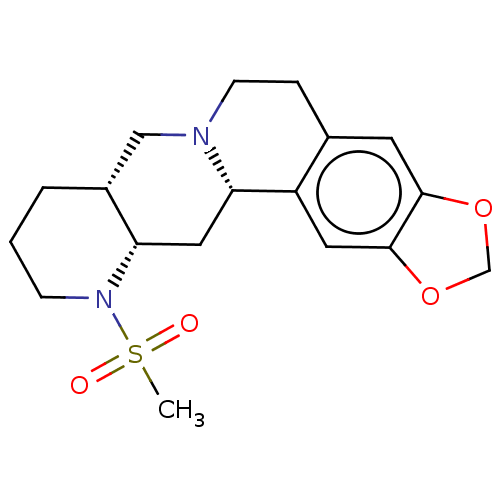
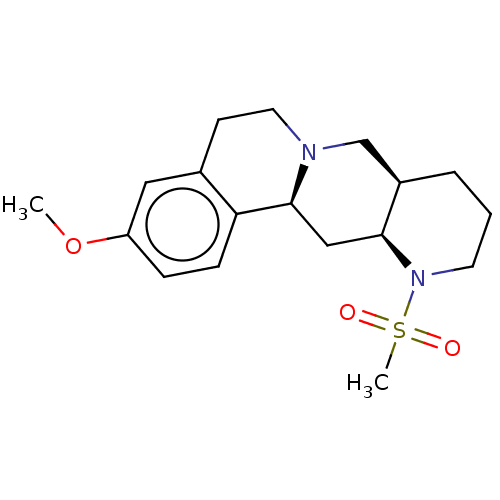
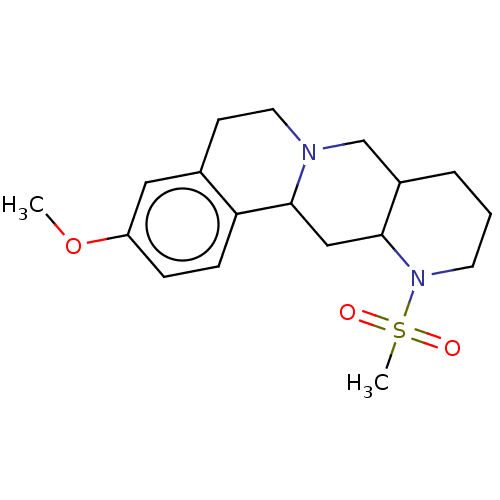
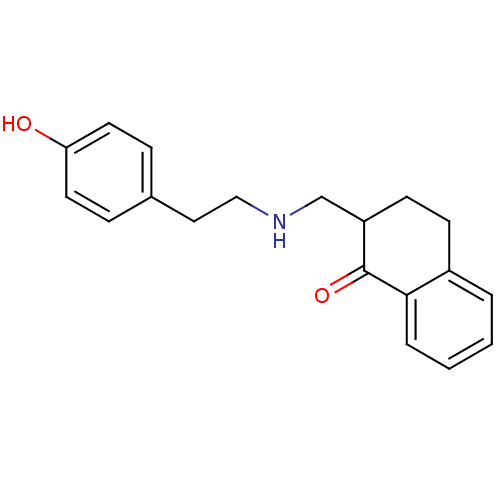
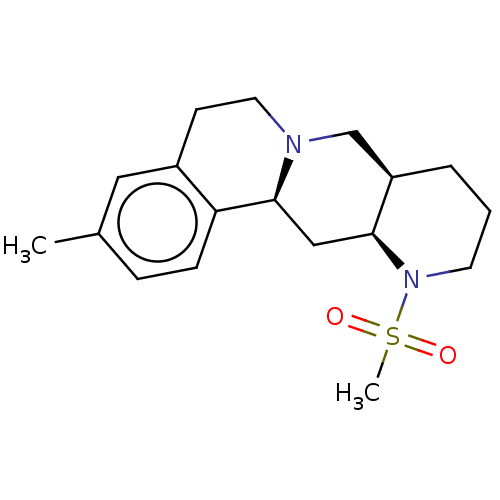
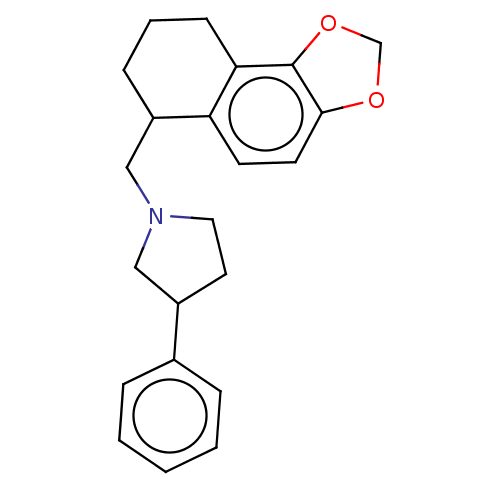
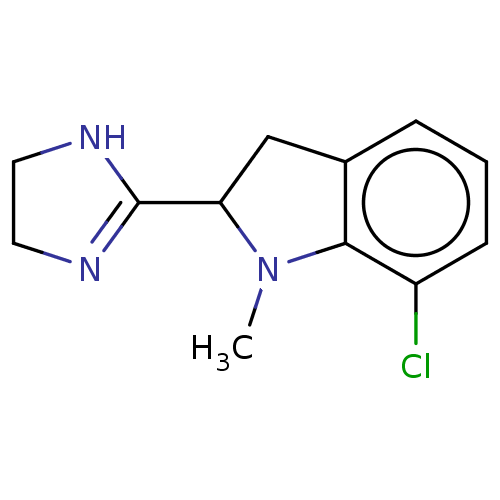
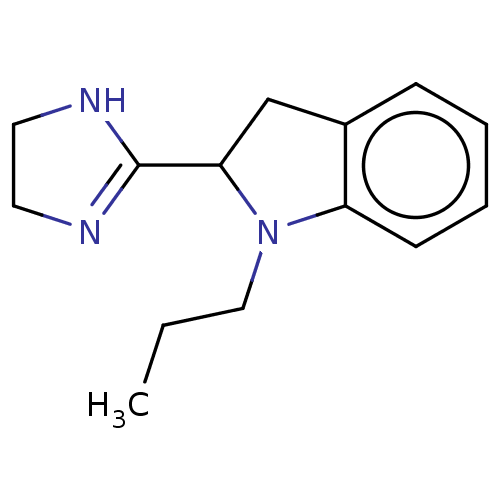
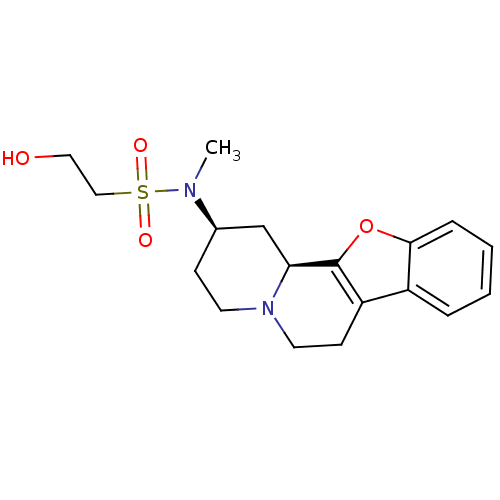
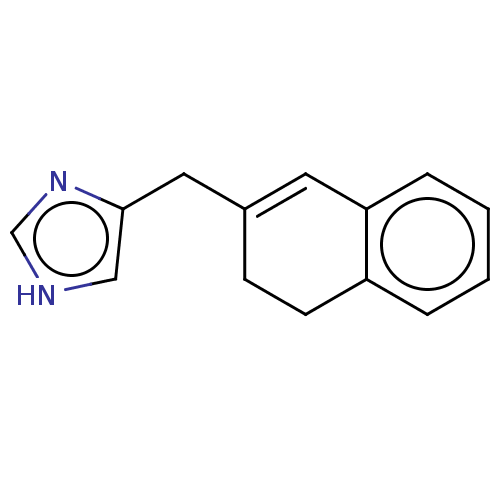
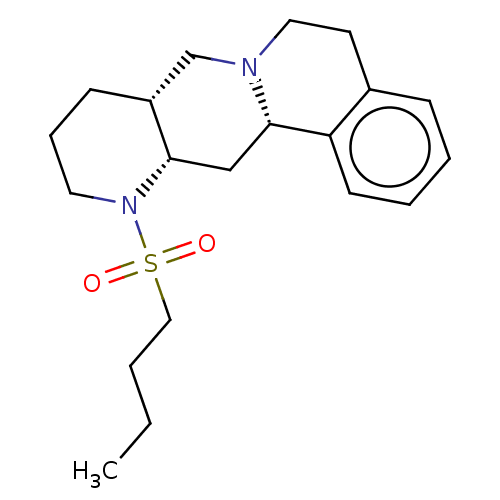
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.0150nMAssay Description:In vitro binding affinity against alpha-2 adrenergic receptor in ratMore data for this Ligand-Target Pair
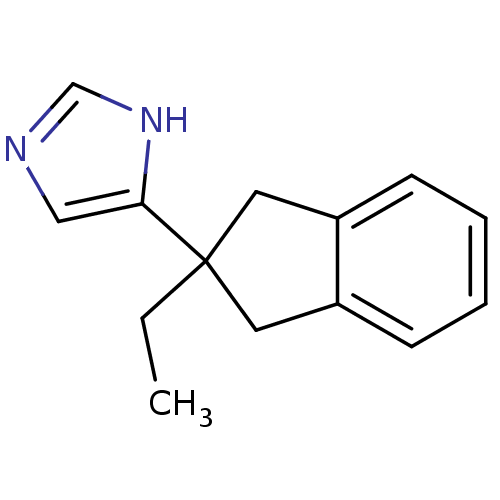
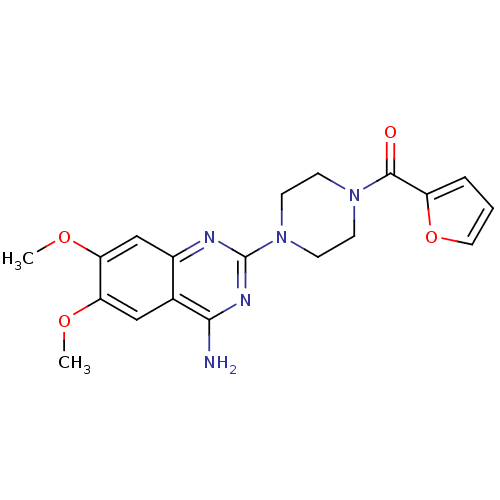
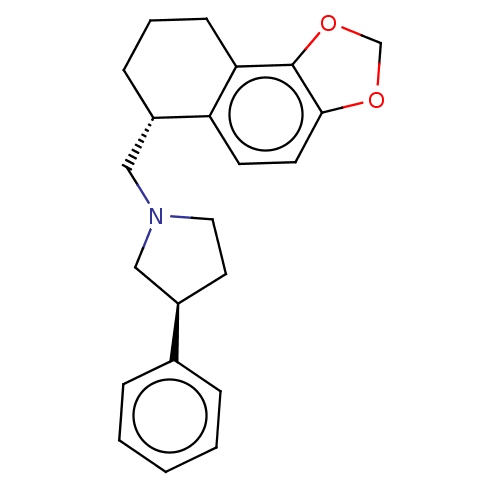
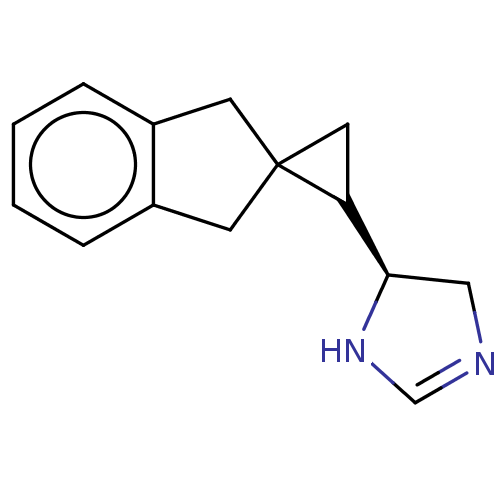
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.126nMAssay Description:In vitro binding affinity against alpha-2 adrenergic receptor of rat in presence of [3H]-RX 821002 radioligandMore data for this Ligand-Target Pair
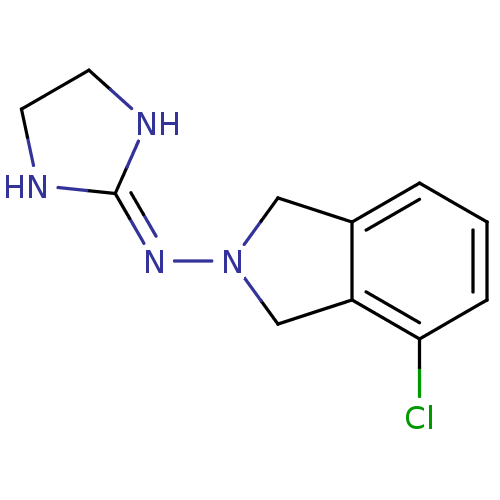
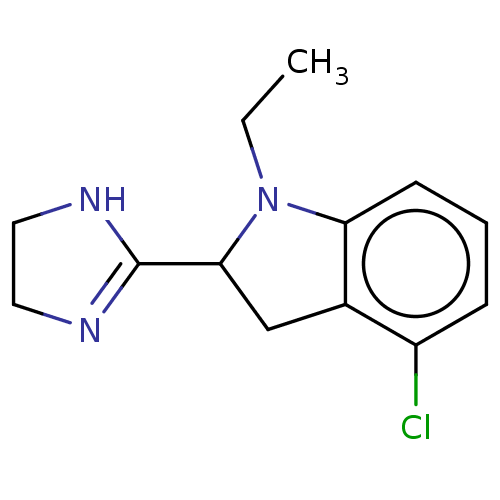
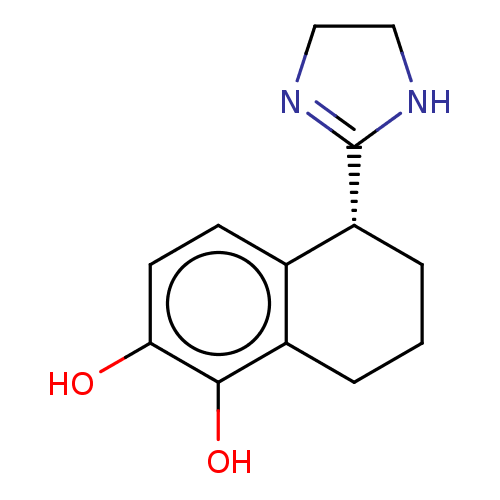
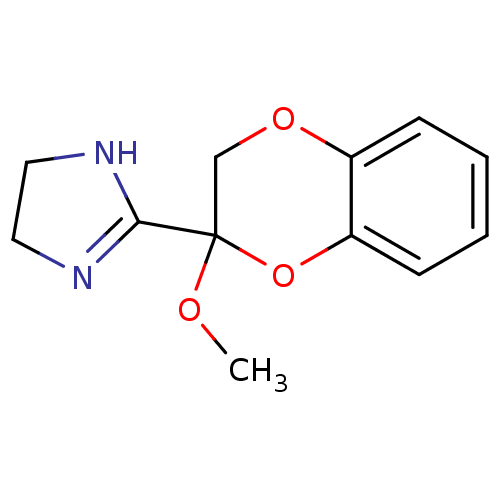
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.160nMAssay Description:Inhibition of saturable binding of [3H]idazoxan to alpha2-site in rat cerebral cortical membranes.More data for this Ligand-Target Pair
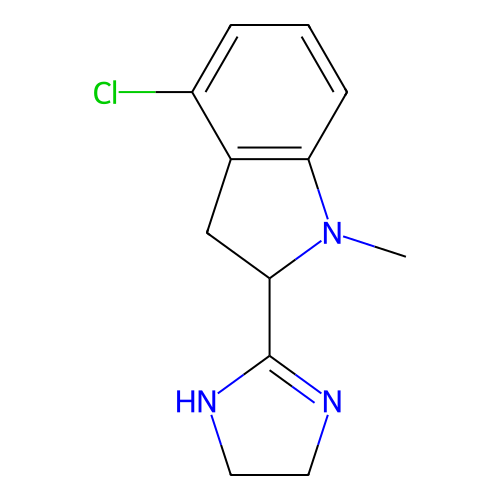
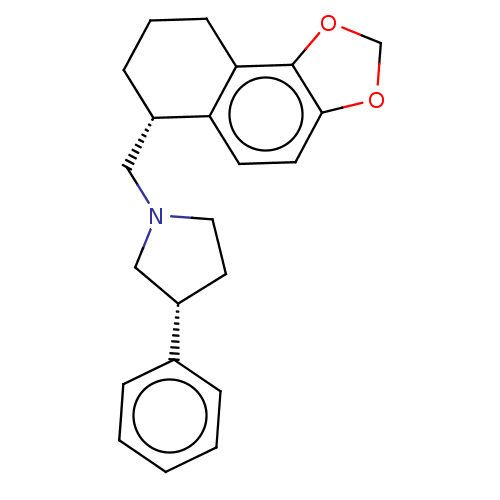
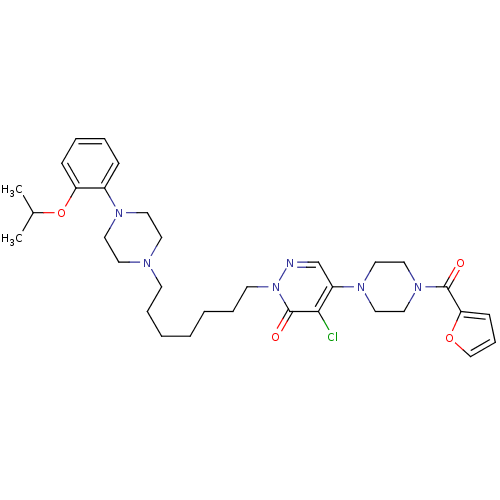
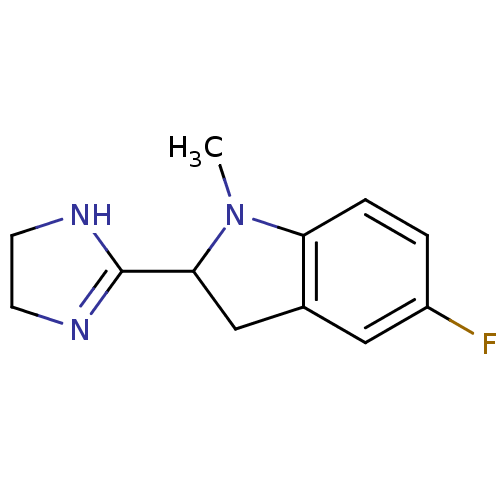
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.240nMAssay Description:Inhibition of saturable binding of [3H]idazoxan to alpha2-site in rat cerebral cortical membranes.More data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.355nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.355nMAssay Description:Binding affinity against alpha-2 adrenergic receptor in rat cerebral cortical membrane, determined using [3H]- yohimbine as the radioligand.More data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.400nMAssay Description:Binding affinity against alpha-2 adrenergic receptor was determined by the displacement of [3H]prazosin from rat brain cortical membranesMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.420nMAssay Description:Inhibition of saturable binding of [3H]idazoxan to alpha2-site in rat cerebral cortical membranes.More data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.428nMAssay Description:In vitro binding affinity at Alpha-2 adrenergic receptor in rat cortex by radioligand binding assay using [3H]rauwolscineMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.450nMAssay Description:Binding affinity against alpha-2 adrenergic receptor was determined by the displacement of [3H]clonidine from rat brain cortical membranesMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.457nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.470nMAssay Description:Binding affinity against alpha-2 adrenergic receptor was determined by the displacement of [3H]clonidine from rat brain cortical membranesMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.5nMAssay Description:In vitro binding affinity at Alpha-2 adrenergic receptor in rat cortex by radioligand binding assay using [3H]rauwolscineMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.550nMAssay Description:Binding affinity for alpha-2 adrenergic receptor in rat cortex using [3H]rauwolscine.More data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.560nMAssay Description:Binding affinity towards Alpha-2 adrenergic receptor in rat cerebral cortex using [3H]rauwolscineMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.603nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.646nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.661nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.661nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.661nMAssay Description:Binding affinity against alpha-2 adrenergic receptor in rat cerebral cortical membrane, determined using [3H]- yohimbine as the radioligand.More data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.690nMAssay Description:Inhibition of saturable binding of [3H]idazoxan to alpha2-site in rat cerebral cortical membranes.More data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.708nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.708nMAssay Description:In vitro binding affinity at Alpha-2 adrenergic receptor in rat cortex by radioligand binding assay using [3H]rauwolscineMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.708nMAssay Description:In vitro binding affinity at Alpha-2 adrenergic receptor in rat cortex by radioligand binding assay using [3H]rauwolscineMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.720nMAssay Description:Binding affinity against alpha-2 adrenergic receptor was determined by the displacement of [3H]clonidine from rat brain cortical membranesMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.794nMAssay Description:In vitro binding affinity against alpha-2 adrenergic receptor of rat in presence of [3H]-RX 821002 radioligandMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.800nMAssay Description:Displacement of [3H]idazoxan from alpha-2 adrenergic receptor from rat brain cerebral cortex.More data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.800nMAssay Description:Displacement of [3H]idazoxan from alpha-2 adrenergic receptor from rat brain cerebral cortex.More data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.910nMAssay Description:Binding affinity against alpha-2 adrenergic receptor was determined by the displacement of [3H]clonidine from rat brain cortical membranesMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.910nMAssay Description:Binding affinity against alpha-2 adrenergic receptor was determined by the displacement of [3H]clonidine from rat brain cortical membranesMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 0.960nMAssay Description:Inhibition of saturable binding of [3H]idazoxan to alpha2-site in rat cerebral cortical membranes.More data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 1.10nMAssay Description:Evaluated for its ability to displace [3H]rauwolscine from alpha-2 adrenergic receptor of rat cerebral cortexMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 1.10nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 1.10nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 1.30nMAssay Description:In vitro binding affinity against alpha-2 adrenergic receptor of rat in presence of [3H]-RX 821002 radioligandMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
The R. W. Johnson Pharmaceutical Research Institute
Curated by ChEMBL
Affinity DataKi: 1.30nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair