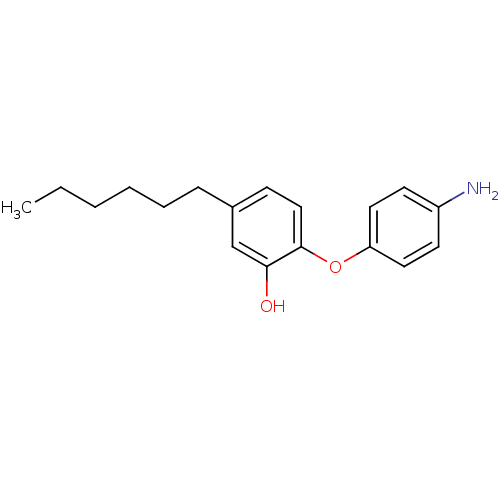
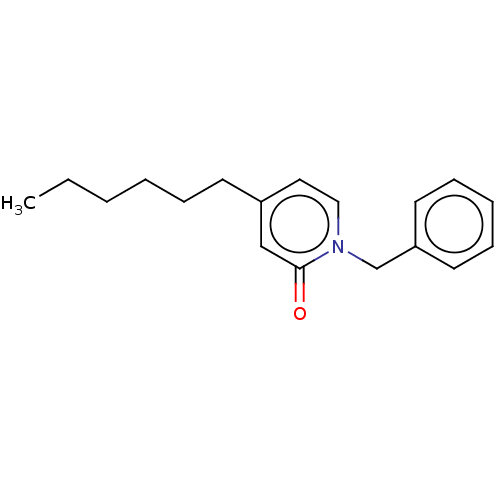
Report error Found 75 with Last Name = 'daryaee' and Initial = 'f'
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 0.510nM ΔG°: -53.0kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 1nM ΔG°: -51.4kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 3nM ΔG°: -48.6kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 4nM ΔG°: -47.9kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 15nM ΔG°: -44.7kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 25nM ΔG°: -43.4kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 31nM ΔG°: -42.9kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 32nM ΔG°: -42.8kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 32nM ΔG°: -42.8kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 32nM ΔG°: -42.8kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 44nM ΔG°: -42.0kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 48nM ΔG°: -41.8kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 55nM ΔG°: -41.4kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 55nM ΔG°: -41.4kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 58nM ΔG°: -41.3kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
Affinity DataKi: 100nM ΔG°: -40.0kJ/molepH: 8.0 T: 25°CAssay Description:Steady-state kinetics were performed on a Cary 100 Bio (Varian) spectrometer at 25 °C using 30 mM PIPES, 150 mM NaCl, and 1.0 mM EDTA (pH 8.0) as...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 105nM ΔG°: -39.8kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 130nM ΔG°: -39.3kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 138nM ΔG°: -39.2kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 153nM ΔG°: -38.9kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 156nM ΔG°: -38.9kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 158nM ΔG°: -38.8kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
Affinity DataKi: 200nM ΔG°: -38.2kJ/molepH: 8.0 T: 25°CAssay Description:Steady-state kinetics were performed on a Cary 100 Bio (Varian) spectrometer at 25 °C using 30 mM PIPES, 150 mM NaCl, and 1.0 mM EDTA (pH 8.0) as...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 405nM ΔG°: -36.5kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
Affinity DataKi: 500nM ΔG°: -36.0kJ/molepH: 8.0 T: 25°CAssay Description:Steady-state kinetics were performed on a Cary 100 Bio (Varian) spectrometer at 25 °C using 30 mM PIPES, 150 mM NaCl, and 1.0 mM EDTA (pH 8.0) as...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Burkholderia pseudomallei)
Stony Brook University
Stony Brook University
Affinity DataKi: 1.13E+3nM ΔG°: -33.9kJ/molepH: 8.0 T: 25°CAssay Description:Slow-onset inhibition kinetics were monitored at 340 nm on a Cary 100 spectrophotometer (Varian) at 25 �C in 30 mM PIPES buffer (pH 8.0) containing 1...More data for this Ligand-Target Pair
Affinity DataKi: 3.00E+3nM ΔG°: -31.5kJ/molepH: 8.0 T: 25°CAssay Description:Steady-state kinetics were performed on a Cary 100 Bio (Varian) spectrometer at 25 °C using 30 mM PIPES, 150 mM NaCl, and 1.0 mM EDTA (pH 8.0) as...More data for this Ligand-Target Pair
Affinity DataKi: 4.00E+3nM ΔG°: -30.8kJ/molepH: 8.0 T: 25°CAssay Description:Steady-state kinetics were performed on a Cary 100 Bio (Varian) spectrometer at 25 °C using 30 mM PIPES, 150 mM NaCl, and 1.0 mM EDTA (pH 8.0) as...More data for this Ligand-Target Pair
Affinity DataKi: 8.00E+3nM ΔG°: -29.1kJ/molepH: 8.0 T: 25°CAssay Description:Steady-state kinetics were performed on a Cary 100 Bio (Varian) spectrometer at 25 °C using 30 mM PIPES, 150 mM NaCl, and 1.0 mM EDTA (pH 8.0) as...More data for this Ligand-Target Pair
Affinity DataKi: 8.00E+3nM ΔG°: -29.1kJ/molepH: 8.0 T: 25°CAssay Description:Steady-state kinetics were performed on a Cary 100 Bio (Varian) spectrometer at 25 °C using 30 mM PIPES, 150 mM NaCl, and 1.0 mM EDTA (pH 8.0) as...More data for this Ligand-Target Pair
Affinity DataKi: 1.60E+4nM ΔG°: -27.4kJ/molepH: 8.0 T: 25°CAssay Description:Steady-state kinetics were performed on a Cary 100 Bio (Varian) spectrometer at 25 °C using 30 mM PIPES, 150 mM NaCl, and 1.0 mM EDTA (pH 8.0) as...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
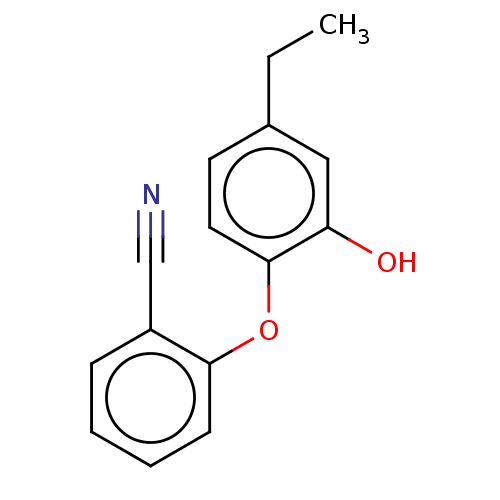
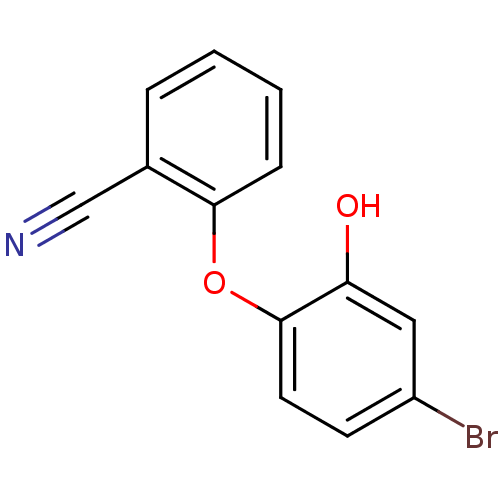
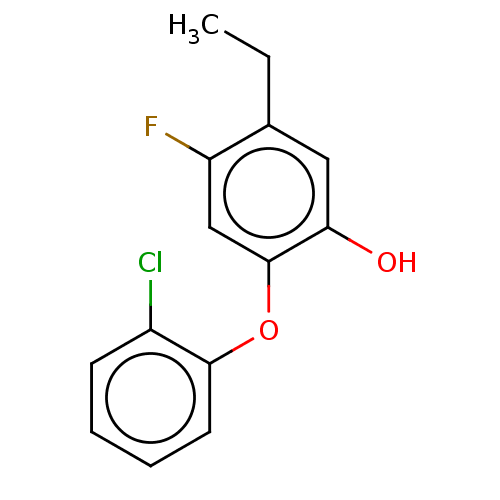
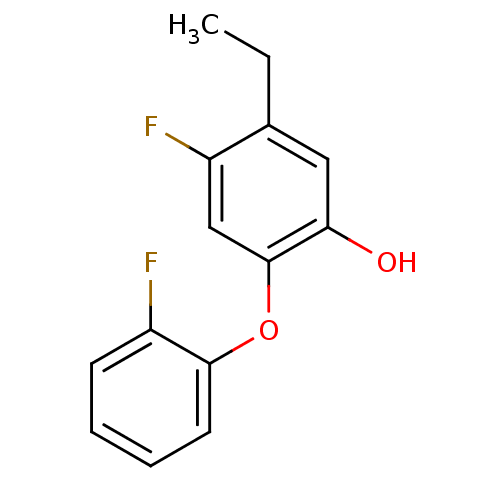

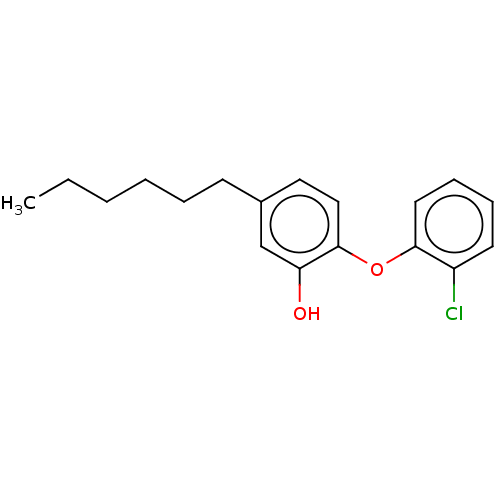
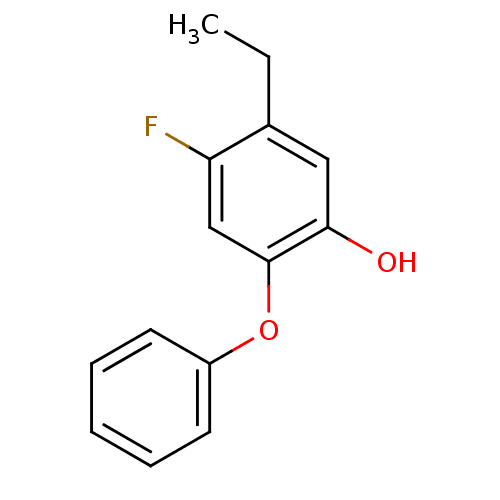
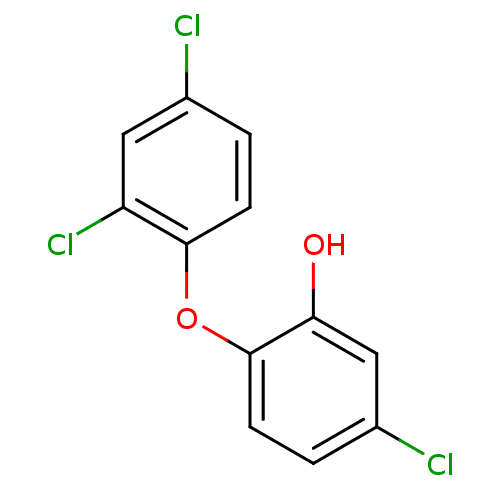
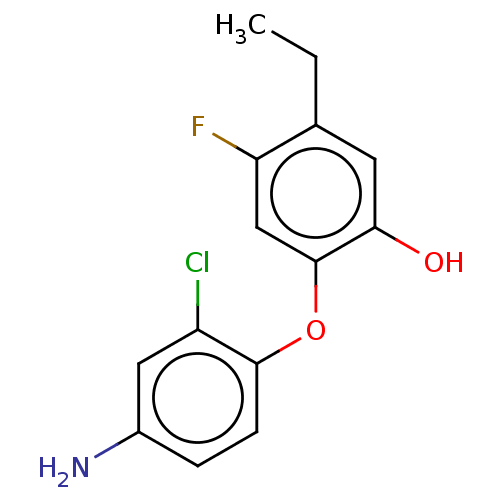
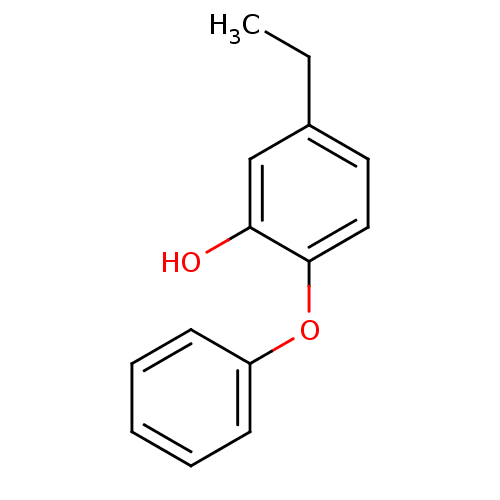
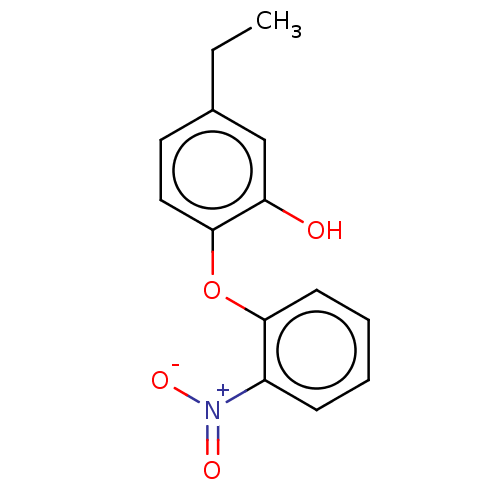
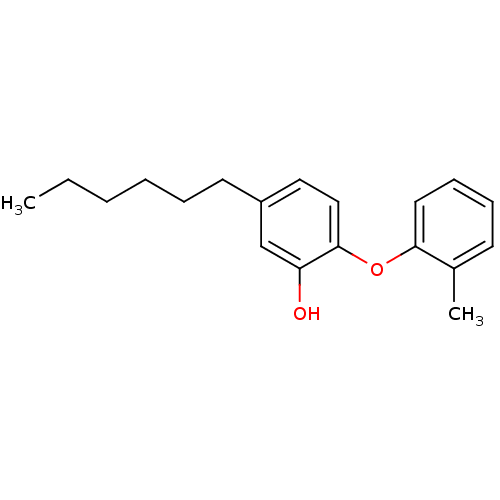
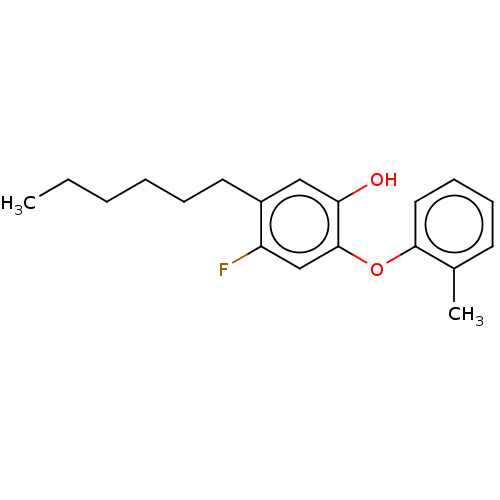
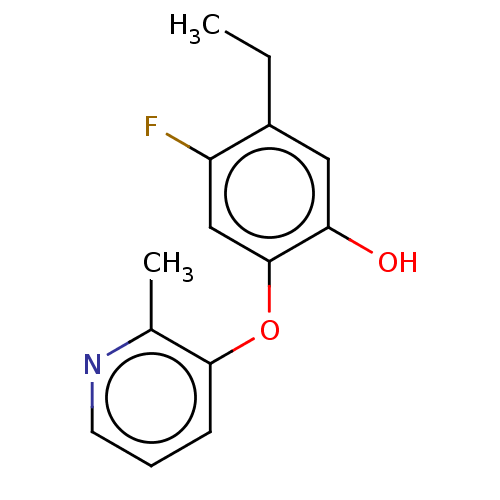
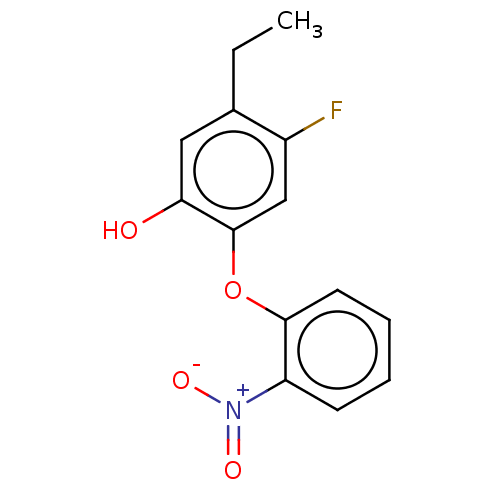
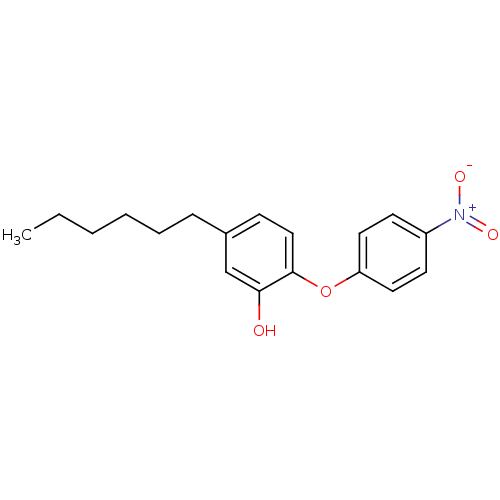
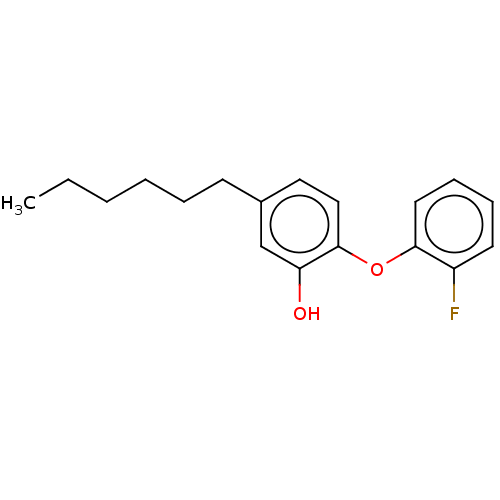
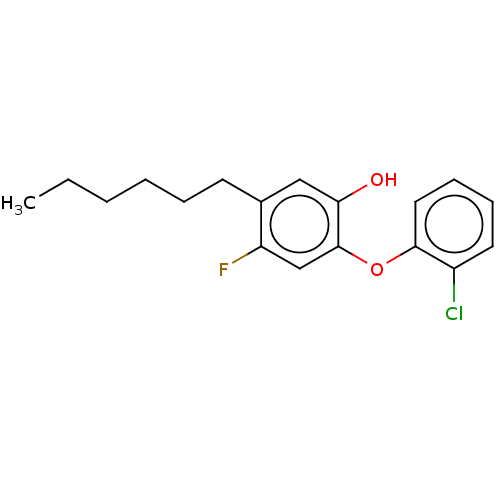
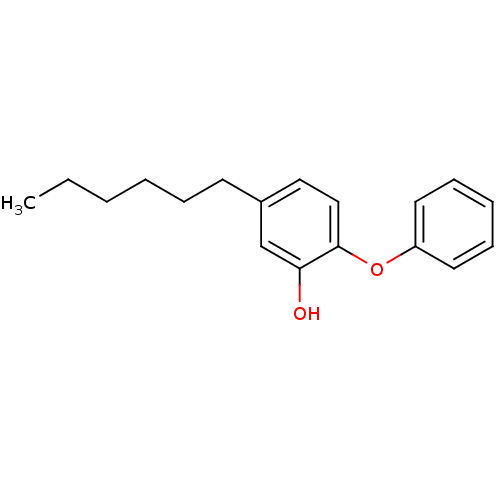
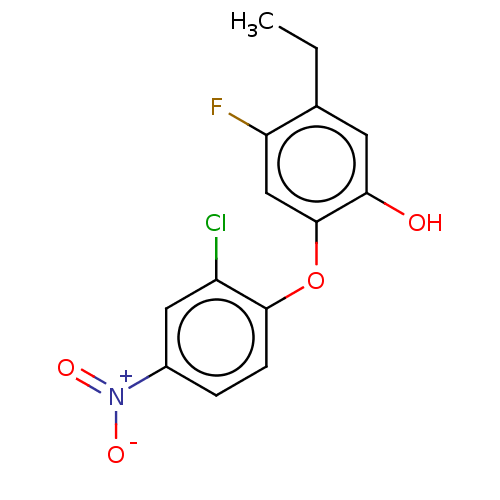
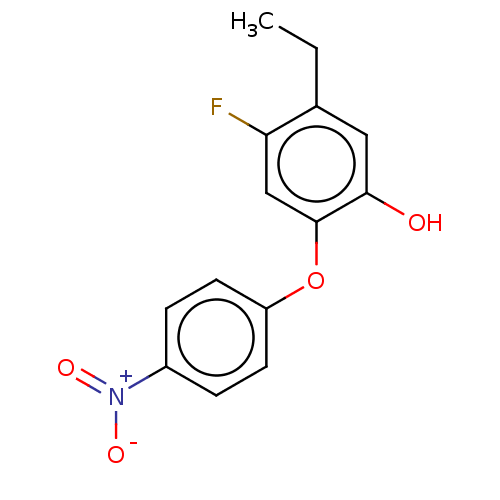

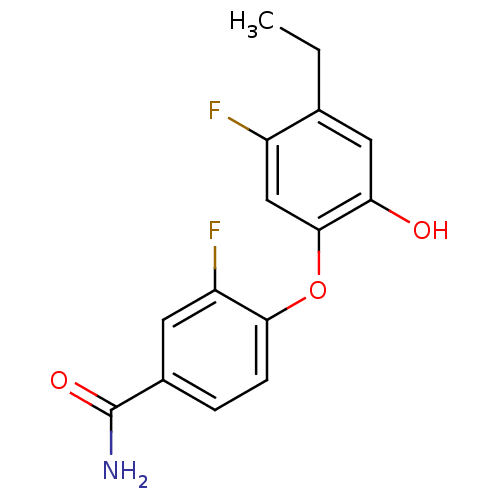
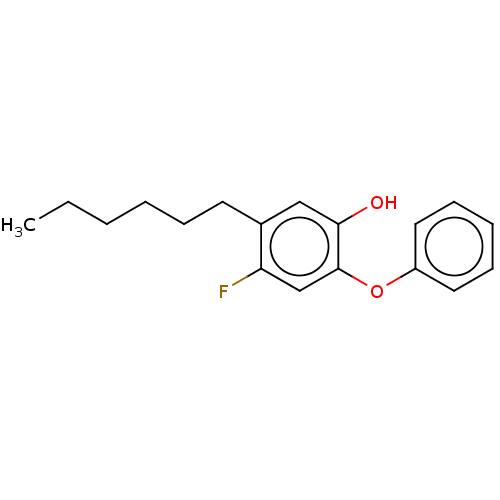
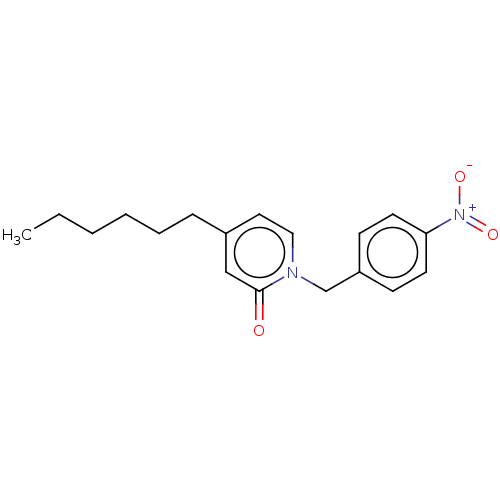
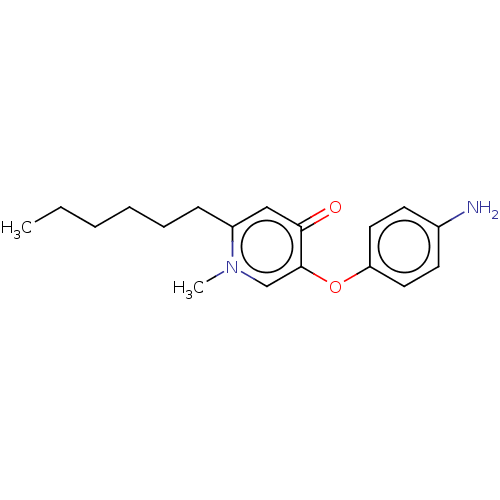
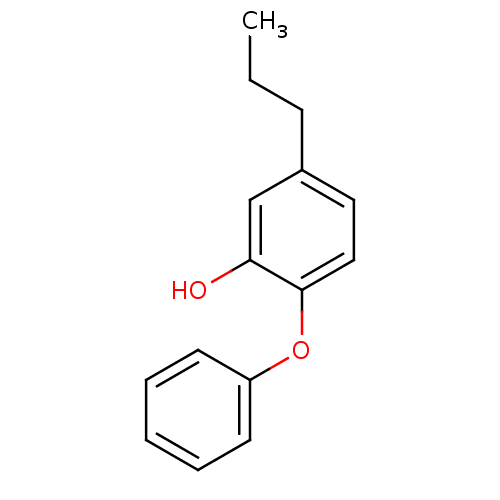
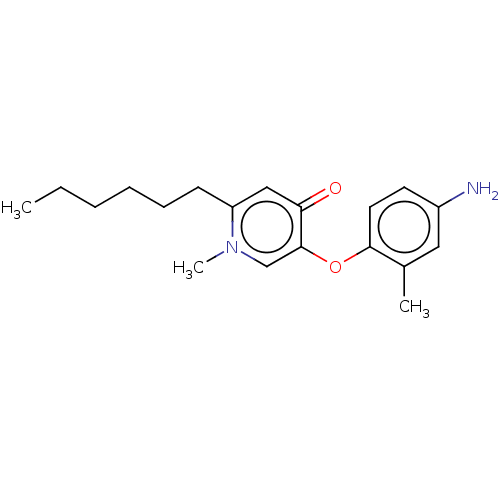
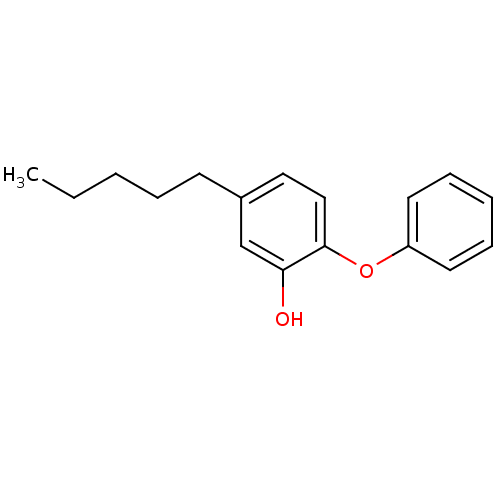
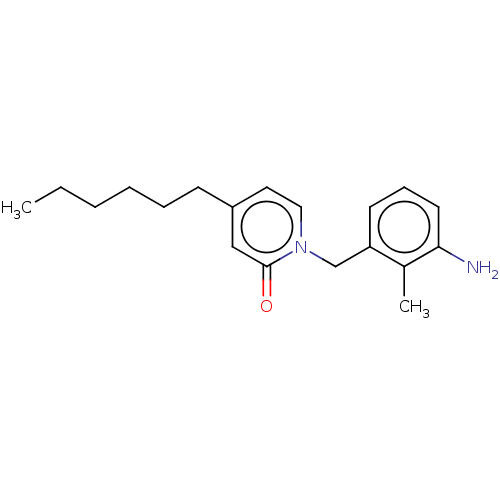
 3D Structure (crystal)
3D Structure (crystal)