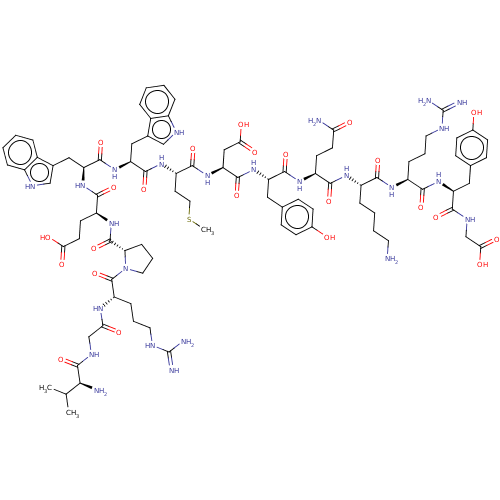
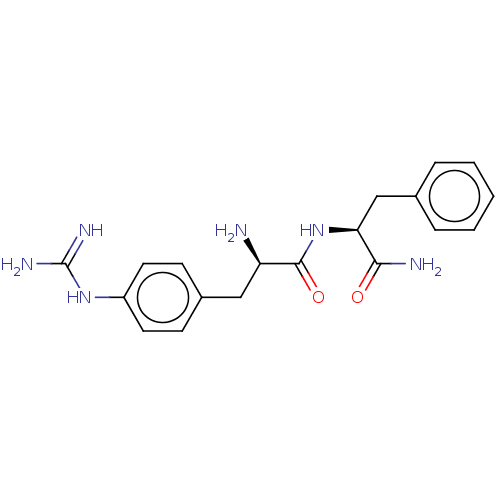
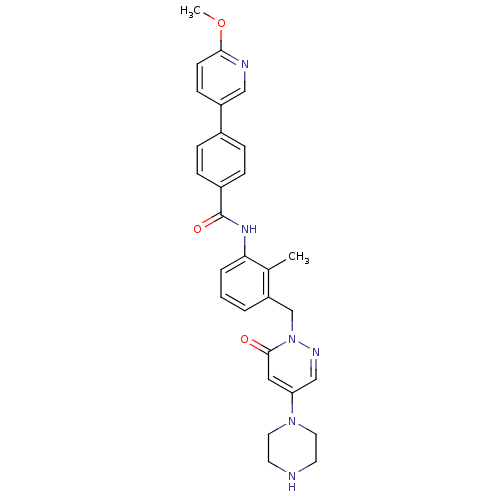
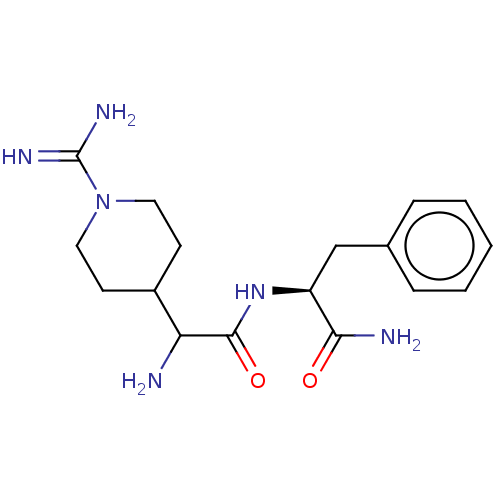
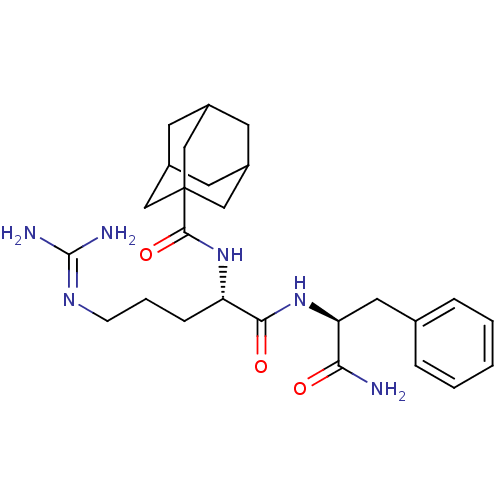
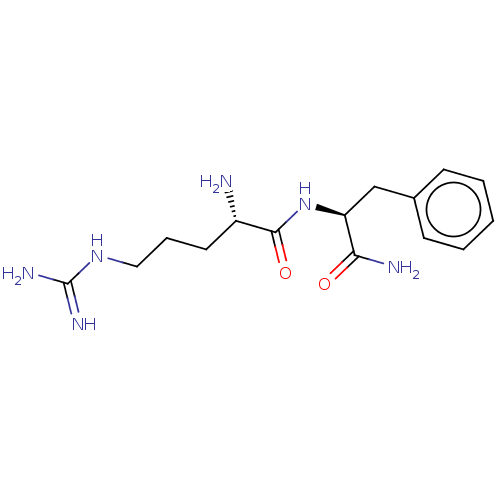
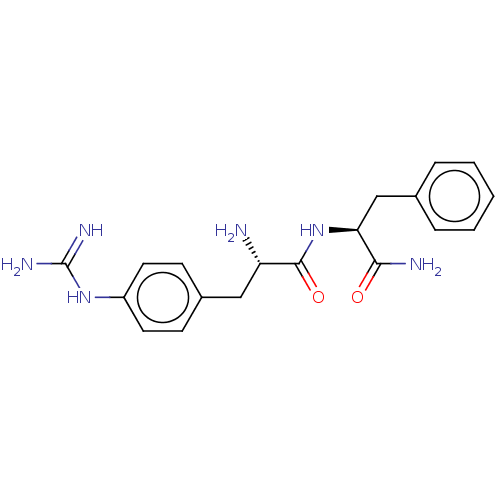
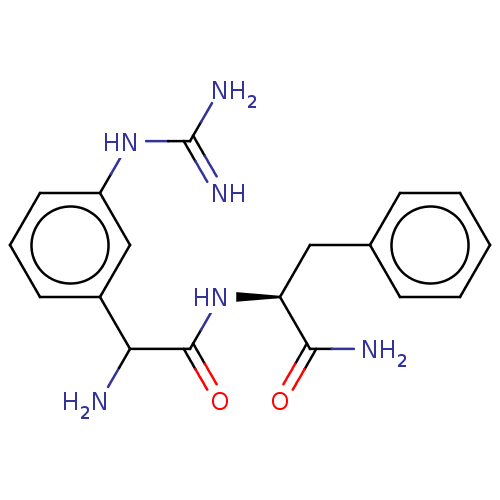
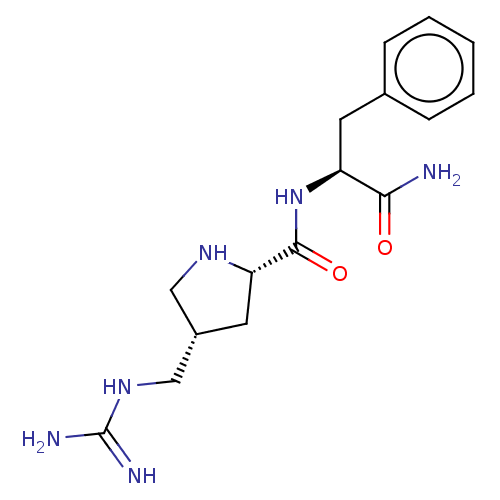
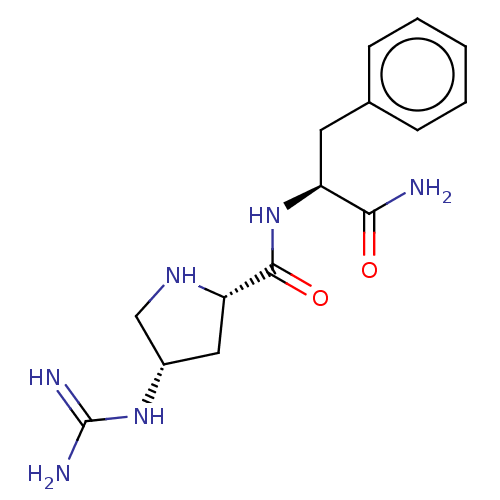
Affinity DataEC50: 300nMAssay Description:Agonist activity at rat MRGPRCMore data for this Ligand-Target Pair
Affinity DataEC50: 316nMAssay Description:Agonist activity at rat MrgX1 transfected in HEK293 cells by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 631nMAssay Description:Agonist activity at rat MrgX1 transfected in HEK293 cells by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: <2.00E+3nMAssay Description:Agonist activity at rat MrgX1 transfected in HEK293 cells by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 3.98E+3nMAssay Description:Agonist activity at rat MrgX1 transfected in HEK293 cells by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 6.31E+3nMAssay Description:Agonist activity at rat MrgX1 transfected in HEK293 cells by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 6.31E+3nMAssay Description:Agonist activity at rat MrgX1 transfected in HEK293 cells by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 5.01E+4nMAssay Description:Agonist activity at rat MrgX1 transfected in HEK293 cells by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: <1.00E+5nMAssay Description:Agonist activity at rat MrgX1 transfected in HEK293 cells by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: <1.00E+5nMAssay Description:Agonist activity at rat MrgX1 transfected in HEK293 cells by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: <1.00E+5nMAssay Description:Agonist activity at rat MrgX1 transfected in HEK293 cells by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: <1.00E+5nMAssay Description:Agonist activity at rat MrgX1 transfected in HEK293 cells by FLIPR assayMore data for this Ligand-Target Pair