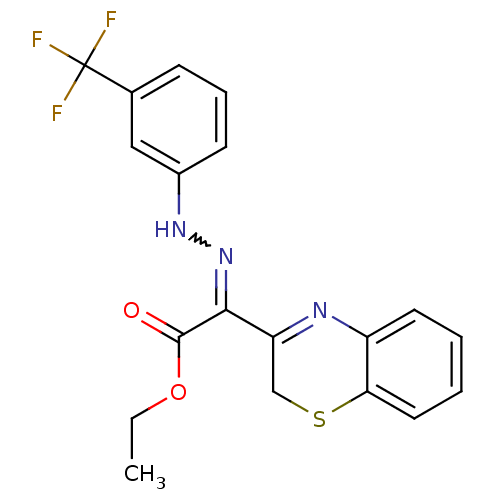
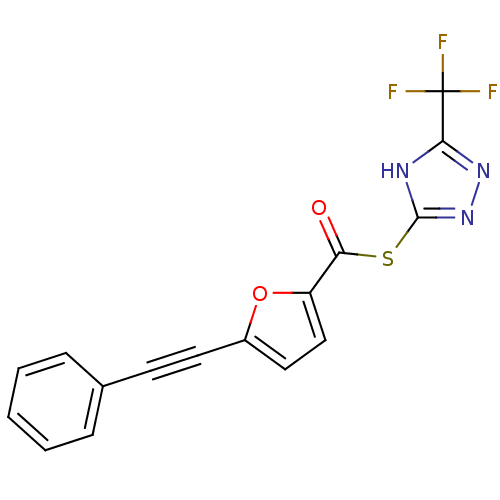
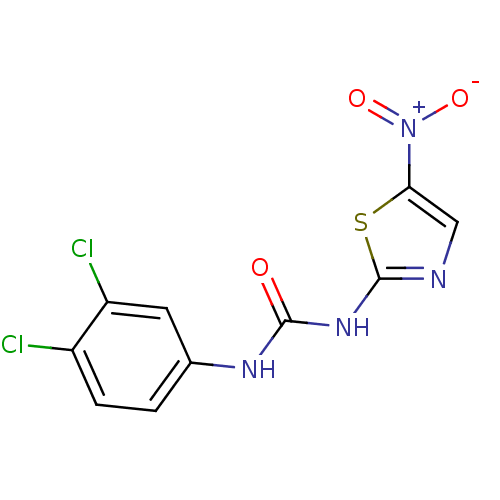
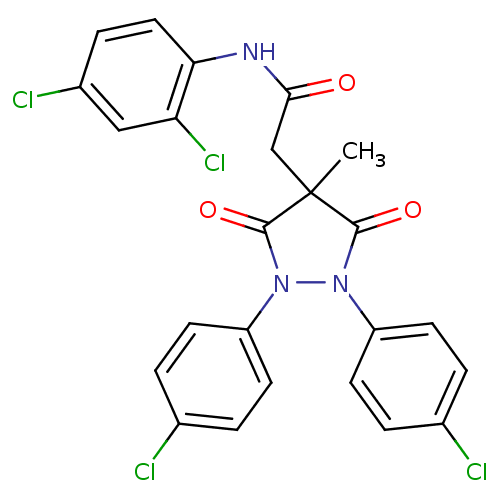
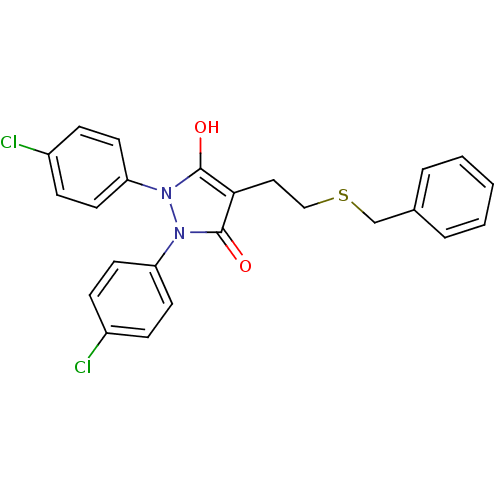
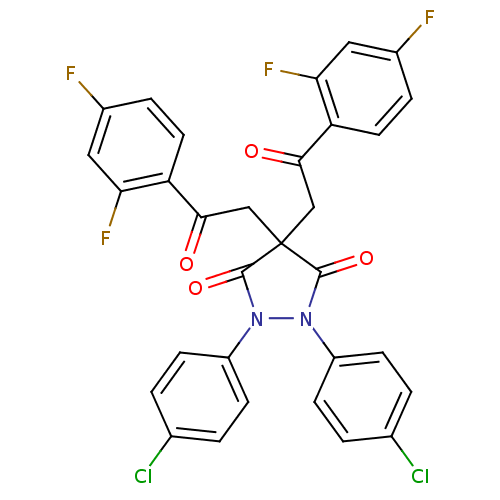
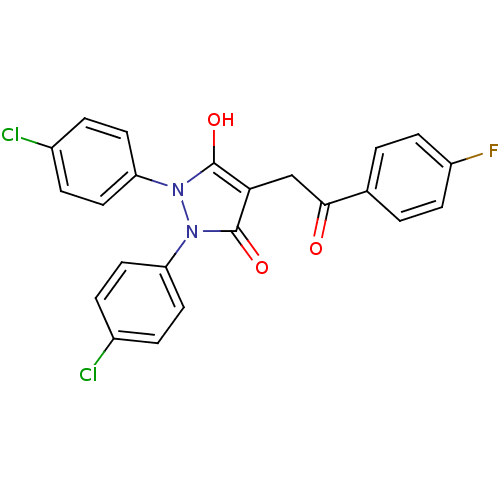
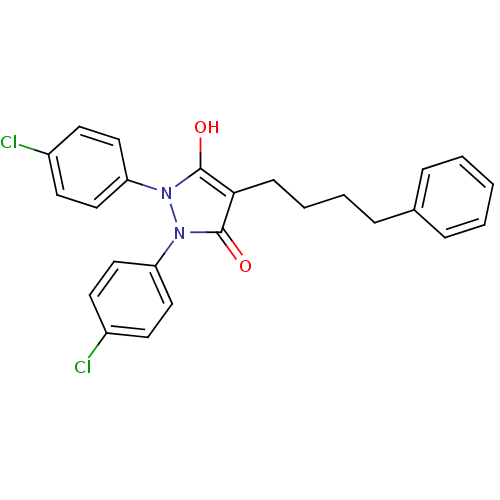
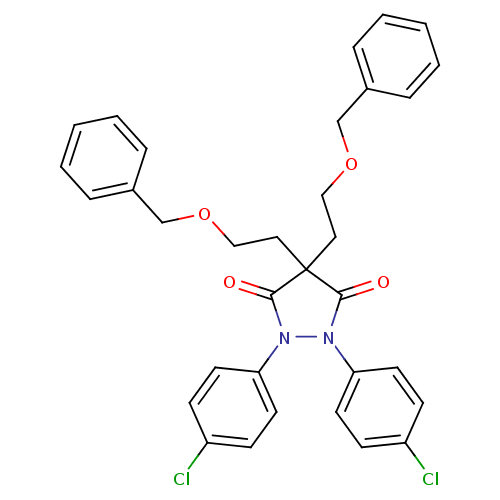
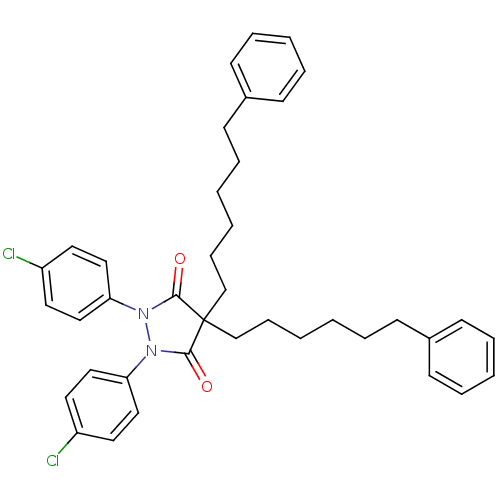
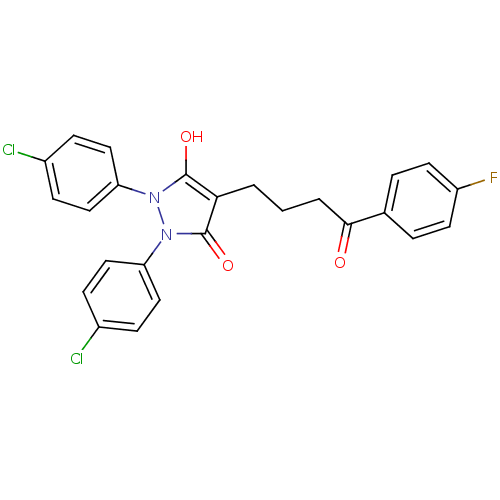
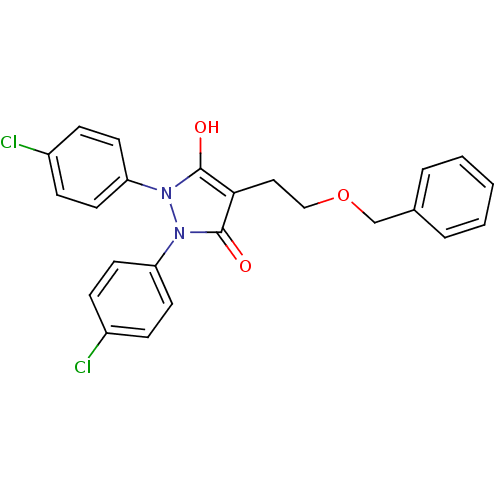
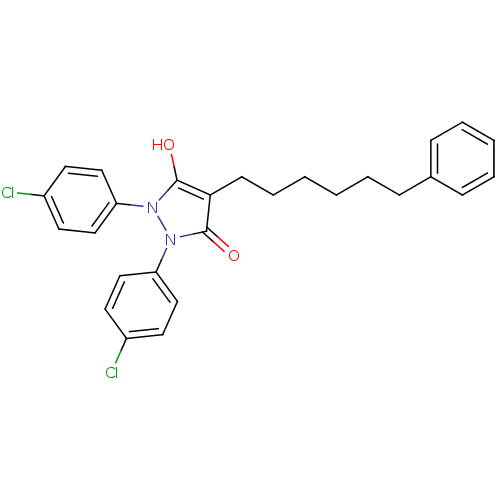
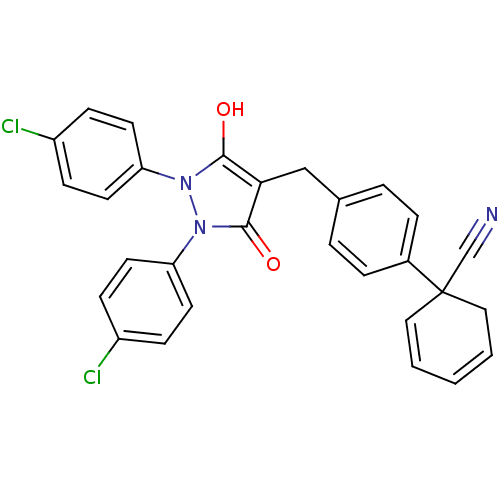
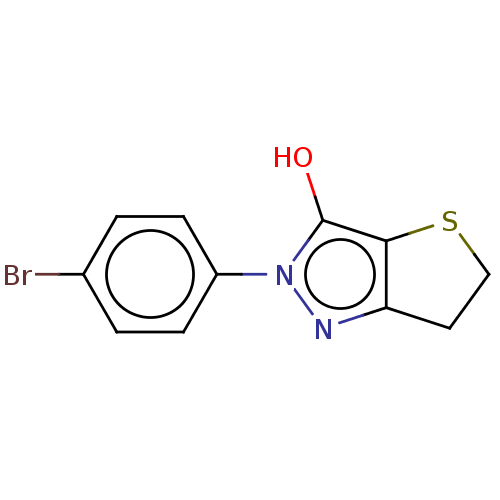
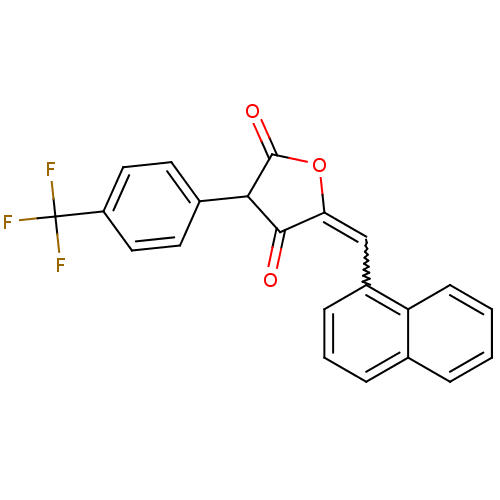
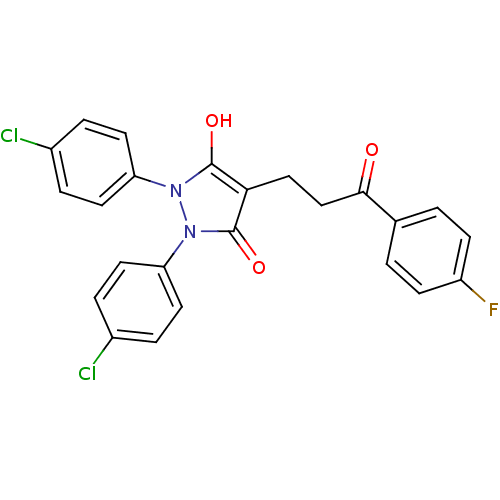
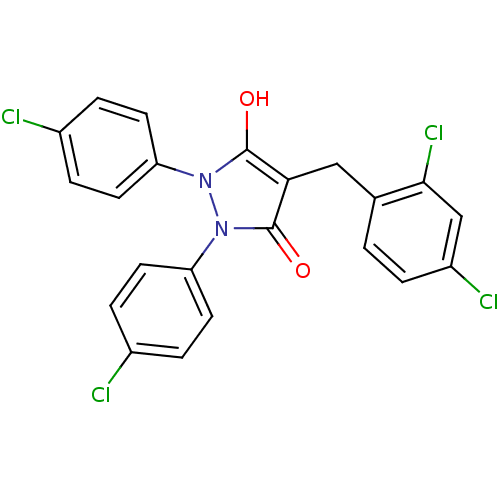
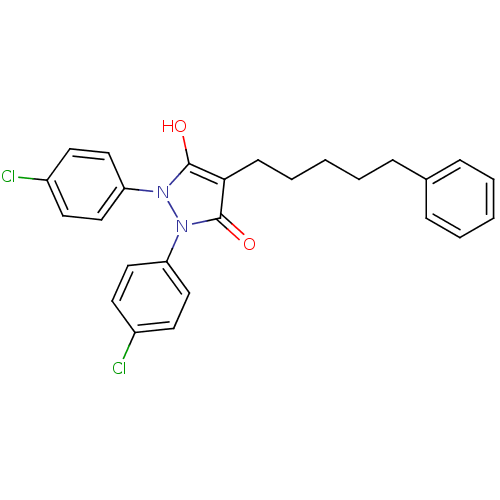
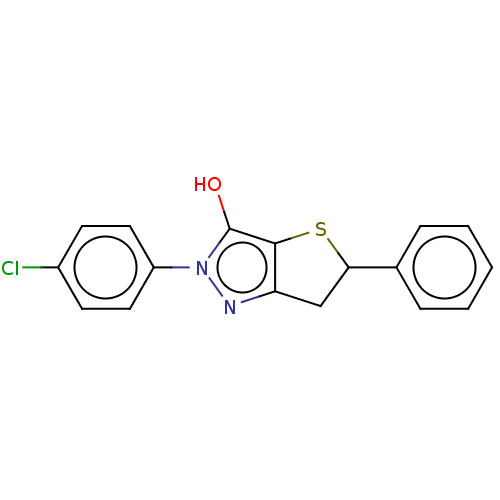
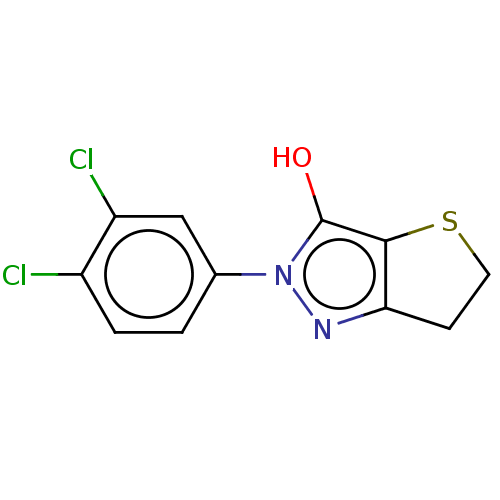
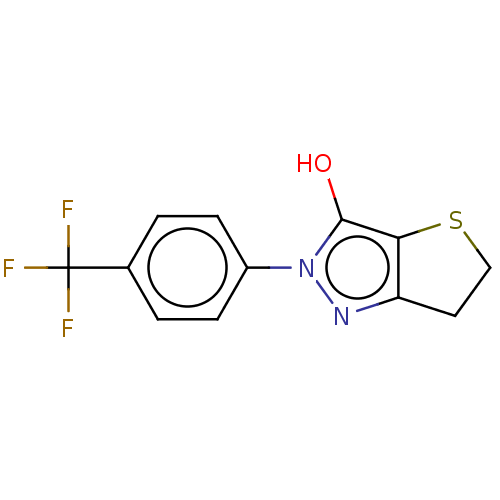
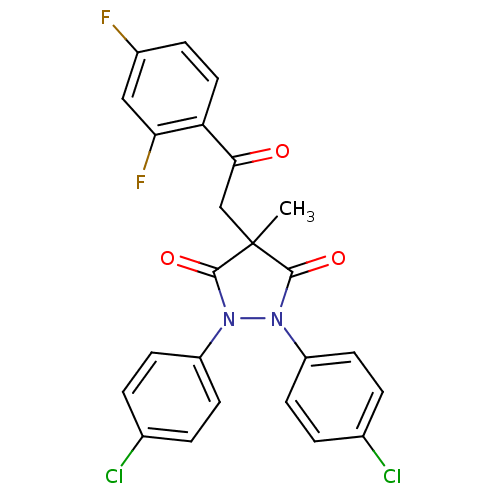
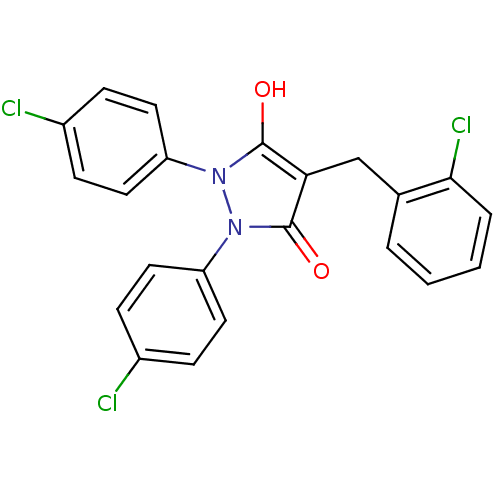
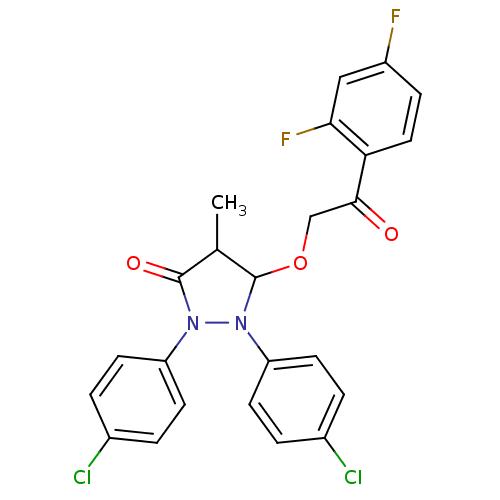
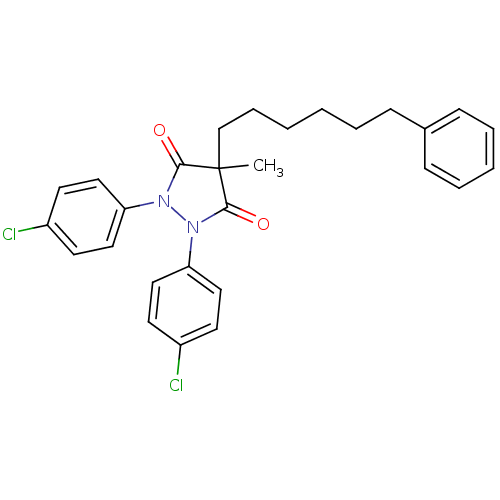
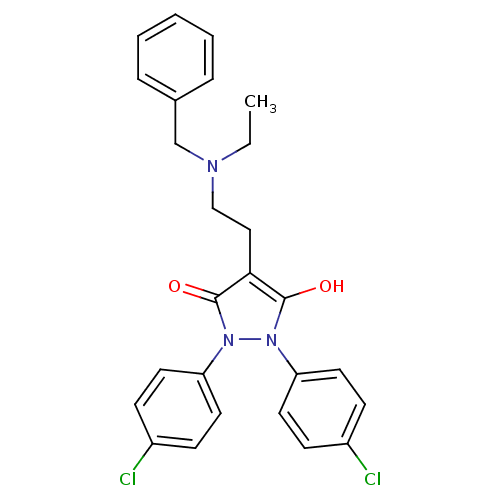
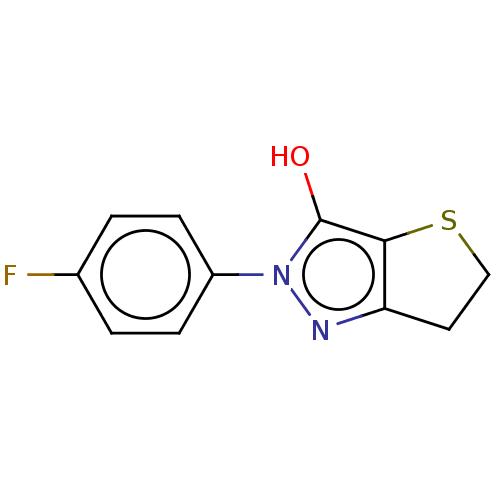
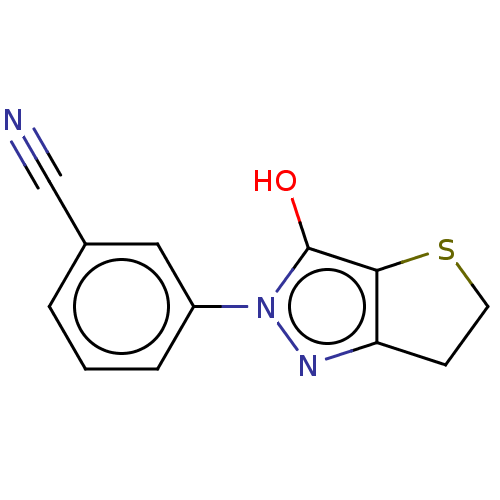
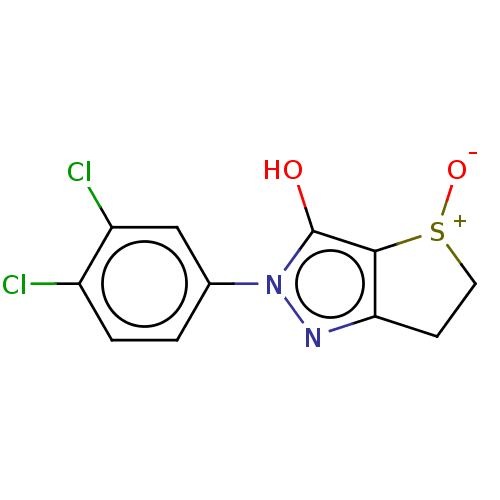
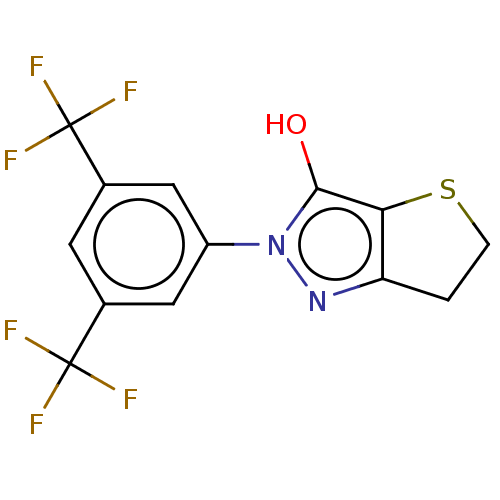
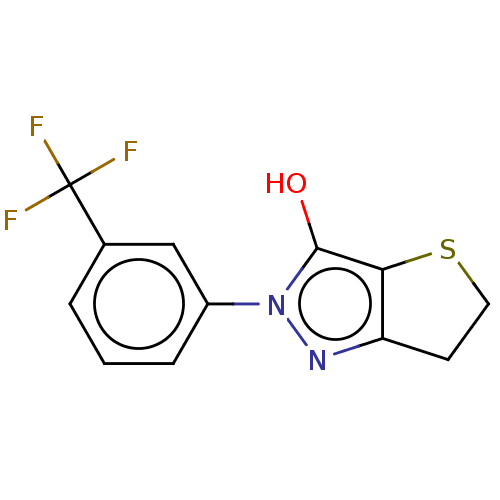
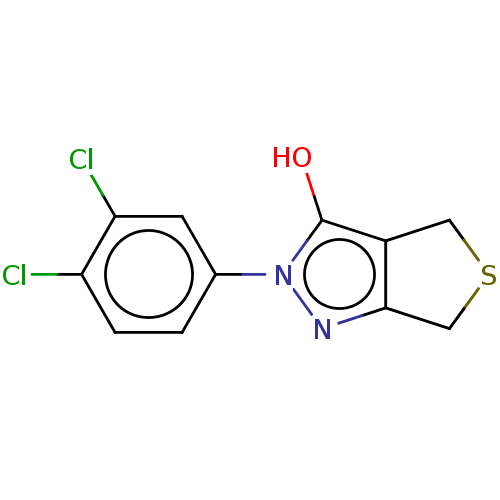
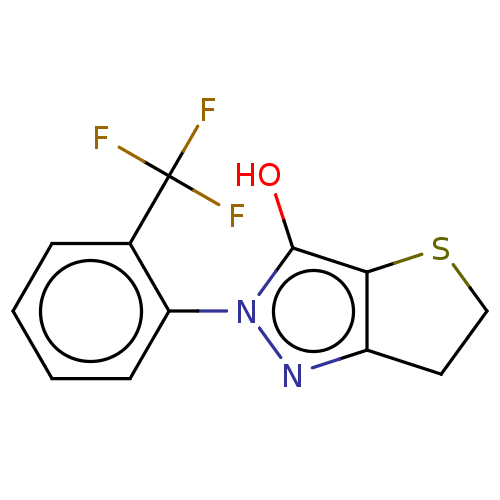
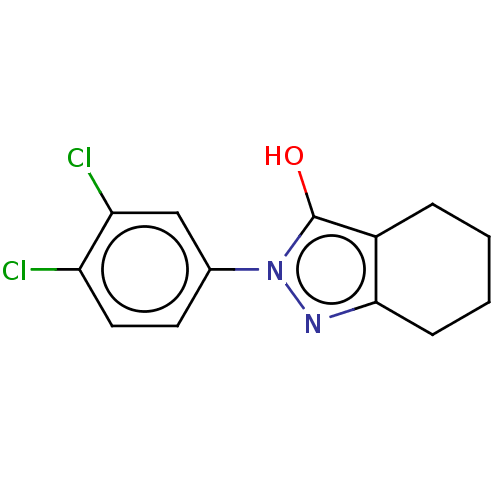
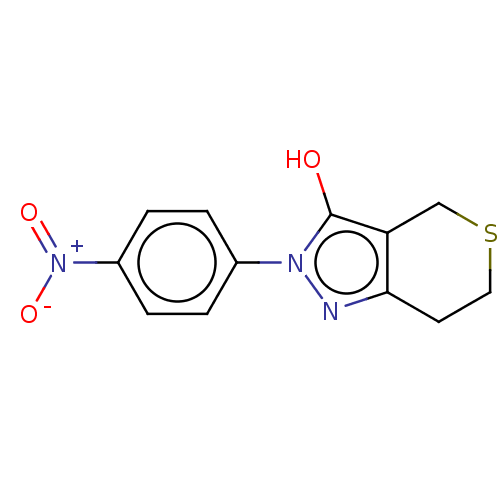
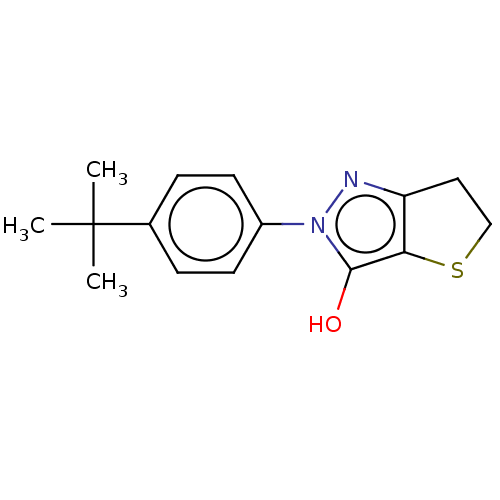
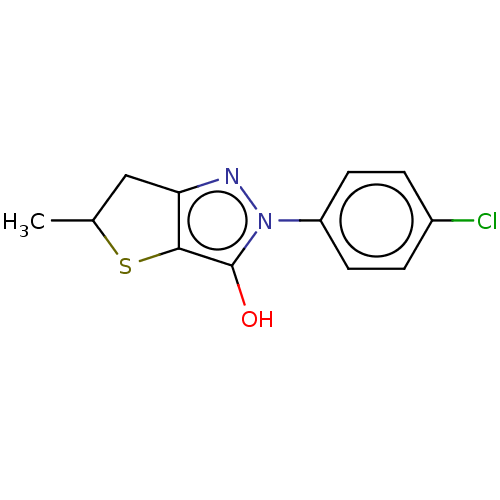
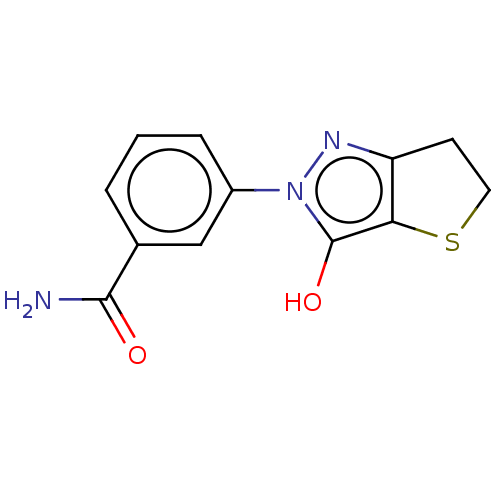
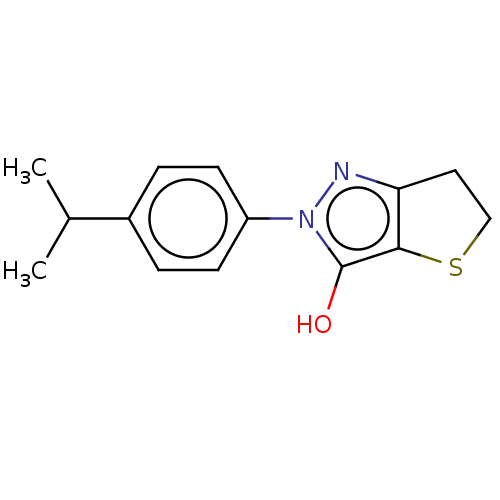
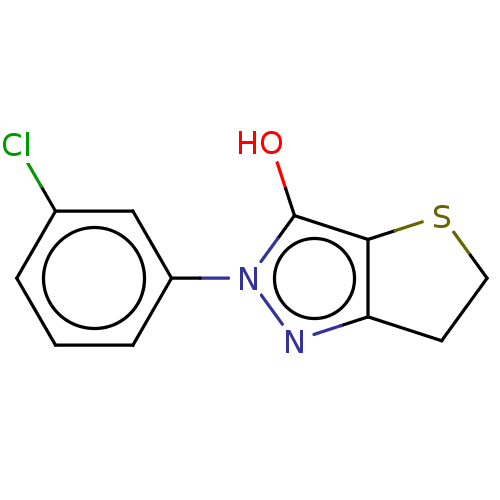
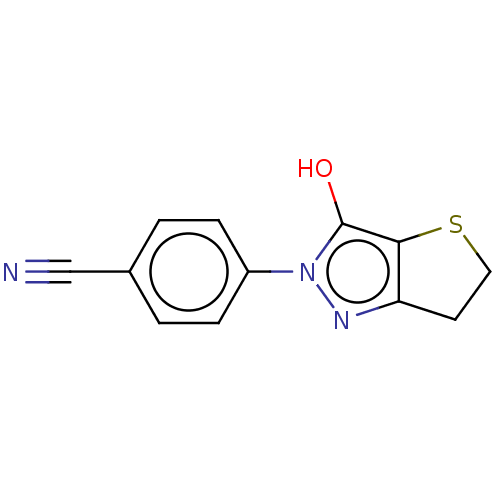
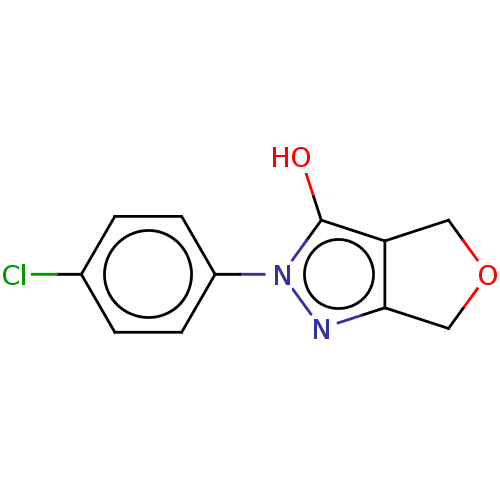
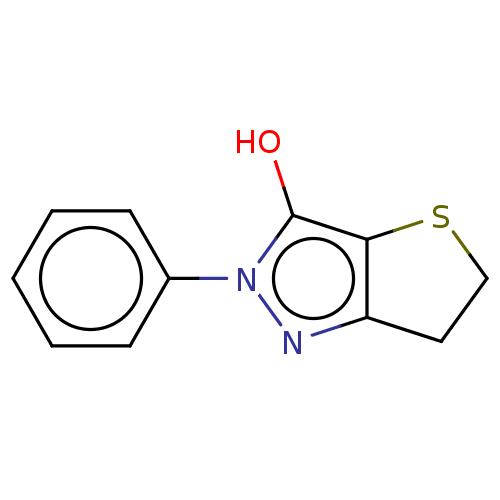
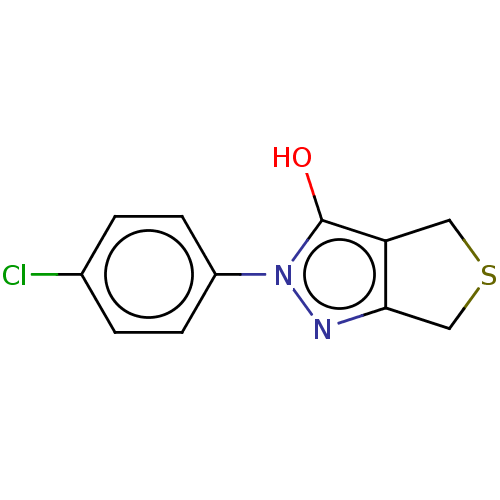
Affinity DataIC50: 5.10E+3nMAssay Description:Inhibitory concentration against MurB enzyme in Staphylococcus aureusMore data for this Ligand-Target Pair
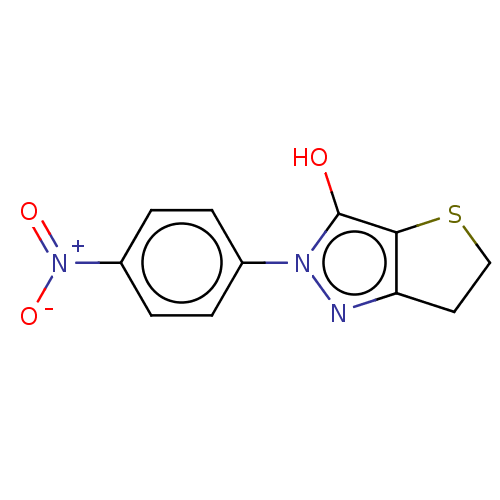
Affinity DataIC50: 5.20E+3nMAssay Description:Inhibitory concentration against MurB enzyme in Staphylococcus aureusMore data for this Ligand-Target Pair
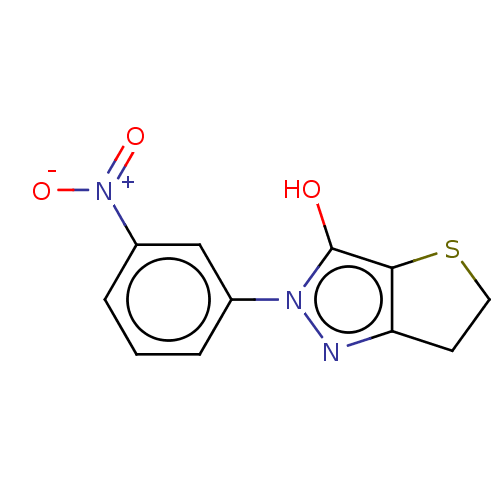
Affinity DataIC50: 5.20E+3nMAssay Description:Inhibitory concentration against MurB enzyme in Staphylococcus aureusMore data for this Ligand-Target Pair
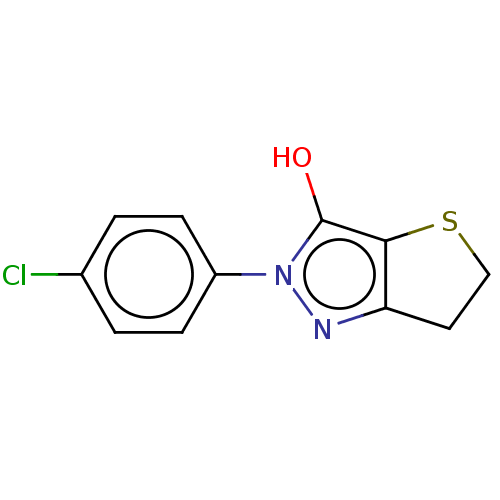
Affinity DataIC50: 6.80E+3nMAssay Description:Inhibitory concentration against MurB enzyme in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 7.50E+3nMAssay Description:Inhibitory concentration against MurB enzyme in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 8.20E+3nMAssay Description:Inhibitory concentration against MurB enzyme in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+4nMAssay Description:Inhibitory concentration against MurB enzyme in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+4nMAssay Description:Inhibitory concentration against MurB enzyme in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibitory concentration against MurB enzyme in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibitory concentration against MurB enzyme in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 1.21E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+4nMAssay Description:Inhibitory concentration against MurB enzyme in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 2.12E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 2.35E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+4nMAssay Description:Inhibitory concentration against MurB enzyme in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+4nMAssay Description:Inhibitory concentration against MurB enzyme in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 3.34E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 3.83E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 3.84E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibitory concentration against MurB enzyme in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibitory concentration against MurB enzyme in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibitory concentration against MurB enzyme in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibitory concentration against MurB enzyme in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibitory concentration against MurB enzyme in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 5.07E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 5.75E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 6.59E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 7.05E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 7.33E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 8.70E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 8.73E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 8.82E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 9.01E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 9.11E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 9.37E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 9.56E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 9.60E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 9.89E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.02E+5nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.05E+5nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.14E+5nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 8.30E+5nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramate dehydrogenaseMore data for this Ligand-Target Pair