Report error Found 1269 with Last Name = 'huang' and Initial = 'a'
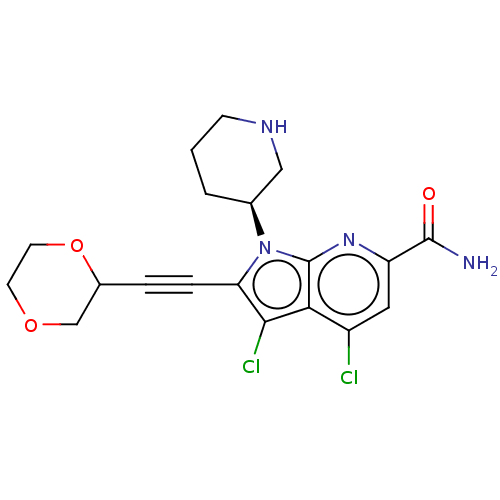
Affinity DataKi: 0.100nMAssay Description:Inhibition of PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser-112 residue by TR-FRET assayMore data for this Ligand-Target Pair
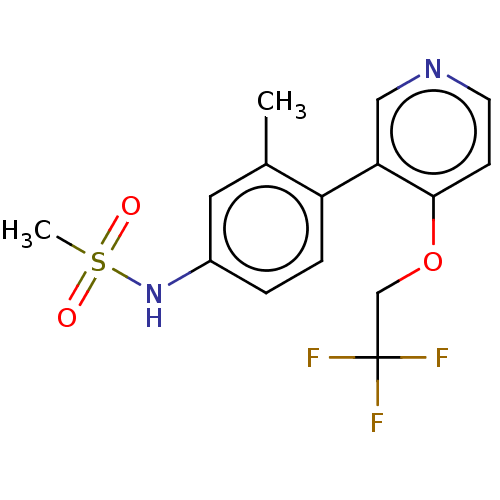
Affinity DataKi: 0.400nMAssay Description:Inhibition of PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser-112 residue by TR-FRET assayMore data for this Ligand-Target Pair
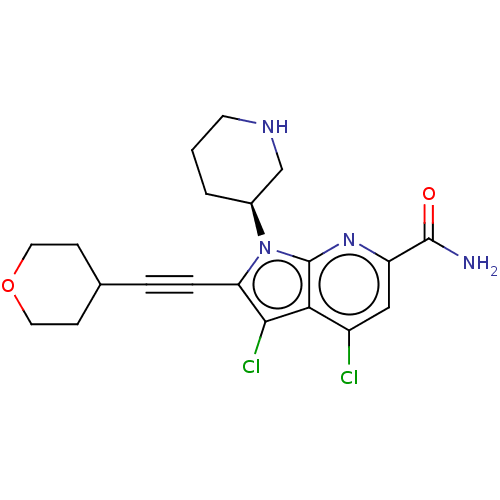
Affinity DataKi: 0.600nMAssay Description:Inhibition of PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser-112 residue by TR-FRET assayMore data for this Ligand-Target Pair
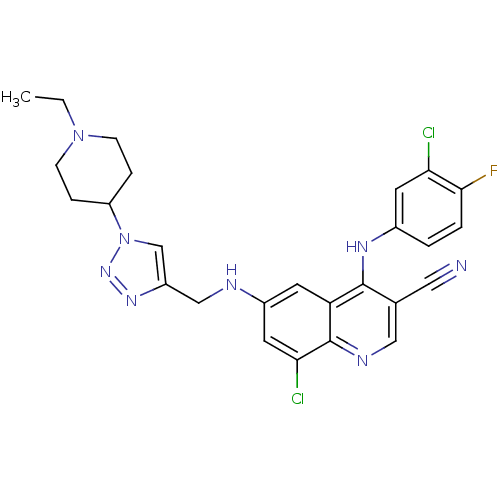
Affinity DataKi: 0.700nMAssay Description:Inhibition of PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser-112 residue by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.900nMAssay Description:Inhibition of PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser-112 residue by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Inhibition of PGD2-induced CRTH2 receptor internalization of CD16 negative granulocytes in human whole blood by flow cytometryMore data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Inhibition of PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser-112 residue by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Inhibition of PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser-112 residue by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Inhibition of PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser-112 residue by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Inhibition of PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser-112 residue by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 53nM ΔG°: -41.5kJ/molepH: 7.5 T: 25°CAssay Description:The effects of compound on enzyme activity were measured by using peptide cleavage assay. Cleavage products were resolved and analyzed with a reverse...More data for this Ligand-Target Pair
Affinity DataKi: 58nM ΔG°: -41.3kJ/molepH: 7.5 T: 25°CAssay Description:The effects of compound on enzyme activity were measured by using peptide cleavage assay. Cleavage products were resolved and analyzed with a reverse...More data for this Ligand-Target Pair
Affinity DataKi: 148nMAssay Description:Antagonist activity at DP receptor in human platelets assessed as inhibition of PGD2-induced cAMP production by competitive ELISAMore data for this Ligand-Target Pair
Affinity DataKi: 660nM ΔG°: -35.3kJ/molepH: 7.5 T: 25°CAssay Description:The effects of compound on enzyme activity were measured by using peptide cleavage assay. Cleavage products were resolved and analyzed with a reverse...More data for this Ligand-Target Pair
Affinity DataKi: 2.26E+3nM ΔG°: -32.2kJ/molepH: 7.5 T: 25°CAssay Description:The effects of compound on enzyme activity were measured by using peptide cleavage assay. Cleavage products were resolved and analyzed with a reverse...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nM ΔG°: >-28.5kJ/molepH: 7.5 T: 25°CAssay Description:The effects of compound on enzyme activity were measured by using peptide cleavage assay. Cleavage products were resolved and analyzed with a reverse...More data for this Ligand-Target Pair
Affinity DataKi: 4.40E+4nMAssay Description:Inhibition of EP2 expressed in HEK293 cells assessed as inhibition of PGE2-induced cAMP productionMore data for this Ligand-Target Pair
Affinity DataKi: 4.40E+4nMAssay Description:Inhibition of EP4 expressed in HEK293 cells assessed as inhibition of PGE2-induced cAMP productionMore data for this Ligand-Target Pair
Affinity DataIC50: 0.25nMAssay Description:Inhibition of mTOR (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 0.600nMAssay Description:Inhibition of PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser-112 residue by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.700nMAssay Description:Inhibition of PIM1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 0.700nMAssay Description:Human IDO1/HEK293 cells were seeded at 10,000 cells per 50 uL per well with RPMI/phenol red free media contains 10% FBS in a 384-well black wall clea...More data for this Ligand-Target Pair
Affinity DataIC50: 0.700nMAssay Description:Inhibition of BTK (unknown origin) by ADP-gloassayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.700nMAssay Description:Inhibition of PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser-112 residue by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.800nMAssay Description:Antagonist activity at human recombinant CRTh2 expressed in CHO-K1 cells by cAMP TR FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.800nMAssay Description:Displacement of [3H]PGD2 from recombinant human CRTH2 receptor expressed in CHO-K1 cells after 30 min by FRET methodMore data for this Ligand-Target Pair
Affinity DataIC50: 0.800nMAssay Description:Binding affinity to human CRTH2 receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 0.800nMAssay Description:Inhibition of PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser-112 residue by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: <1nMpH: 7.4 T: 22°CAssay Description:Compounds were tested for their ability to inhibit the cleavage of the substrate by the purified enzyme in a fluorescence-based fluorescence resonanc...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser-112 residue by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Binding affinity against Melatonin receptor using ovine pars tuberalis membranes of the pituitary.More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of recombinant full length human N-terminal His tagged BTK expressed in baculovirus infected Sf9 cells using poly (4:1 Glu, Tyr) as substr...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of PIM1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of PIM1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Human IDO1/HEK293 cells were seeded at 10,000 cells per 50 uL per well with RPMI/phenol red free media contains 10% FBS in a 384-well black wall clea...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Antagonist activity at human recombinant CRTh2 expressed in CHO-K1 cells by cAMP TR FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of PIM1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.10nMAssay Description:Inhibition of PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser-112 residue by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10nMAssay Description:The assays were performed in U-bottom 384-well optiplates. The final assay volume was 15 μl prepared from 7.5 μl additions of microsomes (p...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10nMAssay Description:Inhibition of human ERGMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10nMpH: 7.2 T: 25°CAssay Description:The assays were performed in U-bottom 384-well optiplates. The final assay volume was 15 μl prepared from 7.5 μl additions of microsomes (p...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20nMAssay Description:Inhibition of CRTH2 receptor-mediated chemotaxis in human basophilsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20nMAssay Description:Inhibition of CRTH2 receptor-mediated chemotaxis in human basophilsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Inhibition of human ERGMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Binding affinity to human CRTH2 receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Inhibition of PD-1-Ig/His-PD-L1 (unknown origin) protein-protein interaction incubated with PD-L1 for 15 mins followed by addition of PD-1 for 15 min...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMpH: 7.2 T: 4°CAssay Description:The assays were performed in U-bottom 384-well optiplates. The final assay volume was 15 ul prepared from 7.5 ul additions of microsomes (prepared as...More data for this Ligand-Target Pair
Affinity DataIC50: 1.5nMAssay Description:Inhibition of PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser-112 residue by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMAssay Description:Inhibition of PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser-112 residue by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues 217 and 221 of GST-MEK1 was detected by ...More data for this Ligand-Target Pair
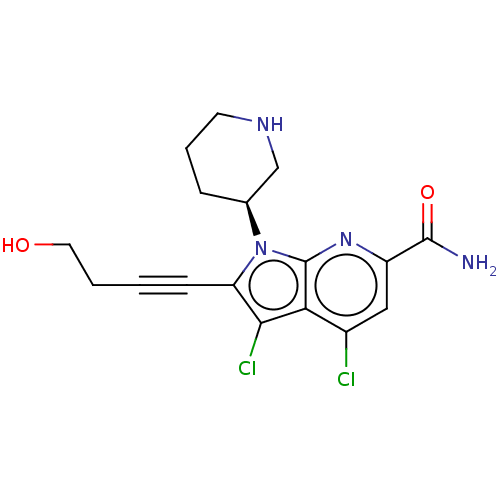
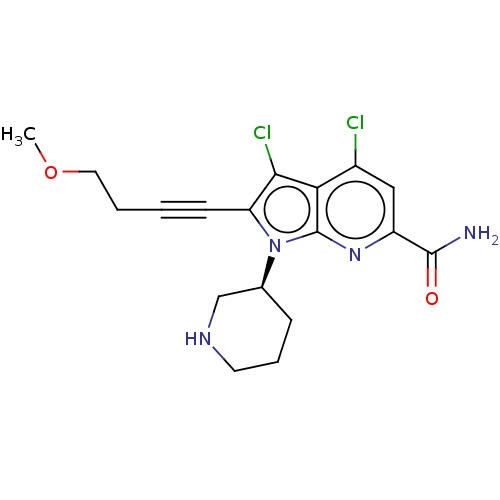
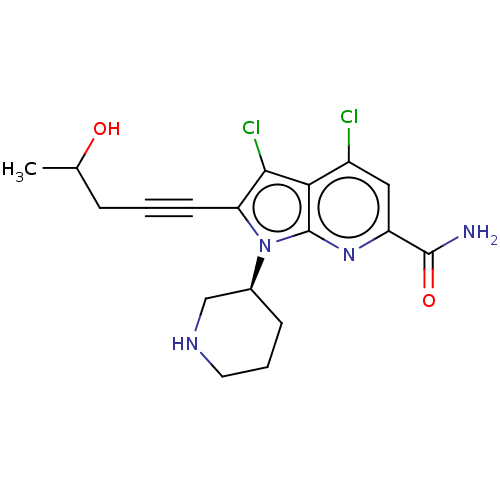
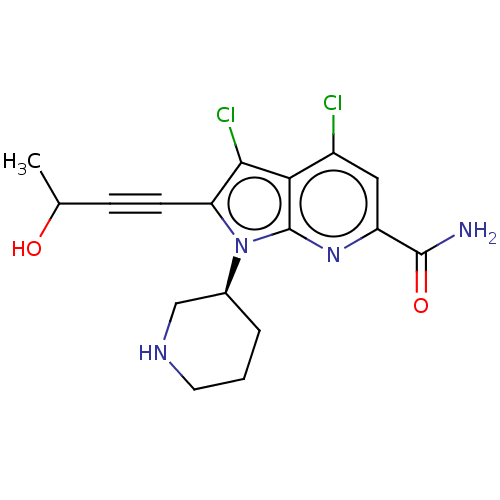

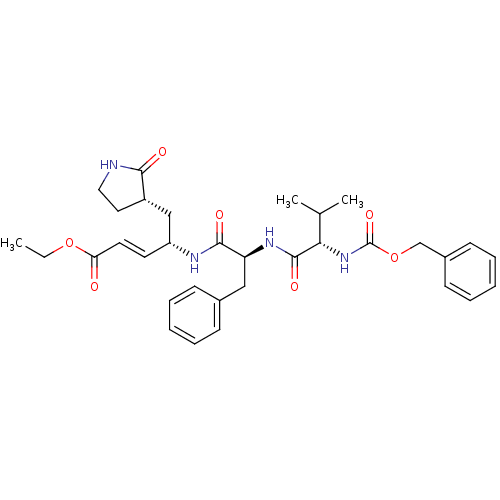
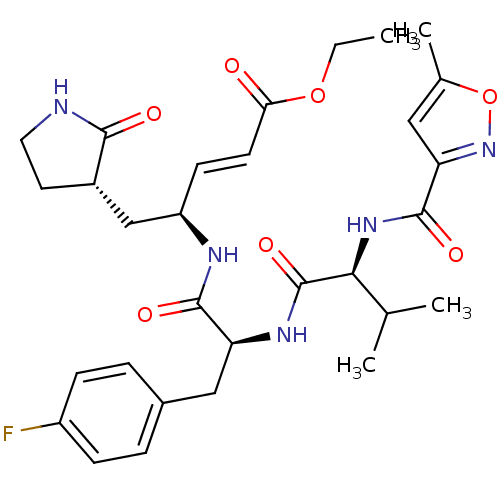

 3D Structure (crystal)
3D Structure (crystal)