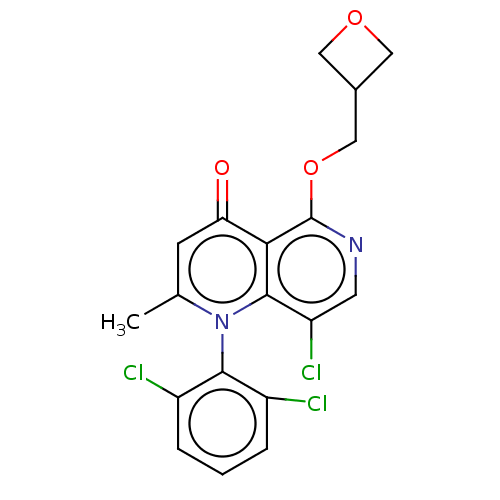
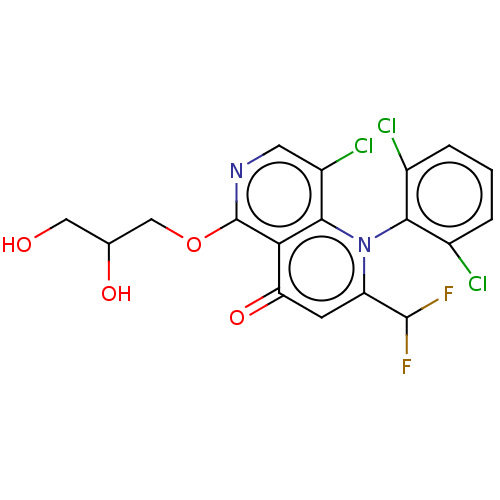
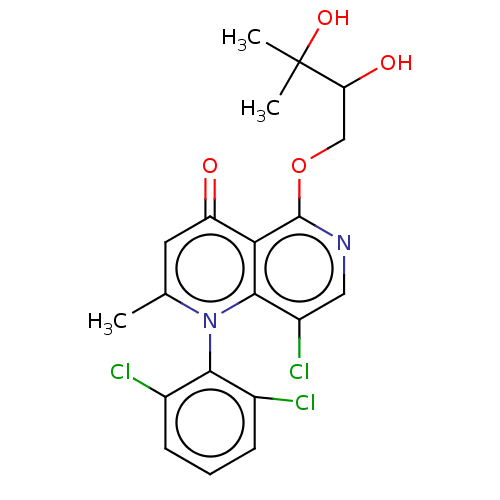
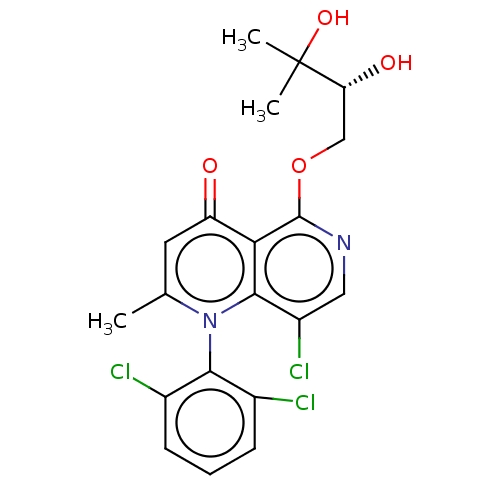
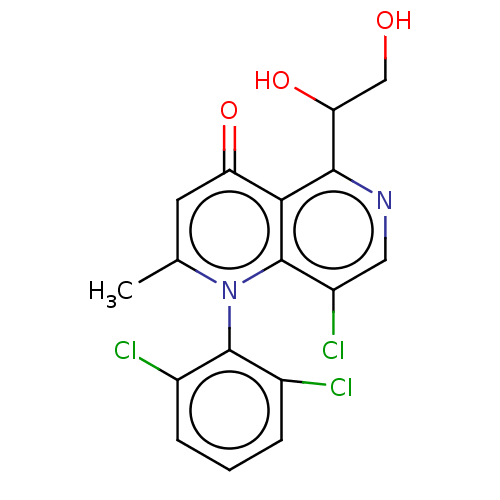
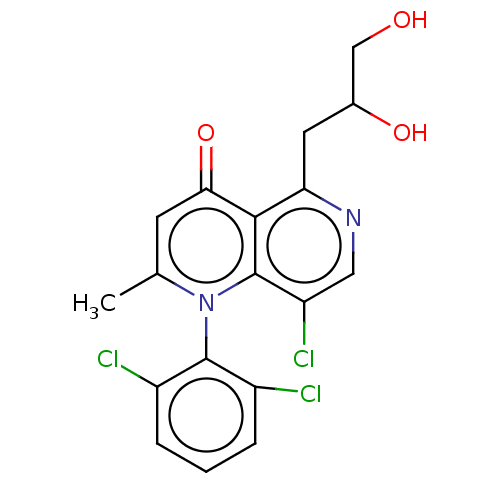
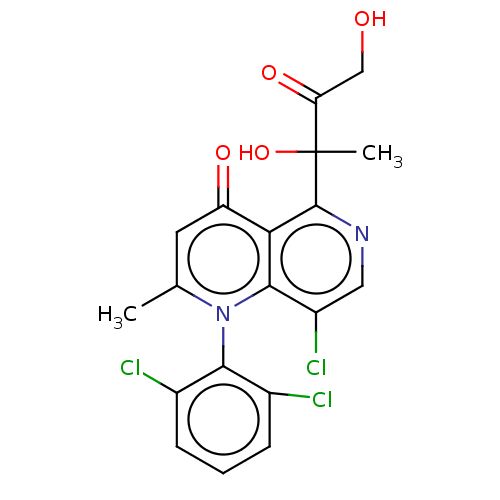
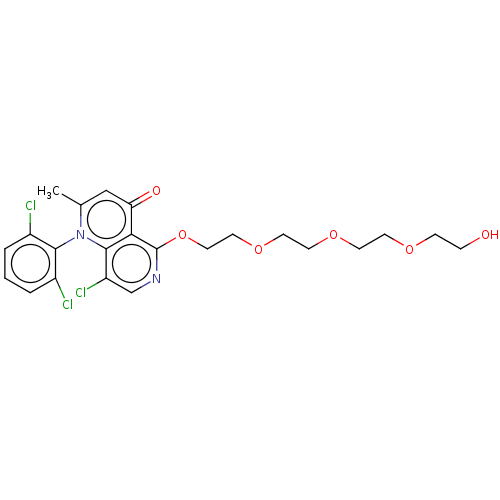
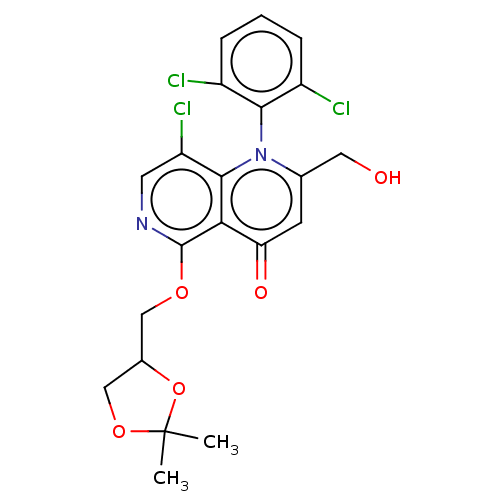
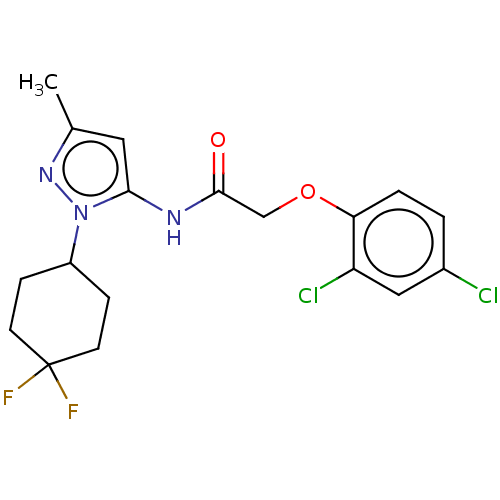
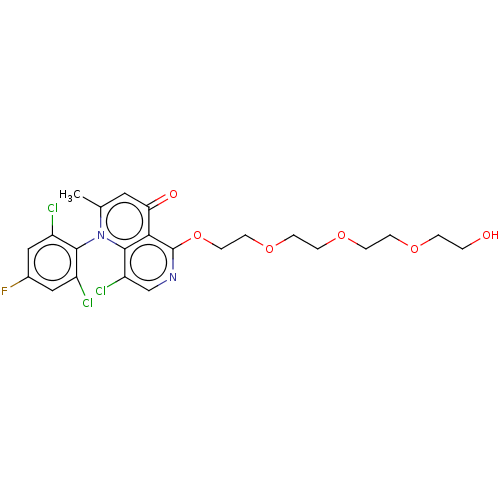
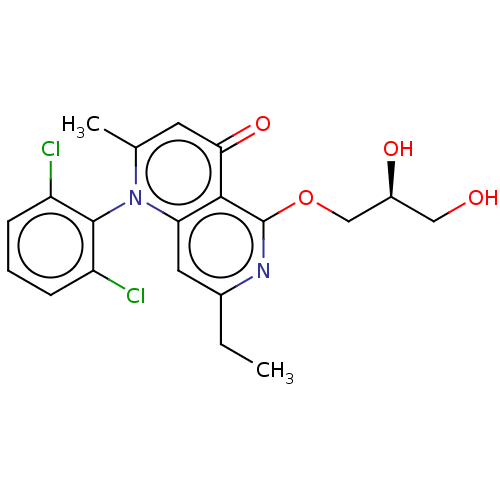
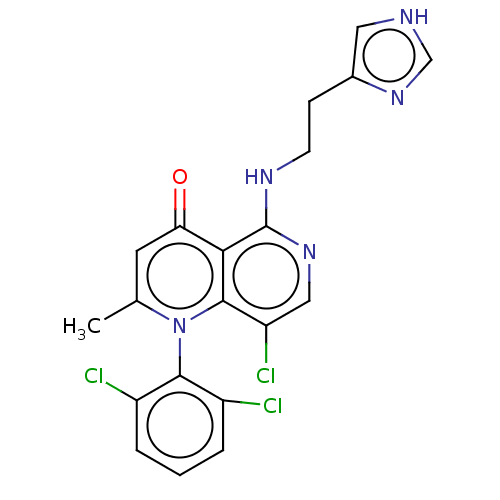
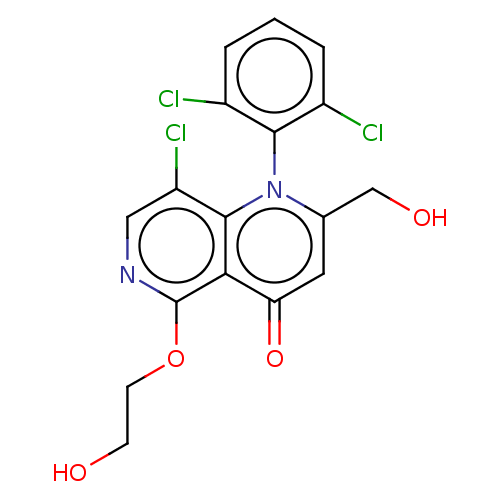
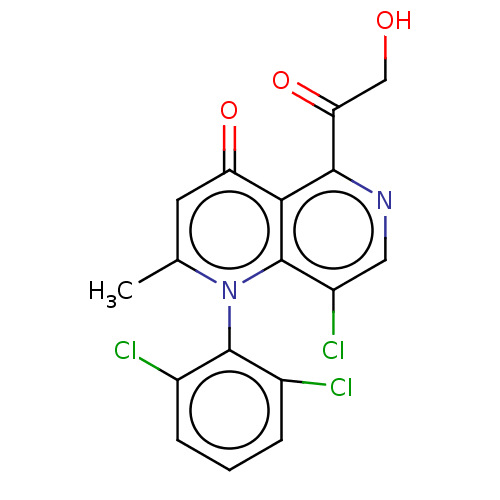
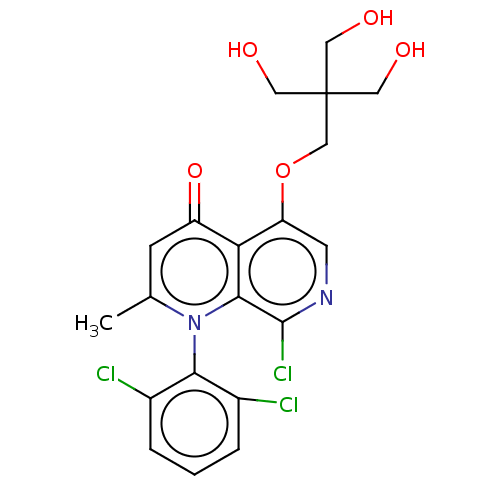
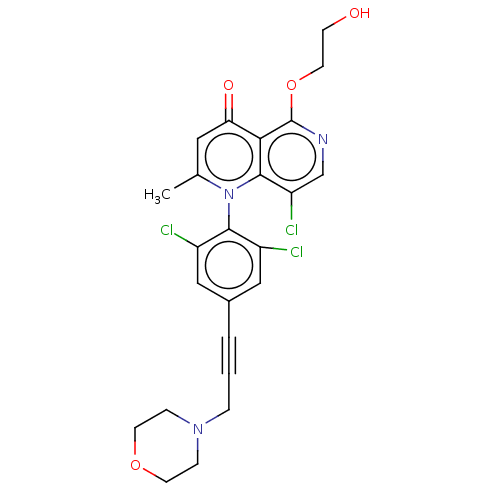
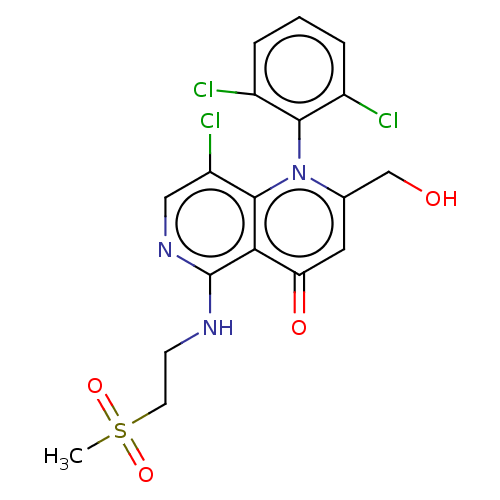
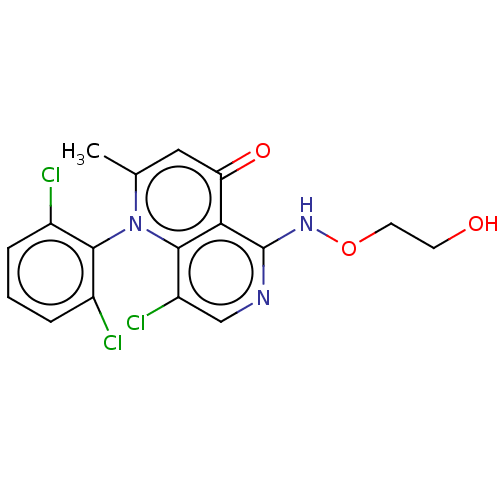
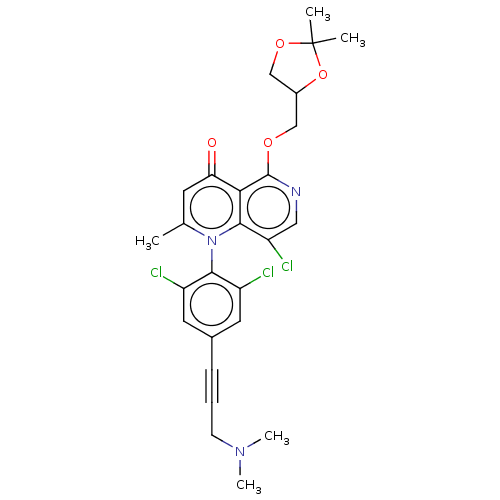
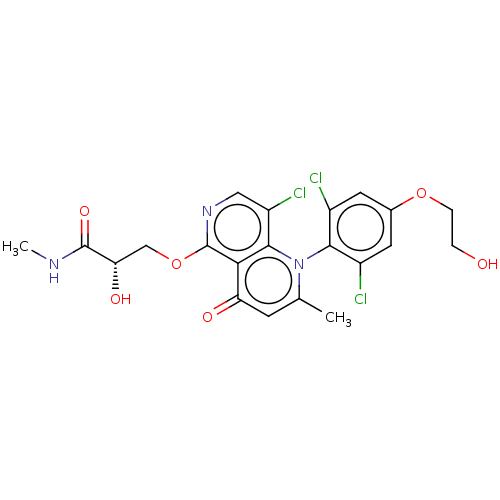
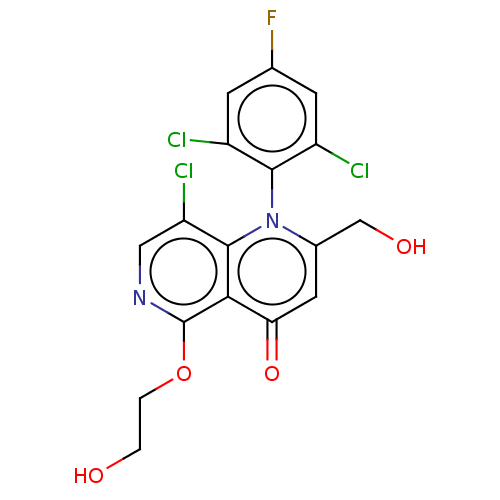
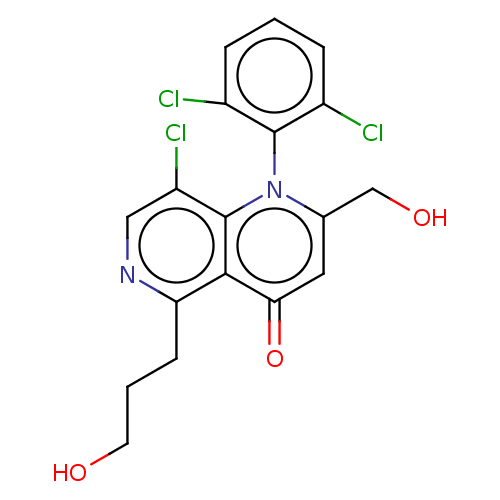
Affinity DataIC50: 6nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
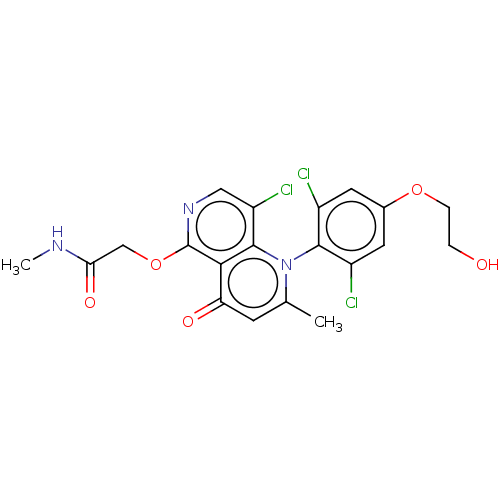
Affinity DataIC50: 8nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
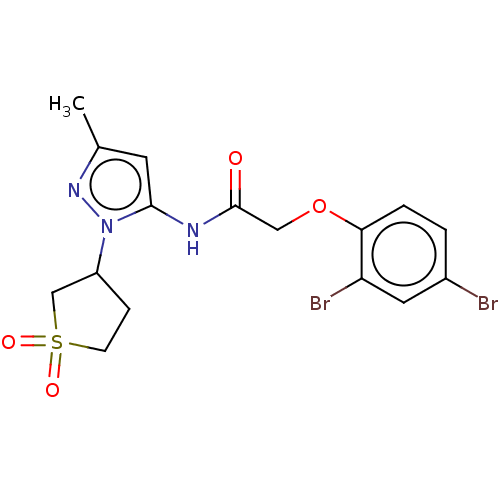
Affinity DataIC50: 9nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
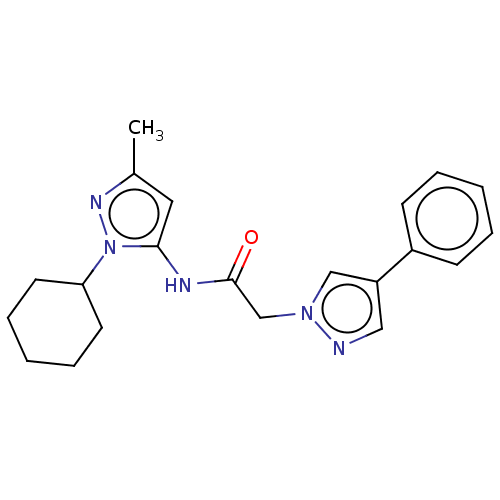
Affinity DataIC50: 9nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 17nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 24nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 25nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 25nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 28nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 28nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 29nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:The GIRK1/4 activity of a compound according to the present invention was assessed by the following in vitro method.Buffers:a. External buffer: 10 mM...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 30nMAssay Description:The GIRK1/4 activity of a compound according to the present invention was assessed by the following in vitro method.Buffers:a. External buffer: 10 mM...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 31nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 32nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
TargetG protein-activated inward rectifier potassium channel 1(Human)
University of Nebraska
Curated by ChEMBL
University of Nebraska
Curated by ChEMBL
Affinity DataEC50: 33nMAssay Description:Activation of GIRK1/2 channel (unknown origin) expressed in HEK293 cells incubated for 6 mins by thallium flux assayMore data for this Ligand-Target Pair
Affinity DataIC50: 33nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 34nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 35nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 36nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 37nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 39nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 41nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 42nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 46nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 47nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:The GIRK1/4 activity of a compound according to the present invention was assessed by the following in vitro method.Buffers:a. External buffer: 10 mM...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 50nMAssay Description:The GIRK1/4 activity of a compound according to the present invention was assessed by the following in vitro method.Buffers:a. External buffer: 10 mM...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 52nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 54nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 56nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 59nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 60nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 60nMAssay Description:The GIRK1/4 activity of a compound according to the present invention was assessed by the following in vitro method.Buffers:a. External buffer: 10 mM...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 60nMAssay Description:The GIRK1/4 activity of a compound according to the present invention was assessed by the following in vitro method.Buffers:a. External buffer: 10 mM...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetG protein-activated inward rectifier potassium channel 1(Human)
University of Nebraska
Curated by ChEMBL
University of Nebraska
Curated by ChEMBL
Affinity DataEC50: 64nMAssay Description:Activation of GIRK1/2 channel (unknown origin) expressed in HEK293 cells incubated for 6 mins by thallium flux assayMore data for this Ligand-Target Pair
TargetG protein-activated inward rectifier potassium channel 1(Human)
University of Nebraska
Curated by ChEMBL
University of Nebraska
Curated by ChEMBL
Affinity DataEC50: 65nMAssay Description:Activation of human GIRK1/2 expressed in HEK293 cells co-expressing rat mGlu8 by thallium flux assayMore data for this Ligand-Target Pair
Affinity DataIC50: 68nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair
Affinity DataIC50: 69nMAssay Description:Quattro is controlled using IonWorks v2 software to perform the following steps:a. Add 3.5 ul cells plus 3.5 ul external buffer to wells of Quattro P...More data for this Ligand-Target Pair