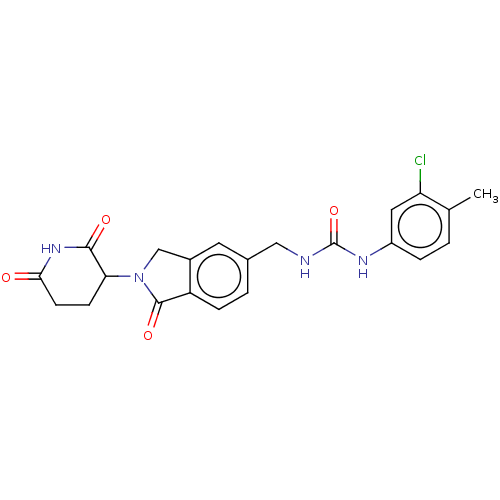
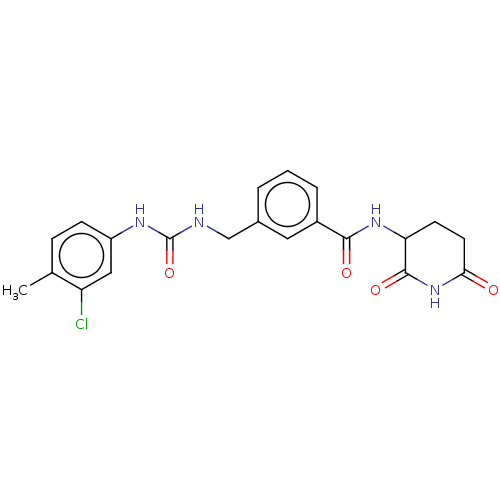
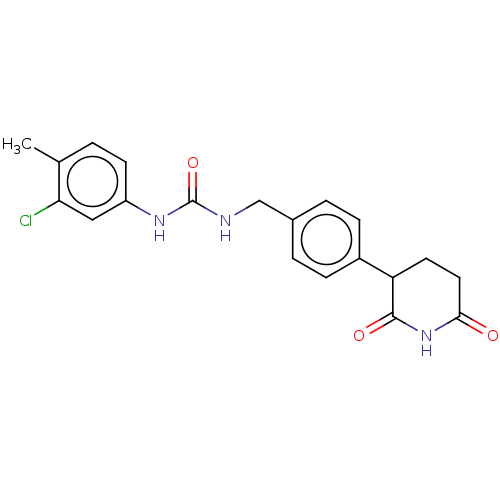
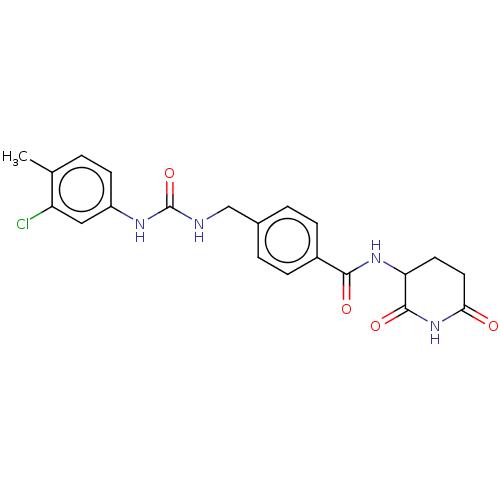
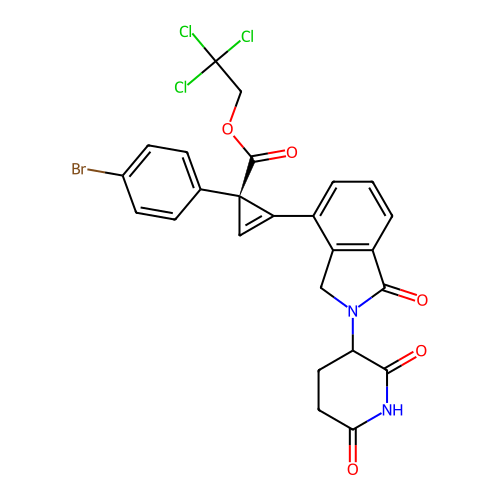
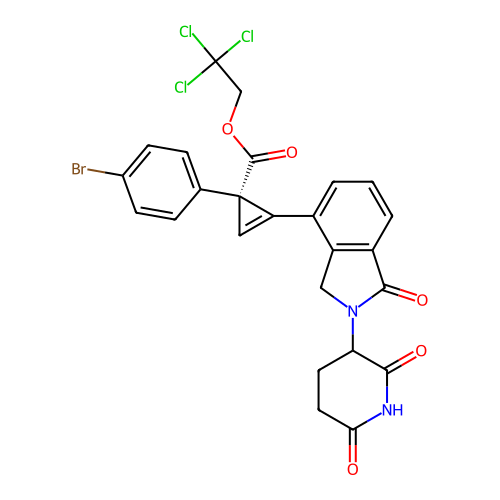
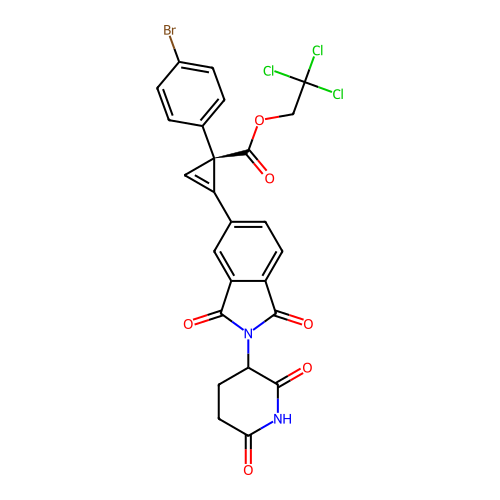
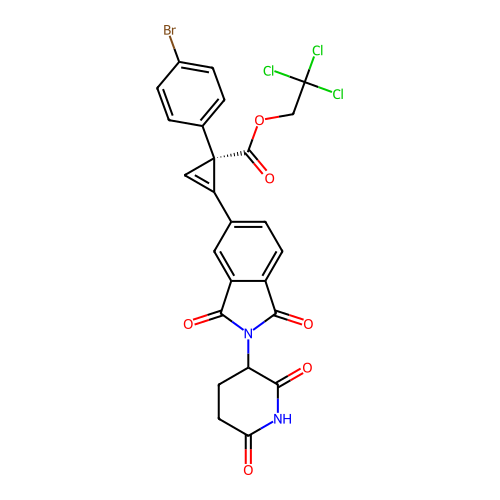
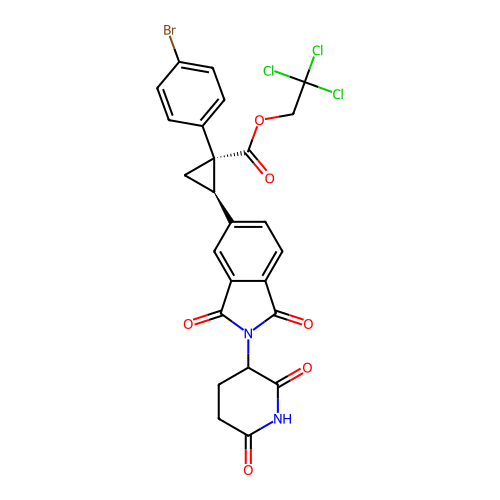
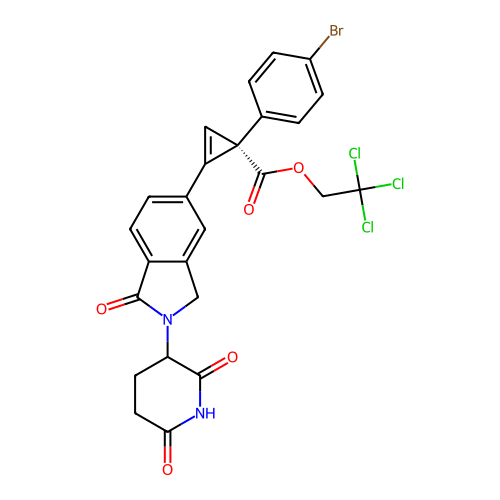
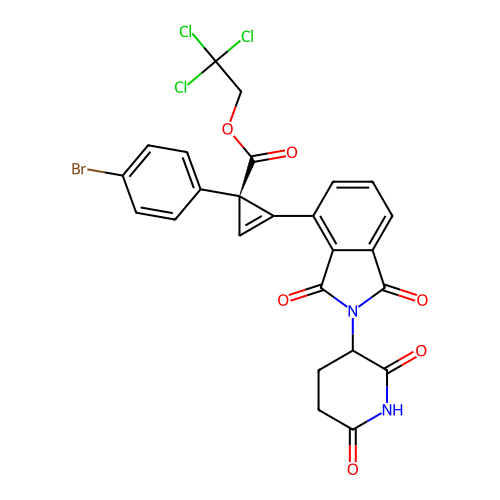
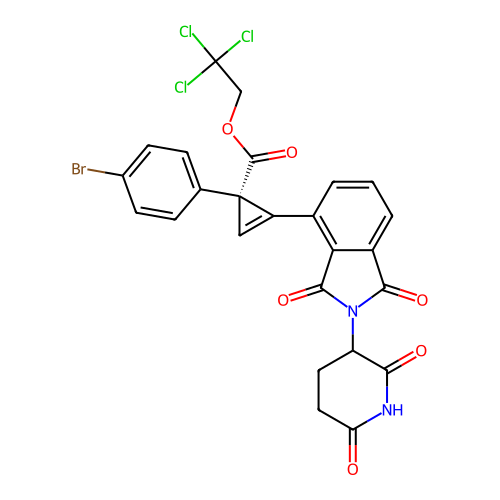
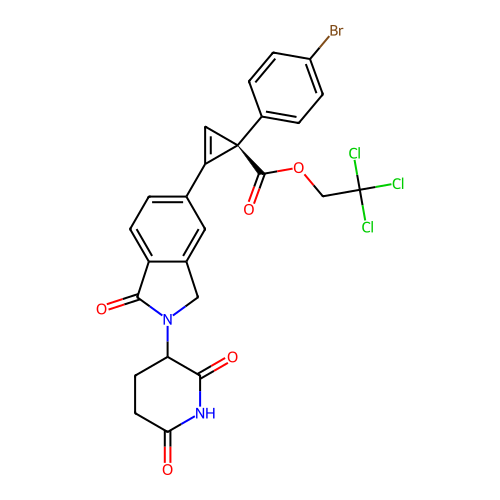
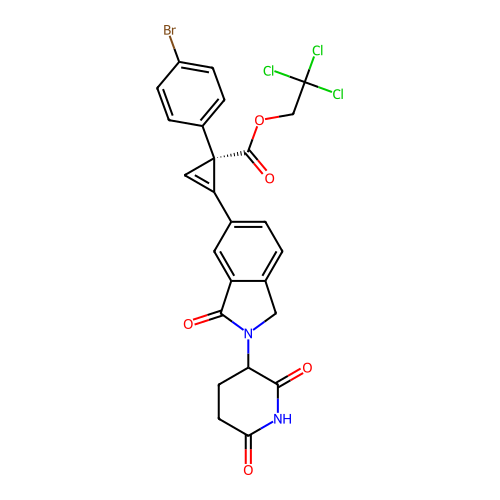
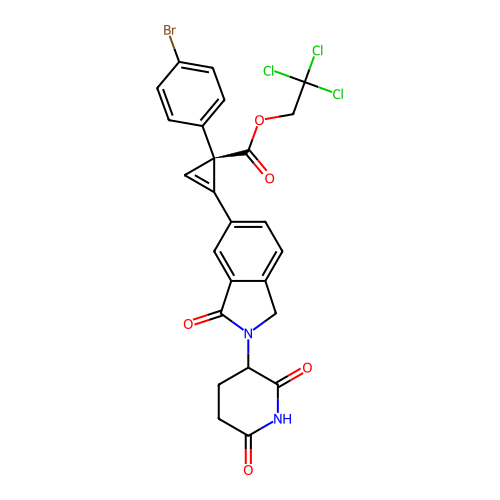
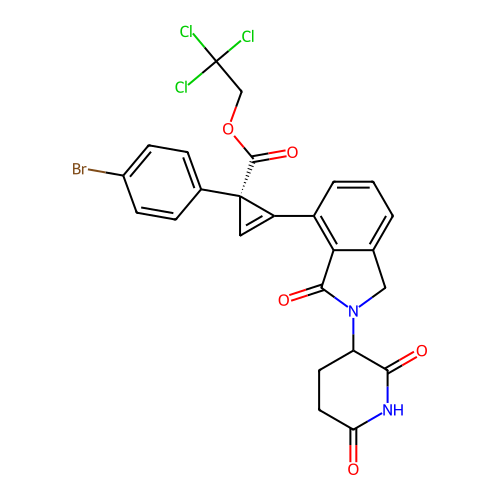
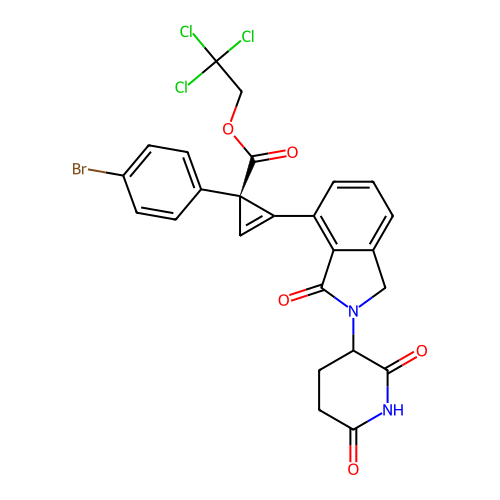
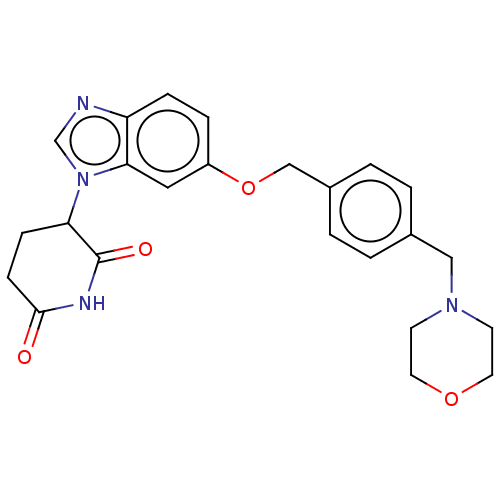
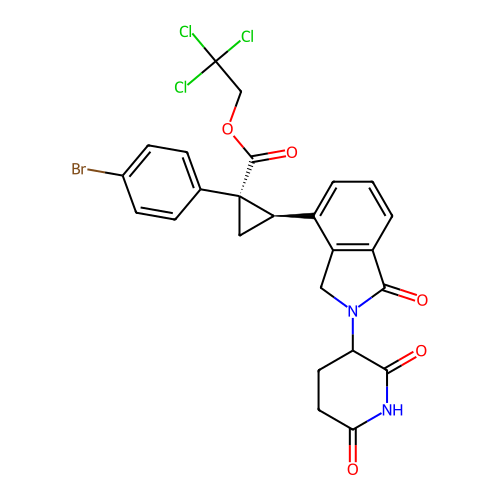
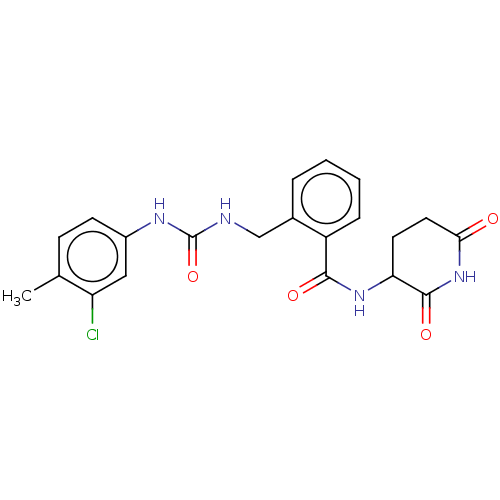
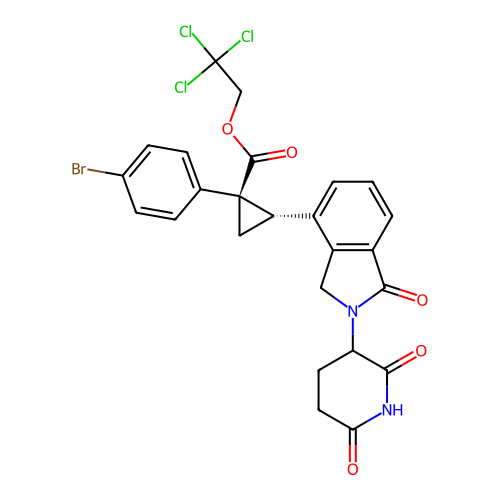
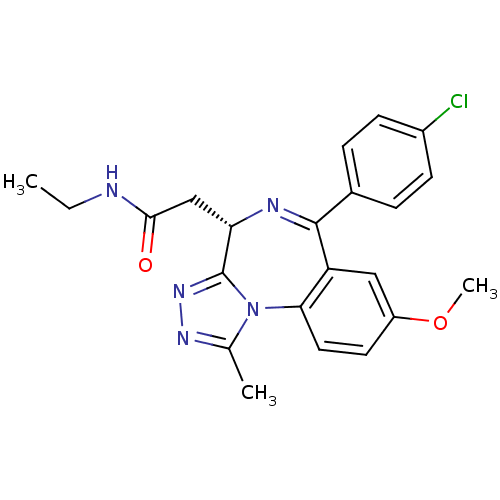
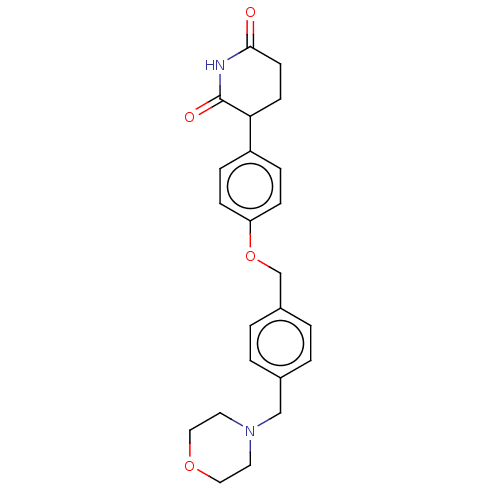
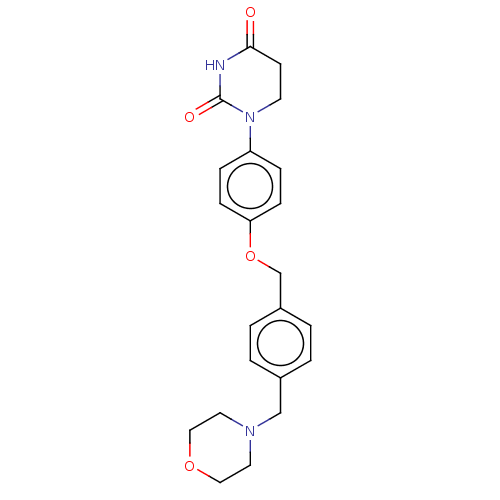
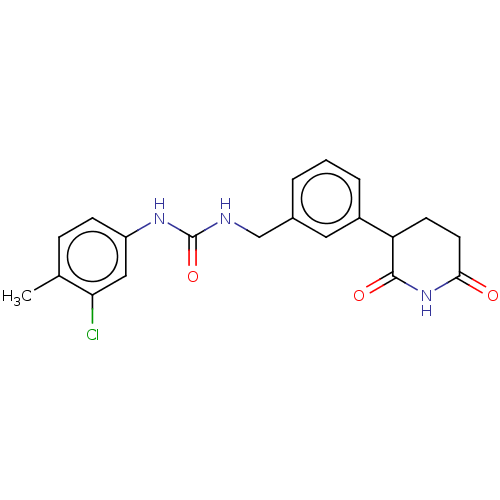
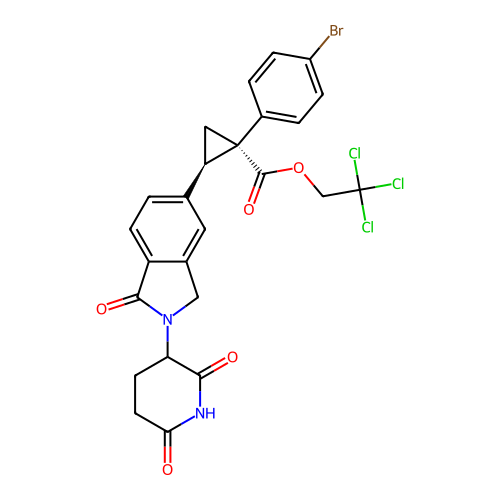
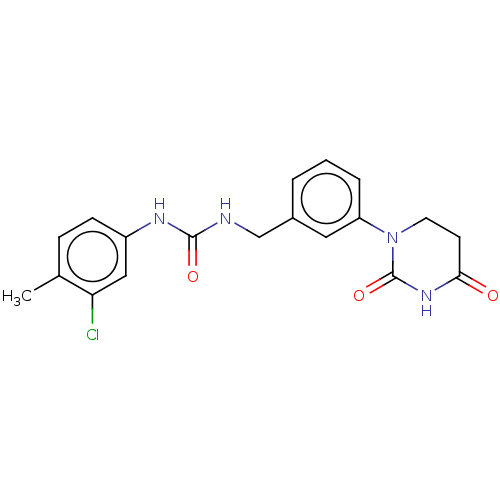
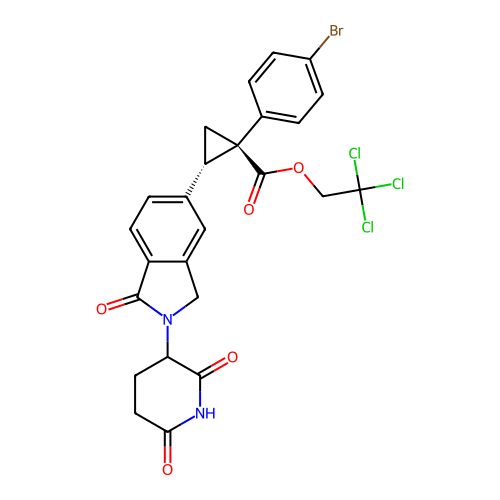
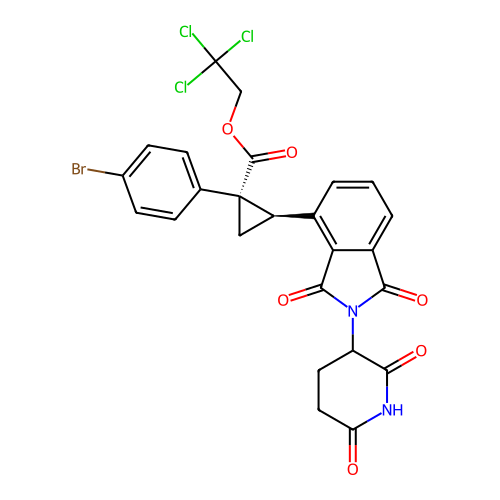
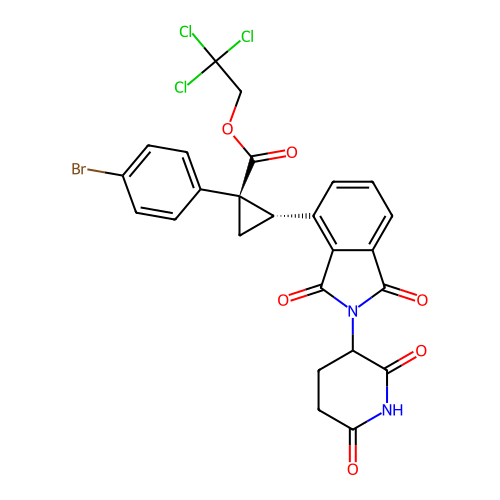
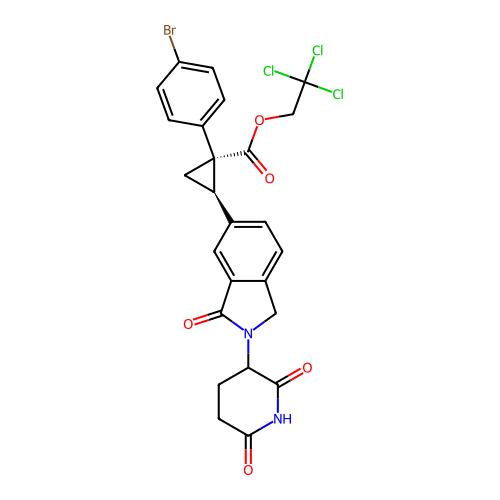
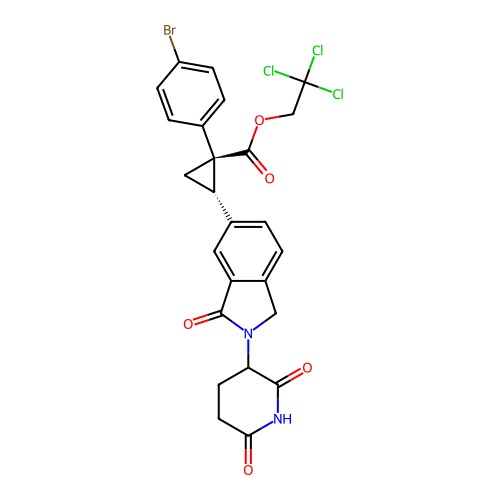
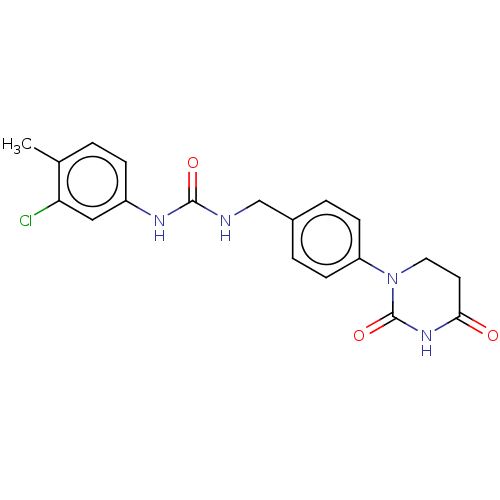
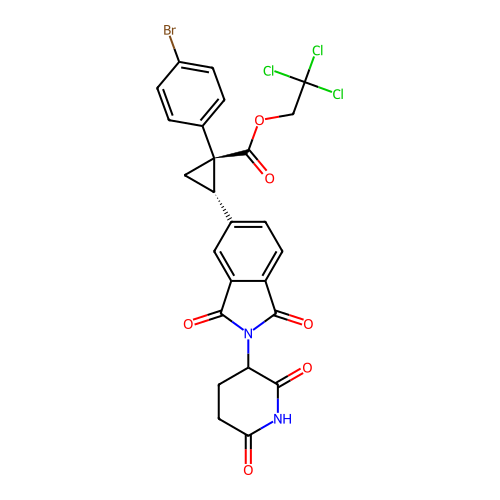
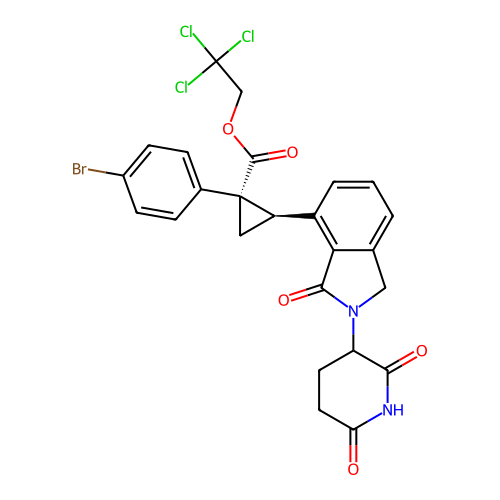
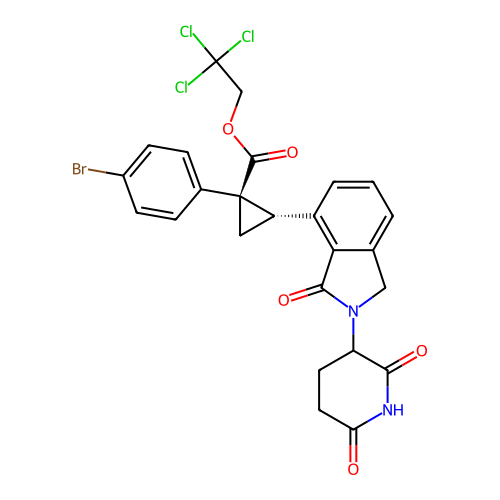
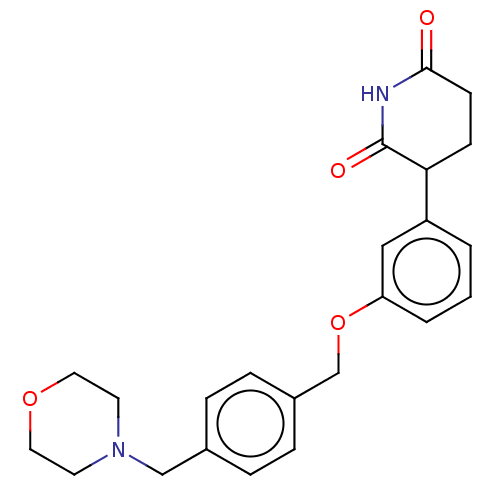
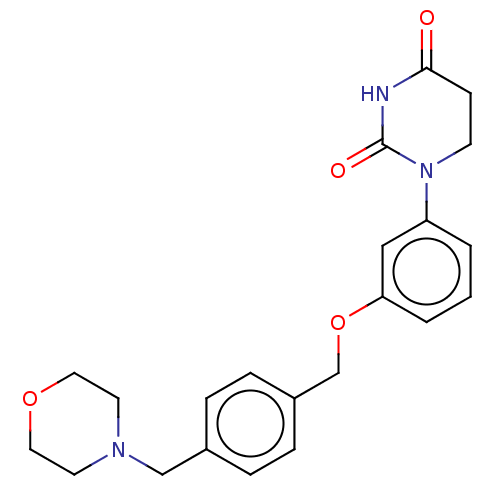
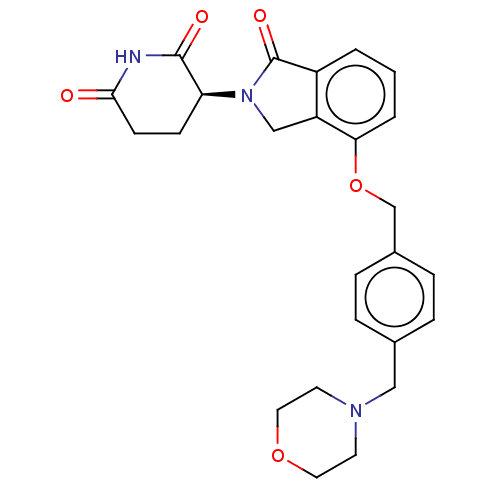
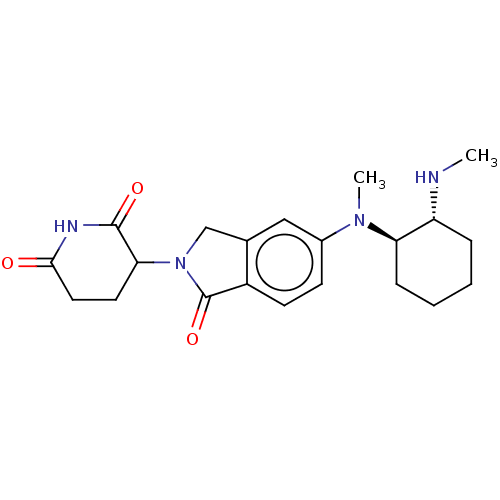
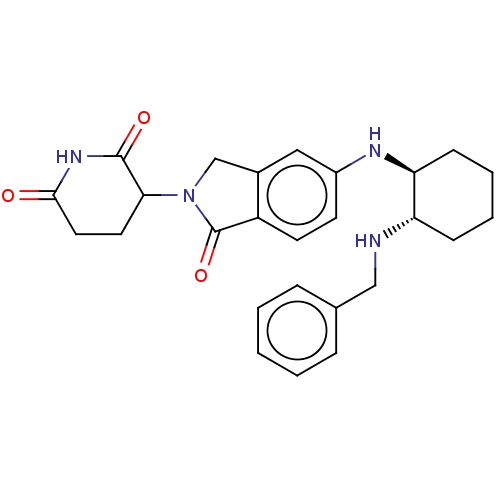
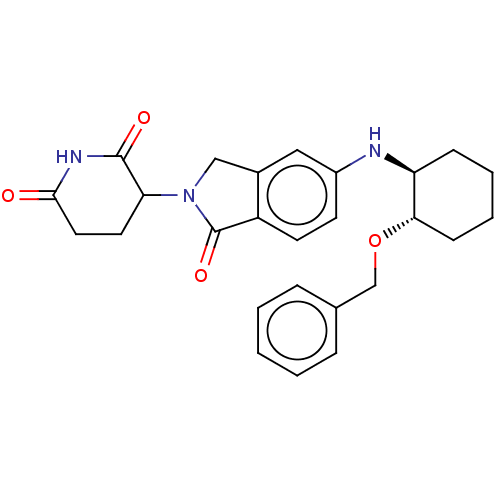
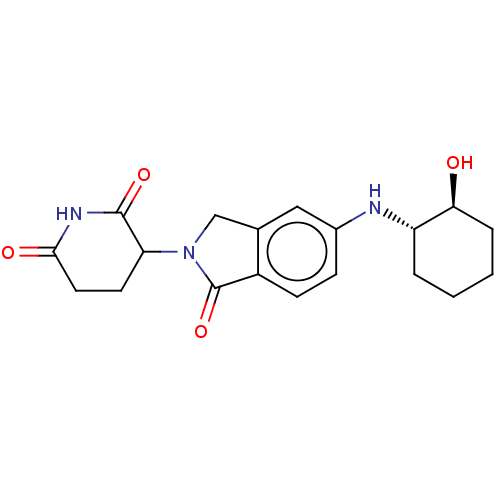
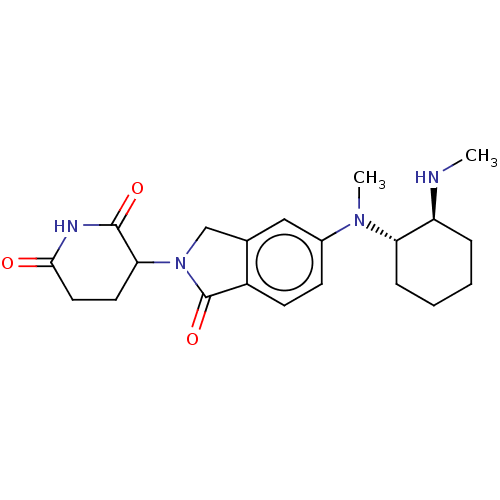
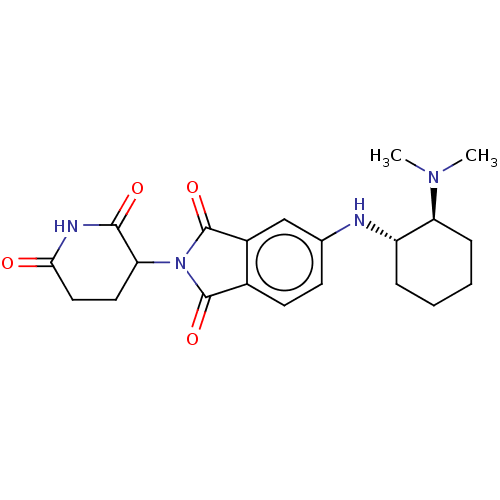
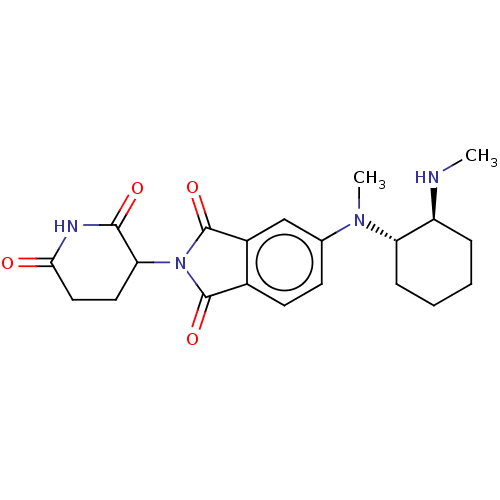
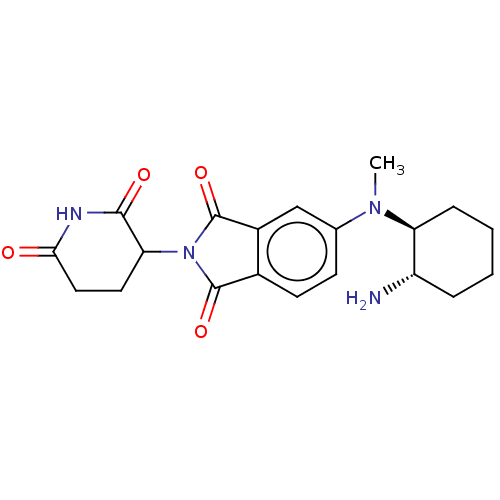
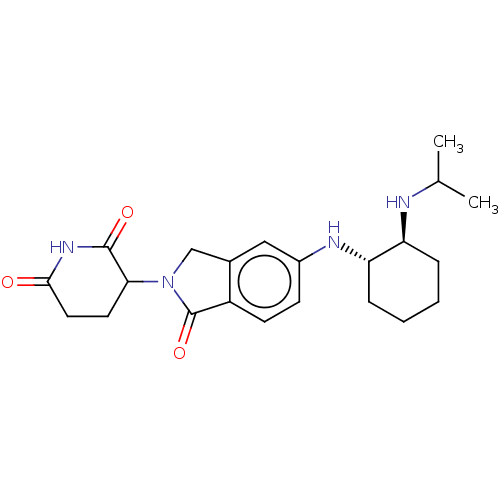
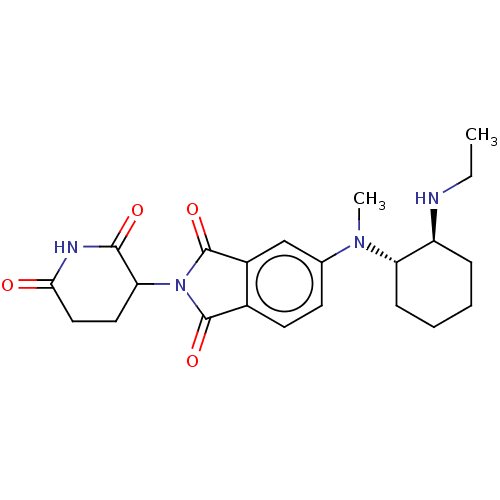
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 0.300nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 20 hrs by luminescence based assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 21nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 20 hrs by luminescence based assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 70nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 20 hrs by luminescence based assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 100nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 20 hrs by luminescence based assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 170nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 550nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 1.10E+3nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 1.40E+3nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >3.10E+3nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 3.30E+3nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 4.30E+3nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 5.10E+3nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 20 hrs by luminescence based assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 20 hrs by luminescence based assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of GSPT1 (unknown origin) incubated for 1 hr by colloidal coomassie staining based LC-MS/MS analysisMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 20 hrs by luminescence based assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 20 hrs by luminescence based assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 20 hrs by luminescence based assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 20 hrs by luminescence based assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 20 hrs by luminescence based assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 60 mins by luminescence reader assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 20 hrs by luminescence based assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 20 hrs by luminescence based assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >1.00E+4nMAssay Description:Induction of ePL tagged GSPT1 degradation in human DF15 cells incubated for 20 hrs by luminescence based assayMore data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >3.00E+4nMAssay Description:The Prolabel system from DiscoverX was used to develop high-throughput and quantitative assays to measure changes in IKZF1, IKZF2, and GSPT1 protein ...More data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >3.00E+4nMAssay Description:The Prolabel system from DiscoverX was used to develop high-throughput and quantitative assays to measure changes in IKZF1, IKZF2, and GSPT1 protein ...More data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >3.00E+4nMAssay Description:The Prolabel system from DiscoverX was used to develop high-throughput and quantitative assays to measure changes in IKZF1, IKZF2, and GSPT1 protein ...More data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >3.00E+4nMAssay Description:The Prolabel system from DiscoverX was used to develop high-throughput and quantitative assays to measure changes in IKZF1, IKZF2, and GSPT1 protein ...More data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >5.00E+4nMAssay Description:The Prolabel system from DiscoverX was used to develop high-throughput and quantitative assays to measure changes in IKZF1, IKZF2, and GSPT1 protein ...More data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >5.00E+4nMAssay Description:The Prolabel system from DiscoverX was used to develop high-throughput and quantitative assays to measure changes in IKZF1, IKZF2, and GSPT1 protein ...More data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >5.00E+4nMAssay Description:The Prolabel system from DiscoverX was used to develop high-throughput and quantitative assays to measure changes in IKZF1, IKZF2, and GSPT1 protein ...More data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >5.00E+4nMAssay Description:The Prolabel system from DiscoverX was used to develop high-throughput and quantitative assays to measure changes in IKZF1, IKZF2, and GSPT1 protein ...More data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >5.00E+4nMAssay Description:The Prolabel system from DiscoverX was used to develop high-throughput and quantitative assays to measure changes in IKZF1, IKZF2, and GSPT1 protein ...More data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >5.00E+4nMAssay Description:The Prolabel system from DiscoverX was used to develop high-throughput and quantitative assays to measure changes in IKZF1, IKZF2, and GSPT1 protein ...More data for this Ligand-Target Pair
TargetEukaryotic peptide chain release factor GTP-binding subunit ERF3A(Human)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataEC50: >5.00E+4nMAssay Description:The Prolabel system from DiscoverX was used to develop high-throughput and quantitative assays to measure changes in IKZF1, IKZF2, and GSPT1 protein ...More data for this Ligand-Target Pair