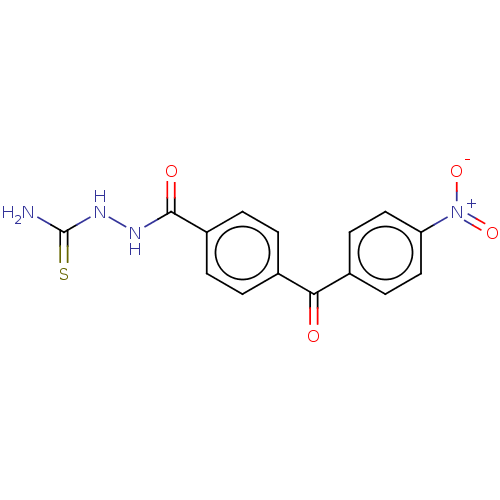
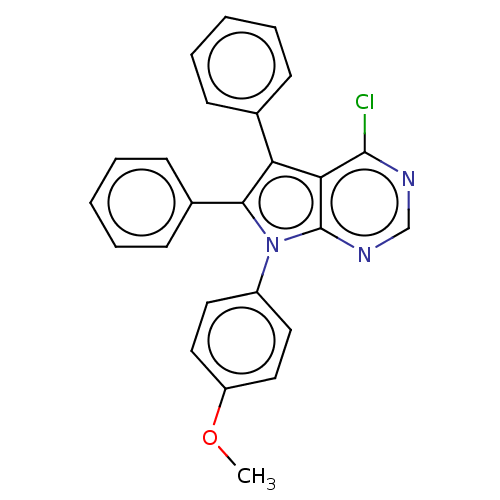
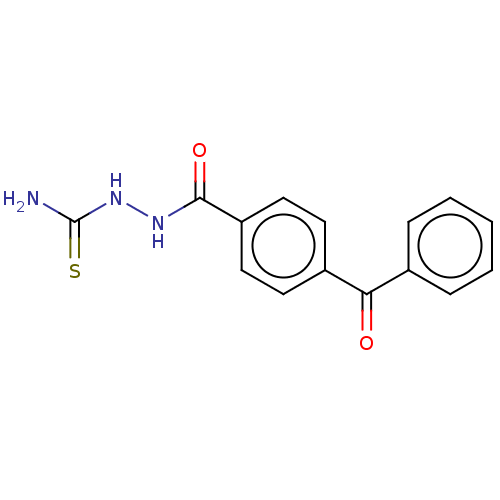
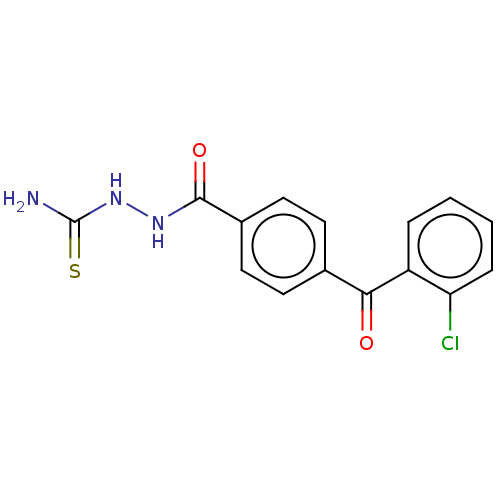
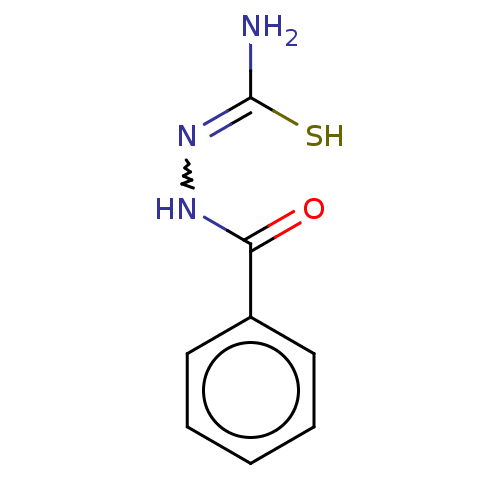
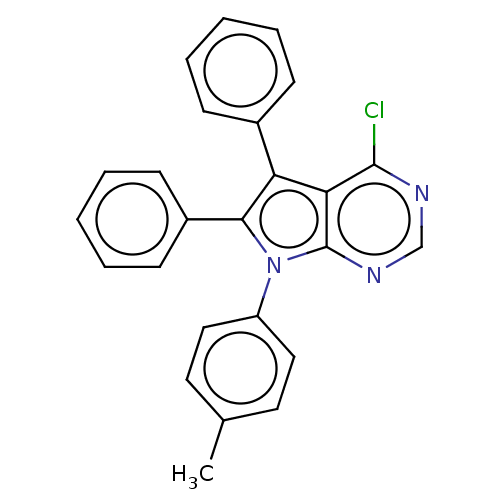
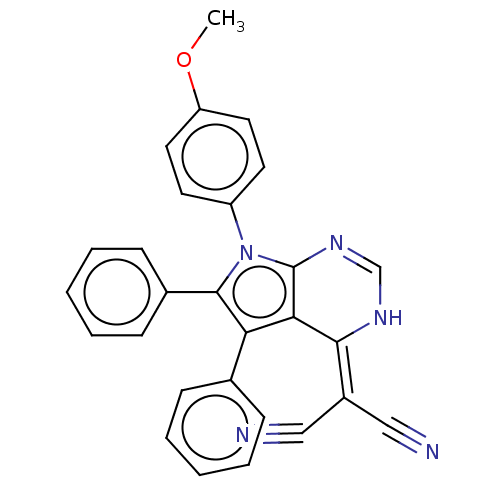
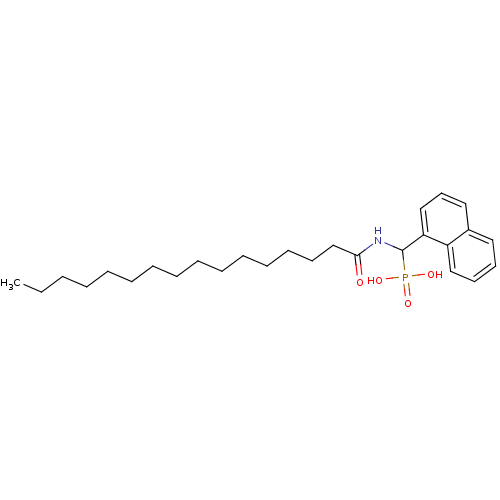
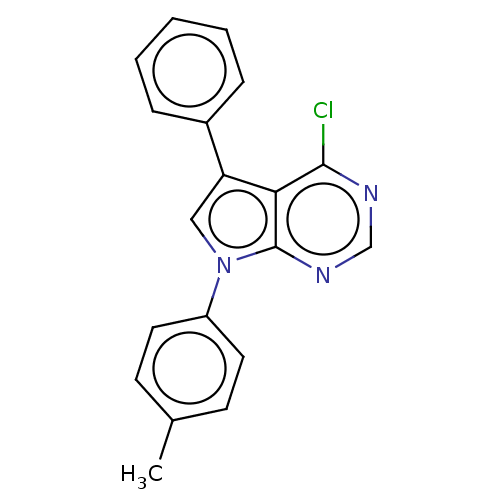
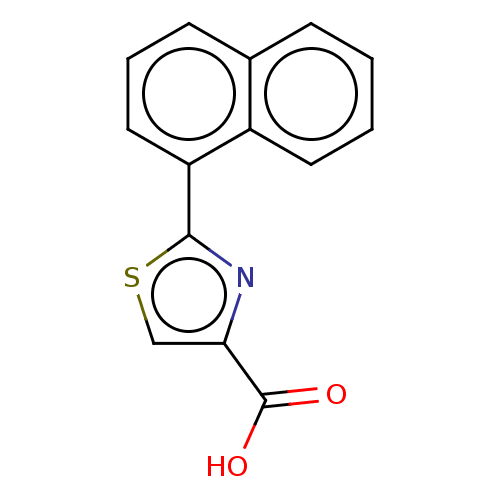
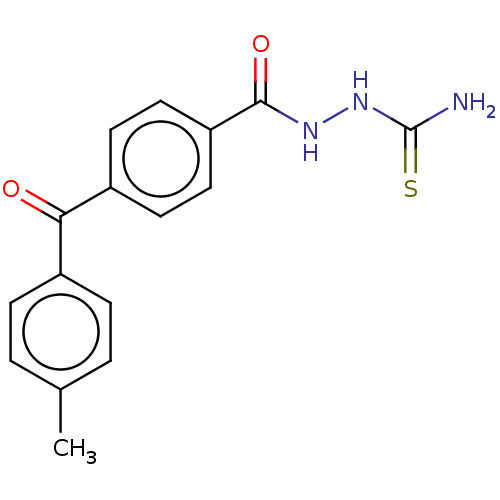
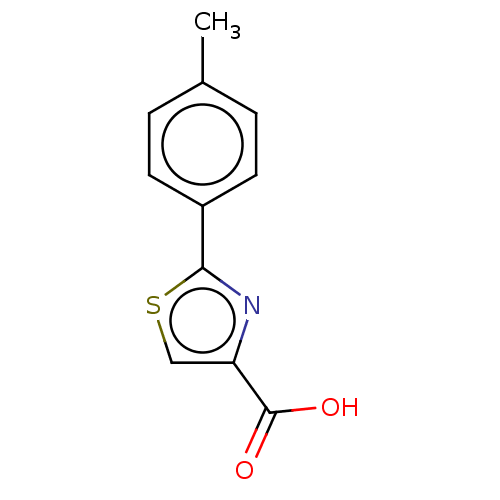
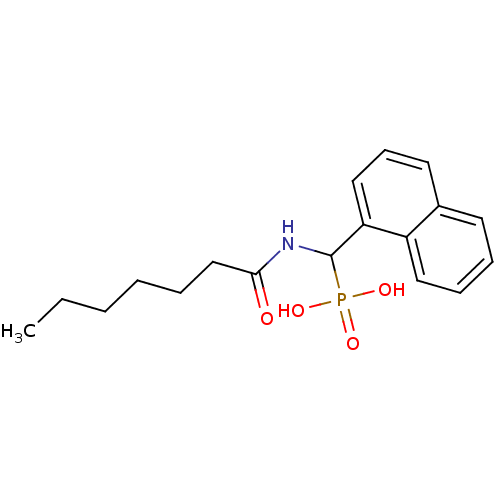
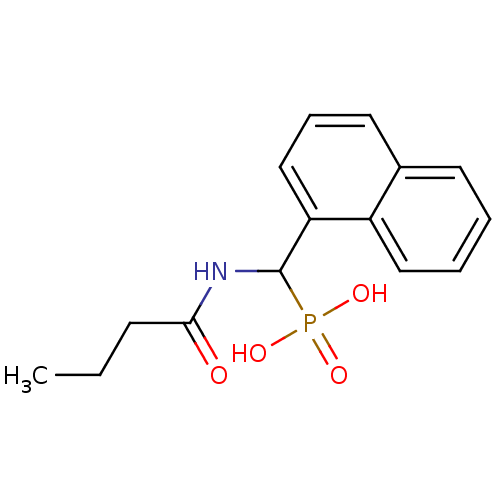
Affinity DataKi: 0.730nMAssay Description:Displacement of [3H]DAMGO from MOP receptor expressed in CHO-K1 cells after overnight incubation by beta counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 0.760nMAssay Description:Displacement of [3H]DAMGO from MOP receptor expressed in CHO-K1 cells after overnight incubation by beta counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Displacement of [3H]DAMGO from MOP receptor expressed in CHO-K1 cells after overnight incubation by beta counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 3.90nMAssay Description:Displacement of [3H]DAMGO from MOP receptor expressed in CHO-K1 cells after overnight incubation by beta counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 132nMAssay Description:Displacement of [3H]DAMGO from MOP receptor expressed in CHO-K1 cells after overnight incubation by beta counting methodMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 200nMAssay Description:Competitive inhibition of red kidney bean PAP using varying levels of pNPP as substrate measured at pH 6.2 by UV-vis spectrophotometric analysisMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 400nMAssay Description:Competitive inhibition of red kidney bean PAP using varying levels of pNPP as substrate measured at pH 6.2 by UV-vis spectrophotometric analysisMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 500nMAssay Description:Competitive inhibition of red kidney bean PAP using varying levels of pNPP as substrate measured at pH 6.2 by UV-vis spectrophotometric analysisMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 800nMAssay Description:Competitive inhibition of red kidney bean PAP using varying levels of pNPP as substrate measured at pH 6.2 by UV-vis spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataKi: 1.23E+3nMAssay Description:Displacement of [3H]DPDPE from DOP receptor expressed in CHO-K1 cells after overnight incubation by beta counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 1.56E+3nMAssay Description:Displacement of [3H]DPDPE from DOP receptor expressed in CHO-K1 cells after overnight incubation by beta counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 1.98E+3nMAssay Description:Displacement of [3H]DPDPE from DOP receptor expressed in CHO-K1 cells after overnight incubation by beta counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 2.25E+3nMAssay Description:Displacement of [3H]DPDPE from DOP receptor expressed in CHO-K1 cells after overnight incubation by beta counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 2.77E+3nMAssay Description:Displacement of [3H]DPDPE from DOP receptor expressed in CHO-K1 cells after overnight incubation by beta counting methodMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 4.50E+3nMAssay Description:Competitive inhibition of red kidney bean PAP using varying levels of pNPP as substrate measured at pH 6.2 by UV-vis spectrophotometric analysisMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 5.00E+3nMpH: 4.9Assay Description:Competitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometryMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University of Queensland (Uq)
Curated by ChEMBL
The University of Queensland (Uq)
Curated by ChEMBL
Affinity DataKi: 1.10E+4nMAssay Description:Non-competitive inhibition of mushroom tyrosinase using L-DOPA as substrate preincubated for 10 mins followed by substrate addition and further incub...More data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.10E+4nMpH: 4.9Assay Description:Competitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometryMore data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Serratia marcescens)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.10E+4nMAssay Description:Competitive inhibition of Pseudomonas aeruginosa beta-lactamase IMP-1 assessed as formation of 4-nitrothiophenolate at pH 7 using CENTA as chromogeni...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Serratia marcescens)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.20E+4nMAssay Description:Competitive inhibition of Pseudomonas aeruginosa metallo-beta-lactamase IMP-1 expressed in Escherichia coli BL21(DE3) using CENTA as substrate by spe...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Serratia marcescens)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.30E+4nMAssay Description:Competitive inhibition of Pseudomonas aeruginosa beta-lactamase IMP-1 assessed as formation of 4-nitrothiophenolate at pH 7 using CENTA as chromogeni...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Serratia marcescens)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.40E+4nMAssay Description:Competitive inhibition of Pseudomonas aeruginosa beta-lactamase IMP-1 assessed as formation of 4-nitrothiophenolate at pH 7 using CENTA as chromogeni...More data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.40E+4nMAssay Description:Competitive inhibition of red kidney bean PAP using varying levels of pNPP as substrate measured at pH 6.2 by UV-vis spectrophotometric analysisMore data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Serratia marcescens)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.50E+4nMAssay Description:Competitive inhibition of Pseudomonas aeruginosa metallo-beta-lactamase IMP-1 expressed in Escherichia coli BL21(DE3) using CENTA as substrate by spe...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Serratia marcescens)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.60E+4nMAssay Description:Competitive inhibition of Pseudomonas aeruginosa beta-lactamase IMP-1 assessed as formation of 4-nitrothiophenolate at pH 7 using CENTA as chromogeni...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Serratia marcescens)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.80E+4nMAssay Description:Competitive inhibition of Pseudomonas aeruginosa beta-lactamase IMP-1 assessed as formation of 4-nitrothiophenolate at pH 7 using CENTA as chromogeni...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Serratia marcescens)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.80E+4nMAssay Description:Competitive inhibition of Pseudomonas aeruginosa metallo-beta-lactamase IMP-1 expressed in Escherichia coli BL21(DE3) using CENTA as substrate by spe...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Serratia marcescens)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.90E+4nMAssay Description:Competitive inhibition of Pseudomonas aeruginosa beta-lactamase IMP-1 assessed as formation of 4-nitrothiophenolate at pH 7 using CENTA as chromogeni...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Serratia marcescens)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.90E+4nMAssay Description:Competitive inhibition of Pseudomonas aeruginosa metallo-beta-lactamase IMP-1 expressed in Escherichia coli BL21(DE3) using CENTA as substrate by spe...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Serratia marcescens)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 2.90E+4nMAssay Description:Competitive inhibition of Pseudomonas aeruginosa metallo-beta-lactamase IMP-1 expressed in Escherichia coli BL21(DE3) using CENTA as substrate by spe...More data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 3.10E+4nMpH: 4.9Assay Description:Competitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometryMore data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Serratia marcescens)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 3.30E+4nMAssay Description:Competitive inhibition of Pseudomonas aeruginosa metallo-beta-lactamase IMP-1 expressed in Escherichia coli BL21(DE3) using CENTA as substrate by spe...More data for this Ligand-Target Pair
TargetTartrate-resistant acid phosphatase type 5(Sus scrofa)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 3.30E+4nMAssay Description:Competitive inhibition of pig purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV/Vis spectro...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Serratia marcescens)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 4.10E+4nMAssay Description:Competitive inhibition of Pseudomonas aeruginosa beta-lactamase IMP-1 assessed as formation of 4-nitrothiophenolate at pH 7 using CENTA as chromogeni...More data for this Ligand-Target Pair
TargetTartrate-resistant acid phosphatase type 5(Sus scrofa)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 4.20E+4nMAssay Description:Competitive inhibition of pig purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV/Vis spectro...More data for this Ligand-Target Pair
TargetTartrate-resistant acid phosphatase type 5(Sus scrofa)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 4.90E+4nMAssay Description:Competitive inhibition of pig purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV/Vis spectro...More data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 5.70E+4nMpH: 4.9Assay Description:Competitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometryMore data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Serratia marcescens)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 7.50E+4nMAssay Description:Competitive inhibition of Pseudomonas aeruginosa beta-lactamase IMP-1 assessed as formation of 4-nitrothiophenolate at pH 7 using CENTA as chromogeni...More data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.02E+5nMpH: 4.9Assay Description:Uncompetitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometr...More data for this Ligand-Target Pair
TargetTartrate-resistant acid phosphatase type 5(Sus scrofa)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.20E+5nMAssay Description:Competitive inhibition of pig purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV/Vis spectro...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University of Queensland (Uq)
Curated by ChEMBL
The University of Queensland (Uq)
Curated by ChEMBL
Affinity DataKi: 1.30E+5nMAssay Description:Non-competitive inhibition of mushroom tyrosinase using L-DOPA as substrate preincubated for 10 mins followed by substrate addition and further incub...More data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.60E+5nMAssay Description:Inhibition of red kidney bean PAP using pNPP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: 1.85E+5nMAssay Description:Competitive inhibition of red kidney bean purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV...More data for this Ligand-Target Pair
TargetTartrate-resistant acid phosphatase type 5(Sus scrofa)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.90E+5nMAssay Description:Competitive inhibition of pig purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV/Vis spectro...More data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.95E+5nMpH: 4.9Assay Description:Competitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometryMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 2.22E+5nMpH: 4.9Assay Description:Competitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometryMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 2.38E+5nMpH: 4.9Assay Description:Competitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometryMore data for this Ligand-Target Pair
Affinity DataKi: 3.40E+5nMAssay Description:Competitive inhibition of red kidney bean purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV...More data for this Ligand-Target Pair
Affinity DataKi: 4.00E+5nMAssay Description:Competitive inhibition of red kidney bean purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV...More data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 4.43E+5nMpH: 4.9Assay Description:Uncompetitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometr...More data for this Ligand-Target Pair

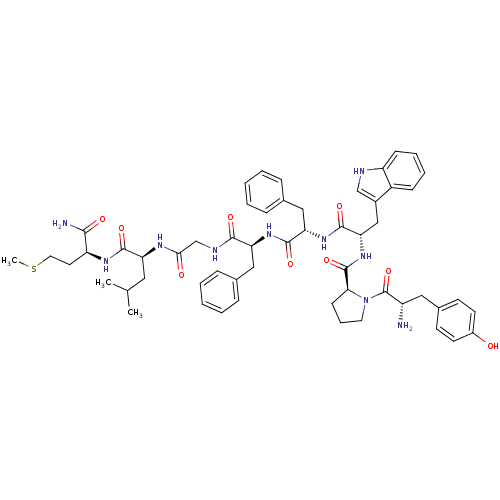
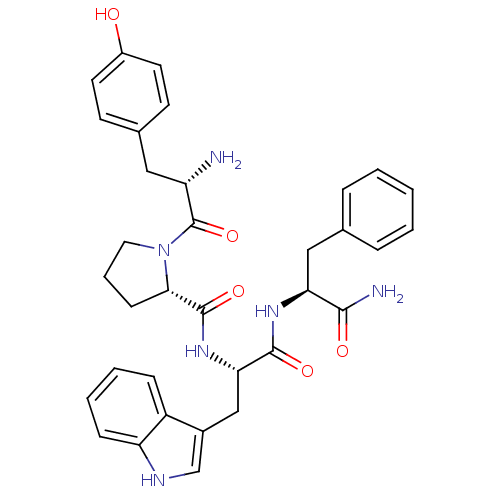
 3D Structure (crystal)
3D Structure (crystal)