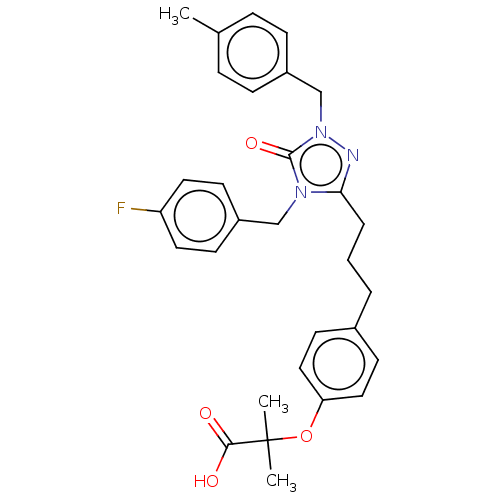
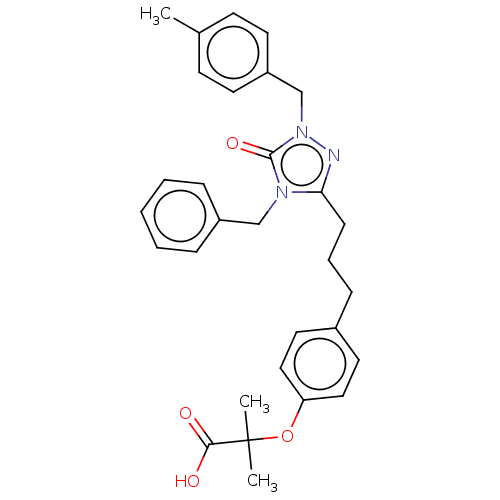
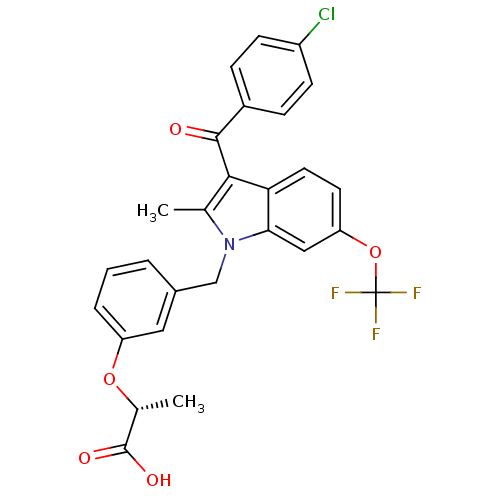
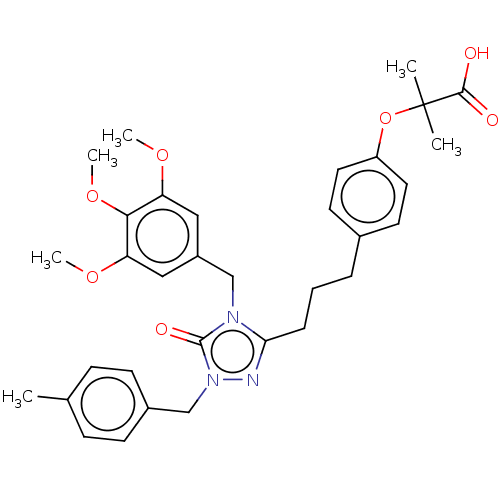
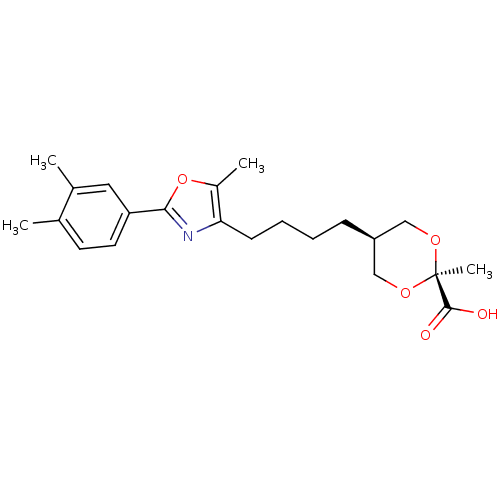
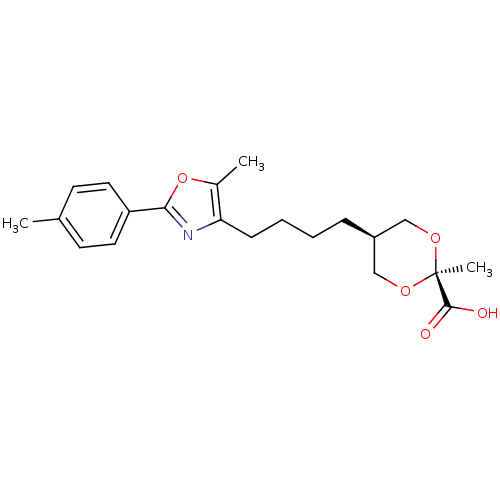
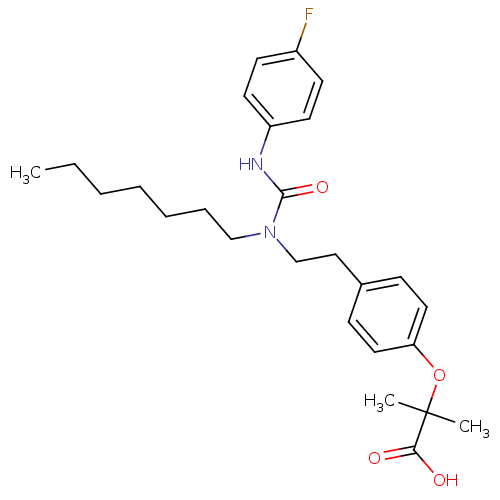
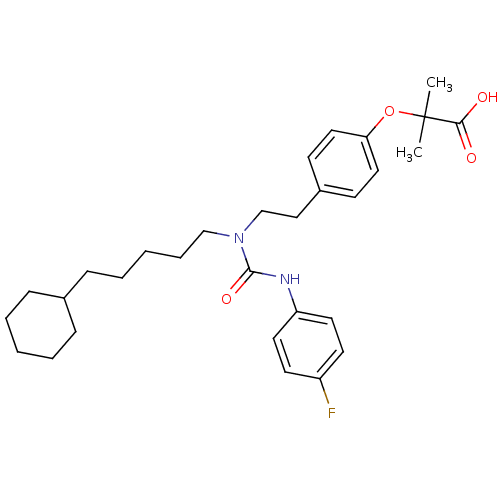
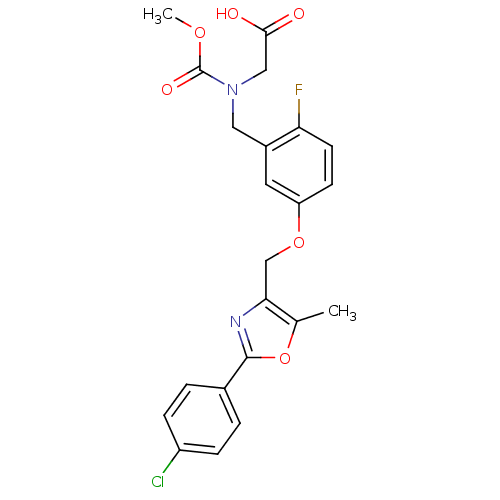
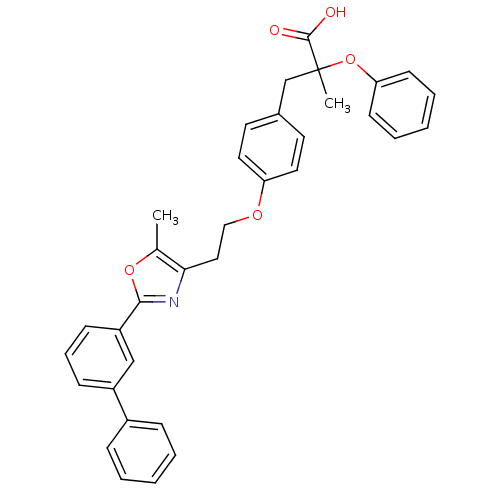
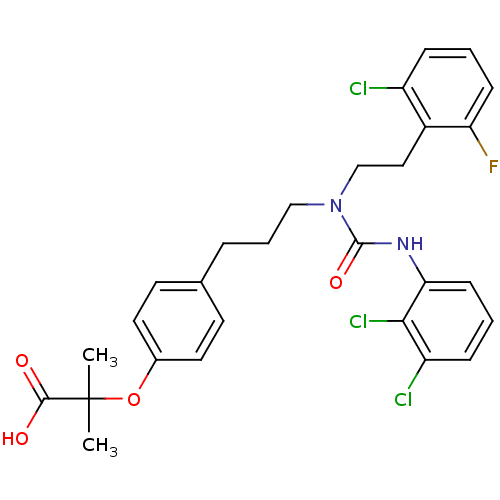
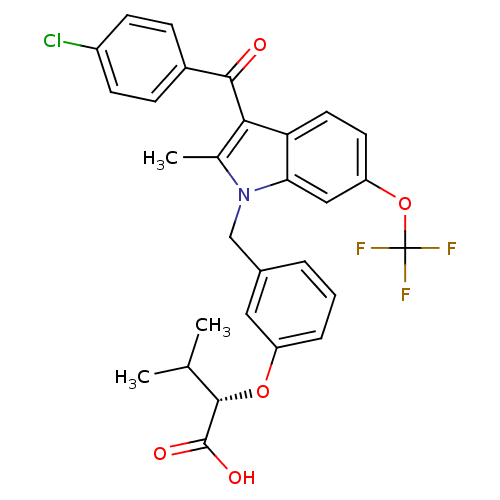
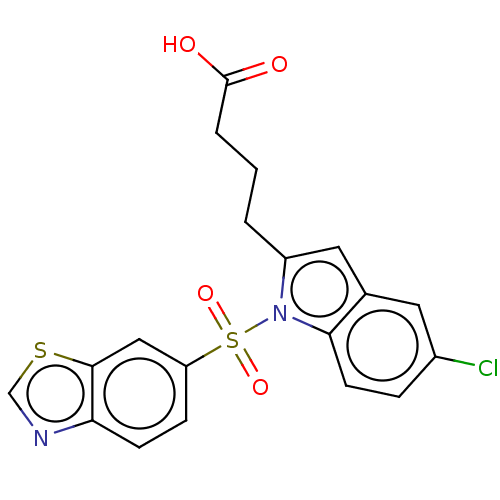
Affinity DataEC50: 0.150nMAssay Description:Effective concentration against murine PPARalpha in transactivation assayMore data for this Ligand-Target Pair
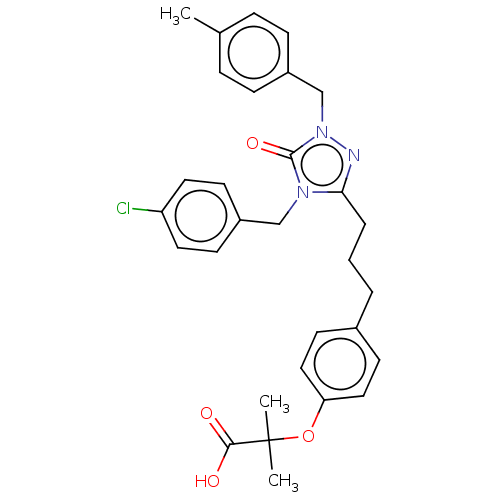
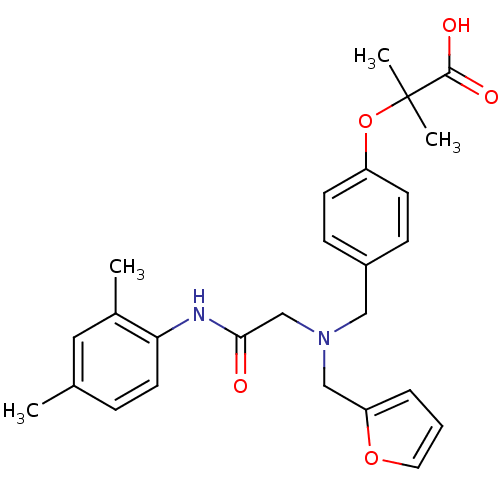
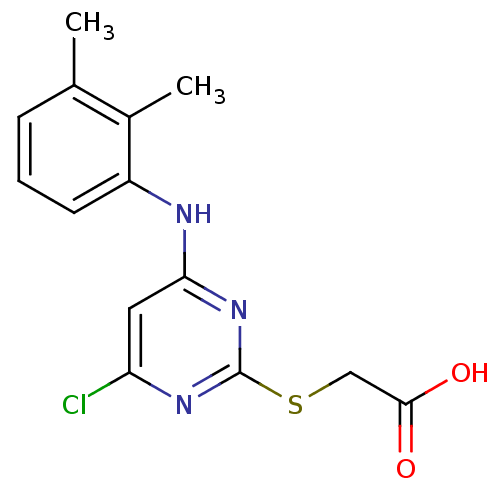
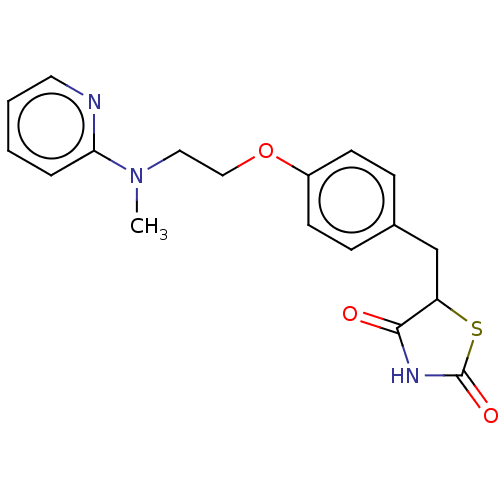
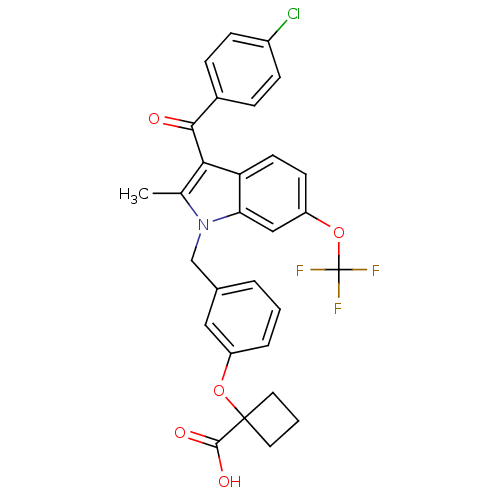
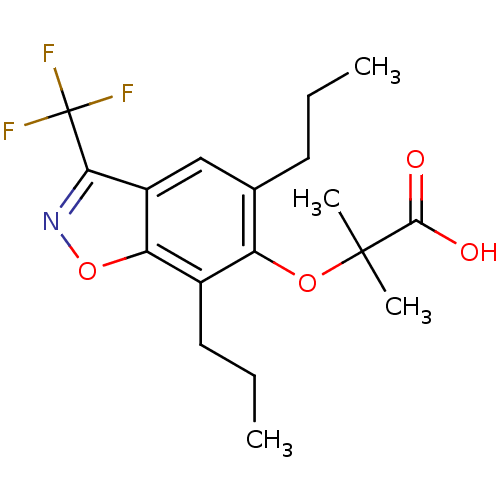
Affinity DataEC50: 0.200nMAssay Description:Effective concentration against murine PPARalpha in transactivation assayMore data for this Ligand-Target Pair
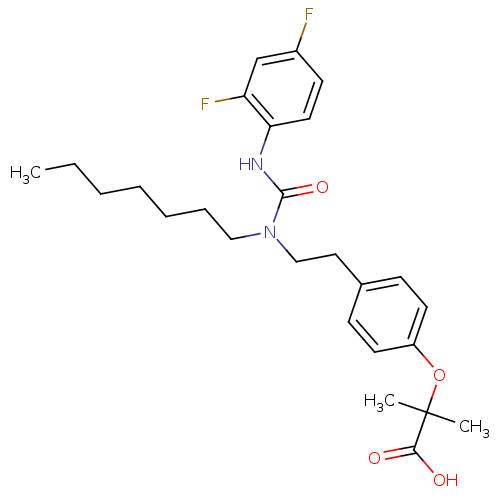
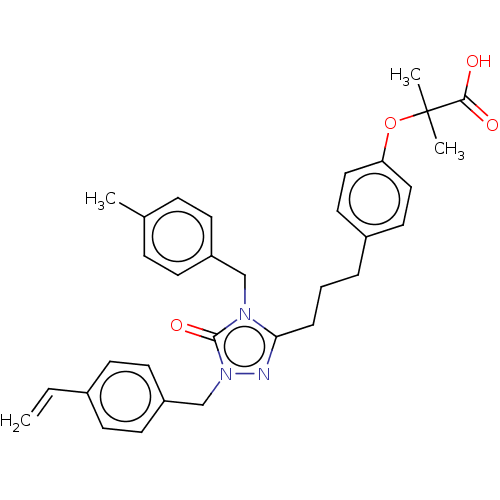
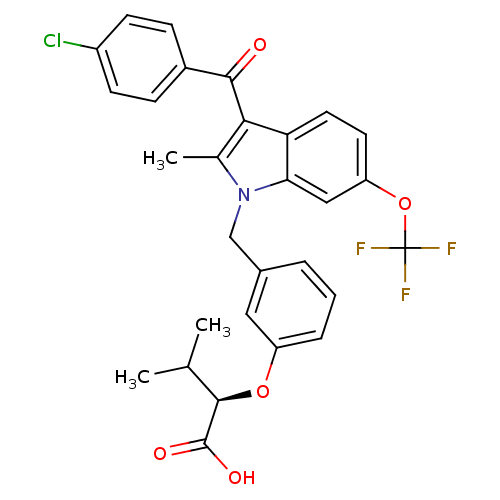
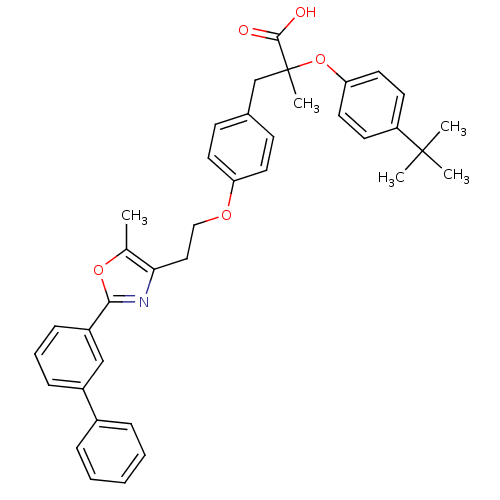
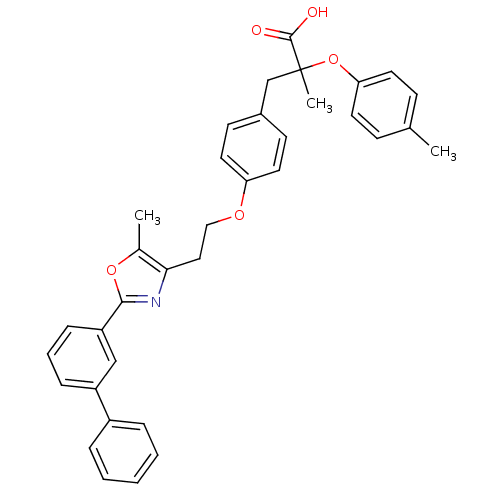
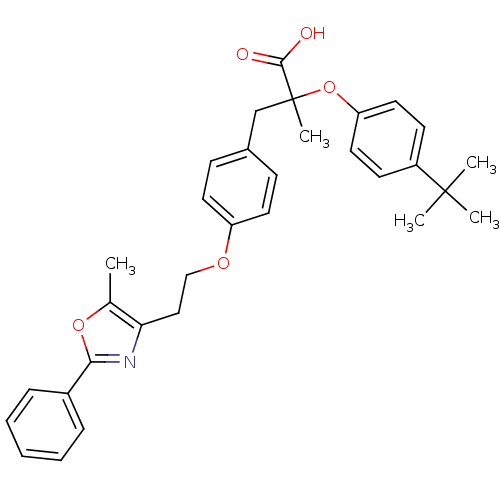
Affinity DataEC50: 0.270nMAssay Description:Effective concentration against murine PPARalpha in transactivation assayMore data for this Ligand-Target Pair
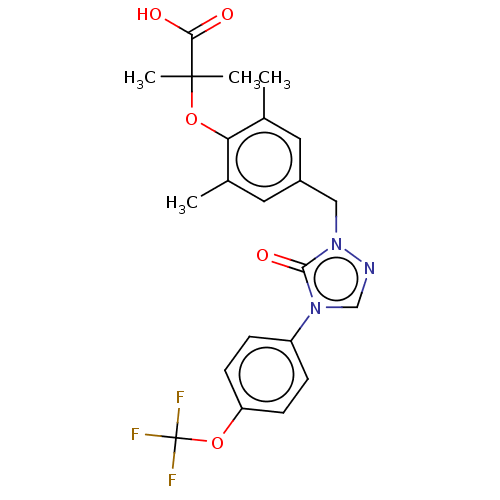
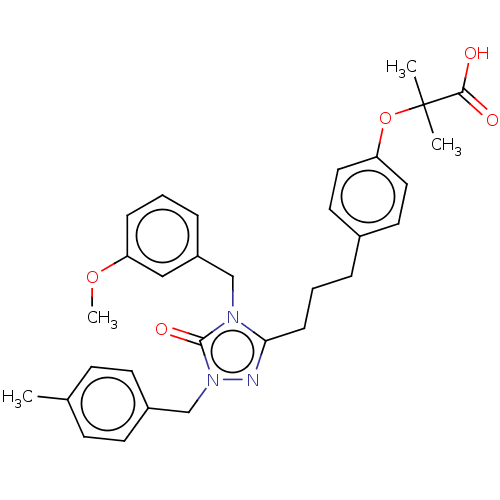
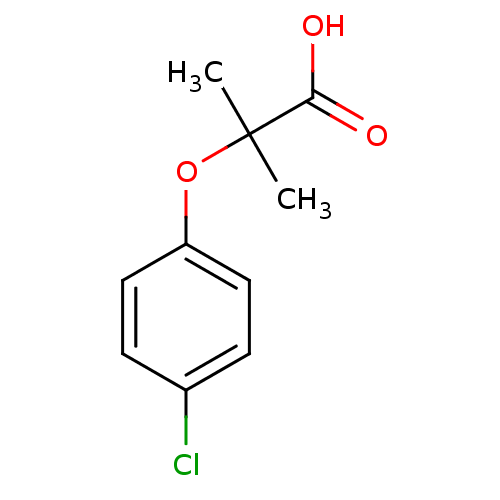
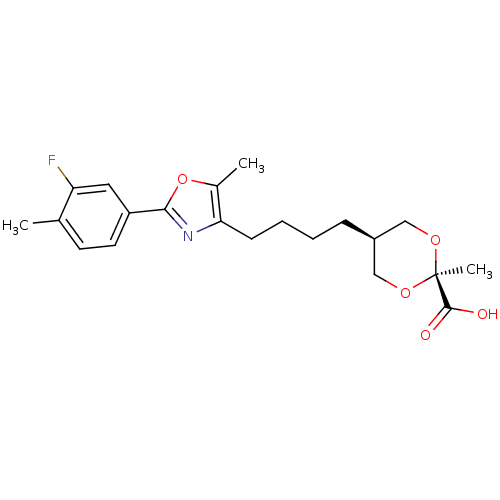
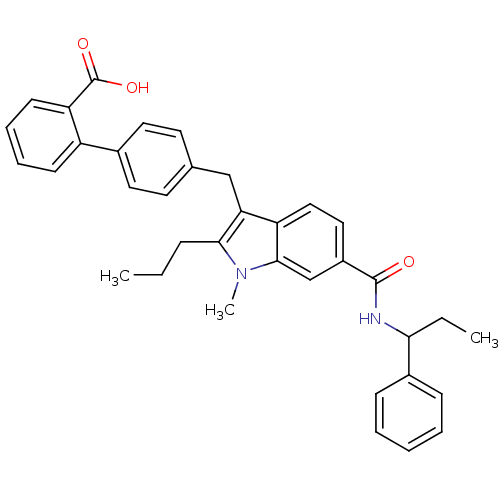
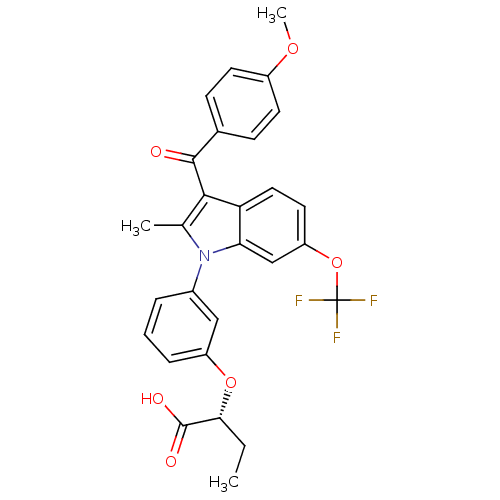
Affinity DataEC50: 0.350nMAssay Description:Effective concentration against murine PPARalpha in transactivation assayMore data for this Ligand-Target Pair
Affinity DataEC50: 1nMAssay Description:Activation of murine PPAR alpha ligand binding domainMore data for this Ligand-Target Pair
Affinity DataEC50: 1.30nMAssay Description:Effective concentration against murine PPARalpha in transactivation assayMore data for this Ligand-Target Pair
Affinity DataEC50: 3nMAssay Description:Agonist activity at Gal4-fused mouse PPARalpha co-expressed with RXRalpha in monkey CV1 cells assessed as PPRE transcriptional activation incubated f...More data for this Ligand-Target Pair
Affinity DataEC50: 3nMAssay Description:Agonist activity at Gal4-fused mouse PPARalpha co-expressed with RXRalpha in monkey CV1 cells assessed as PPRE transcriptional activation incubated f...More data for this Ligand-Target Pair
Affinity DataEC50: 5nMAssay Description:Agonist activity for murine PPAR alpha receptor in transcriptional activation assayMore data for this Ligand-Target Pair
Affinity DataEC50: 5nMAssay Description:Compound was tested for its agonist activity against murine Peroxisome proliferator activated receptor alpha-Gal4 chimeric receptor transfected CV-1 ...More data for this Ligand-Target Pair
Affinity DataEC50: 10nMAssay Description:Agonist activity at Gal4-fused mouse PPARalpha co-expressed with RXRalpha in monkey CV1 cells assessed as PPRE transcriptional activation incubated f...More data for this Ligand-Target Pair
Affinity DataEC50: 10nMAssay Description:Agonist activity at Gal4-fused mouse PPARalphaMore data for this Ligand-Target Pair
Affinity DataEC50: 10nMAssay Description:Agonist activity at Gal4-fused mouse PPARalpha co-expressed with RXRalpha in monkey CV1 cells assessed as PPRE transcriptional activation incubated f...More data for this Ligand-Target Pair
Affinity DataEC50: 10nMAssay Description:Compound was tested for its agonist activity against murine Peroxisome proliferator activated receptor alpha-Gal4 chimeric receptor transfected CV-1 ...More data for this Ligand-Target Pair
Affinity DataEC50: 17nMAssay Description:Agonist activity at mouse PPARalpha transfected in Cos7 cells co-expressing with pBIND and pGL4.35 Gal4 UAS Luc reporter plasmid by measuring lucifer...More data for this Ligand-Target Pair
Affinity DataEC50: 20nMAssay Description:Agonist activity at Gal4-fused mouse PPARalpha co-expressed with RXRalpha in monkey CV1 cells assessed as PPRE transcriptional activation incubated f...More data for this Ligand-Target Pair
Affinity DataEC50: 20nMAssay Description:Agonist activity at Gal4-fused mouse PPARalphaMore data for this Ligand-Target Pair
Affinity DataEC50: 20nMAssay Description:Agonist activity at Gal4-fused mouse PPARalpha co-expressed with RXRalpha in monkey CV1 cells assessed as PPRE transcriptional activation incubated f...More data for this Ligand-Target Pair
Affinity DataEC50: 20nMAssay Description:Agonist activity at Gal4-fused mouse PPARalpha co-expressed with RXRalpha in monkey CV1 cells assessed as PPRE transcriptional activation incubated f...More data for this Ligand-Target Pair
Affinity DataEC50: 28nMAssay Description:Agonist activity at mouse PPARalpha receptor expressed in african green monkey COS1 cells co-transfected with fused yeast Gal4-DBD by transactivation...More data for this Ligand-Target Pair
Affinity DataEC50: 30nMAssay Description:Agonist activity at Gal4-fused mouse PPARalpha co-expressed with RXRalpha in monkey CV1 cells assessed as PPRE transcriptional activation incubated f...More data for this Ligand-Target Pair
Affinity DataEC50: 30nMAssay Description:Agonist activity at mouse PPARalpha expressed in CV1 cells by transactivation assayMore data for this Ligand-Target Pair
Affinity DataEC50: 30nMAssay Description:Agonist activity at mouse PPARalpha expressed in CV1 cells by transactivation assayMore data for this Ligand-Target Pair
Affinity DataEC50: 33nMAssay Description:Compound was tested for its agonist activity against murine Peroxisome proliferator activated receptor alpha-Gal4 chimeric receptor transfected CV-1 ...More data for this Ligand-Target Pair
Affinity DataEC50: 40nMAssay Description:Effective concentration against murine PPARalpha in transactivation assayMore data for this Ligand-Target Pair
Affinity DataEC50: 40nMAssay Description:Agonist activity at mouse PPARalpha receptor expressed in african green monkey COS1 cells co-transfected with fused yeast Gal4-DBD by transactivation...More data for this Ligand-Target Pair
Affinity DataEC50: 42nMAssay Description:Effective concentration against murine PPARalpha in transactivation assayMore data for this Ligand-Target Pair
Affinity DataEC50: 53nMAssay Description:Compound was tested for its agonist activity against murine Peroxisome proliferator activated receptor alpha-Gal4 chimeric receptor transfected CV-1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 80nMAssay Description:Activation of mouse liver PPARalphaMore data for this Ligand-Target Pair
Affinity DataEC50: 91nMAssay Description:Transactivation activity for mouse Peroxisome proliferator activated receptor alphaMore data for this Ligand-Target Pair
Affinity DataEC50: 100nMAssay Description:Agonist activity at mouse PPARalpha expressed in CV1 cells by transactivation assayMore data for this Ligand-Target Pair
Affinity DataKi: 100nMAssay Description:Binding affinity towards peroxisome proliferator activated receptor alpha (murinePPAR alpha)More data for this Ligand-Target Pair
Affinity DataEC50: 100nMAssay Description:Agonist activity at mouse PPARalpha by transactivation assayMore data for this Ligand-Target Pair
Affinity DataEC50: 110nMAssay Description:Agonist activity at mouse PPARalpha receptor expressed in african green monkey COS1 cells co-transfected with fused yeast Gal4-DBD by transactivation...More data for this Ligand-Target Pair
Affinity DataEC50: 120nMAssay Description:Agonist activity at mouse PPARalpha receptor expressed in african green monkey COS1 cells co-transfected with fused yeast Gal4-DBD by transactivation...More data for this Ligand-Target Pair
Affinity DataEC50: 120nMAssay Description:Compound was tested for its agonist activity against murine Peroxisome proliferator activated receptor alpha-Gal4 chimeric receptor transfected CV-1 ...More data for this Ligand-Target Pair
Affinity DataEC50: 144nMAssay Description:Agonist activity at mouse PPAR-alphaMore data for this Ligand-Target Pair
Affinity DataIC50: 146nMAssay Description:In vitro inhibition of mouse Peroxisome proliferator activated receptor alphaMore data for this Ligand-Target Pair
Affinity DataEC50: 150nMAssay Description:Agonist activity at mouse PPARalpha receptor expressed in african green monkey COS1 cells co-transfected with fused yeast Gal4-DBD by transactivation...More data for this Ligand-Target Pair
Affinity DataEC50: 151nMAssay Description:Transactivation activity for mouse Peroxisome proliferator activated receptor alphaMore data for this Ligand-Target Pair
Affinity DataEC50: 170nMAssay Description:Agonist activity at mouse PPARalpha expressed in CV-1 cells by reporter gene assayMore data for this Ligand-Target Pair
Affinity DataEC50: 173nMAssay Description:Transactivation activity for mouse Peroxisome proliferator activated receptor alphaMore data for this Ligand-Target Pair
Affinity DataEC50: 180nMAssay Description:Agonist activity against mouse Peroxisome proliferator activated receptor alphaMore data for this Ligand-Target Pair
Affinity DataEC50: 211nMAssay Description:Transactivation activity for mouse Peroxisome proliferator activated receptor alphaMore data for this Ligand-Target Pair
Affinity DataEC50: 240nMAssay Description:Agonist activity at mouse PPARalphaMore data for this Ligand-Target Pair
Affinity DataEC50: 270nMAssay Description:Agonist activity for murine PPAR alpha receptor in transcriptional activation assayMore data for this Ligand-Target Pair
Affinity DataEC50: 350nMAssay Description:Agonist activity at mouse PPARalpha receptor expressed in african green monkey COS1 cells co-transfected with fused yeast Gal4-DBD by transactivation...More data for this Ligand-Target Pair
Affinity DataEC50: 360nMAssay Description:Agonist activity at mouse PPARalpha receptor expressed in african green monkey COS1 cells co-transfected with fused yeast Gal4-DBD by transactivation...More data for this Ligand-Target Pair
Affinity DataEC50: 366nMAssay Description:Transactivation of mouse GAL4-fused PPARalpha LBD expressed in African green monkey COS7 cells co-expressing 5Gal4 pGL3 TK Luc after overnight incuba...More data for this Ligand-Target Pair
Affinity DataIC50: 404nMAssay Description:In vitro inhibition of mouse Peroxisome proliferator activated receptor alphaMore data for this Ligand-Target Pair
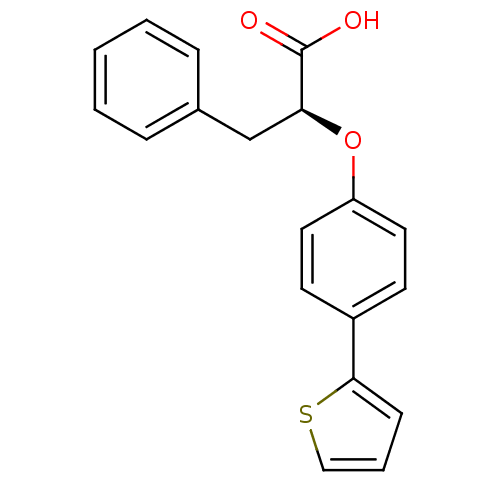

 3D Structure (crystal)
3D Structure (crystal)