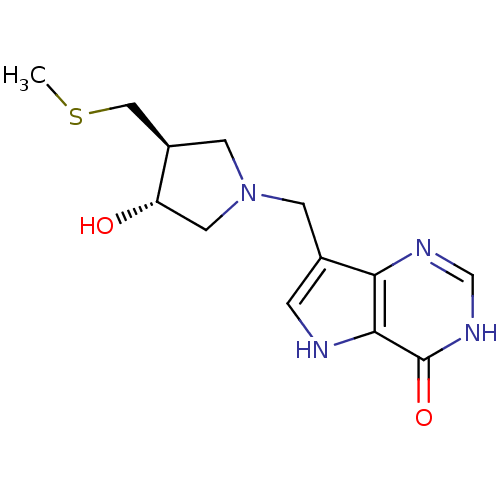
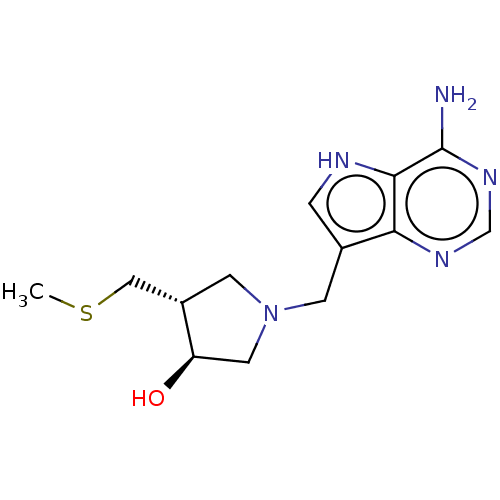
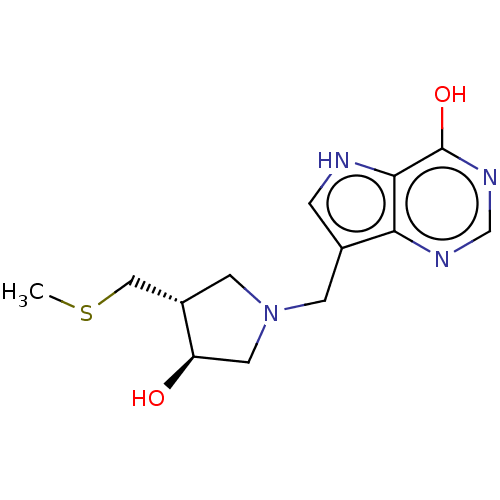
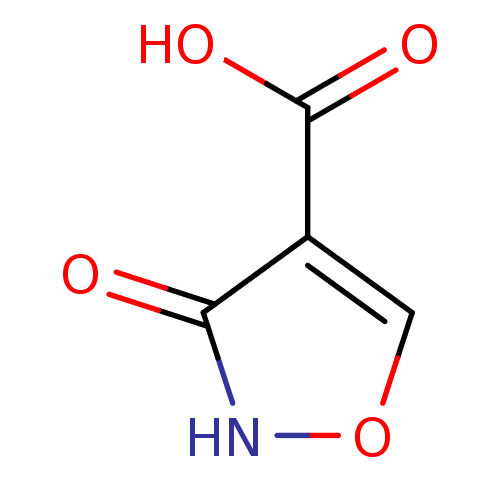
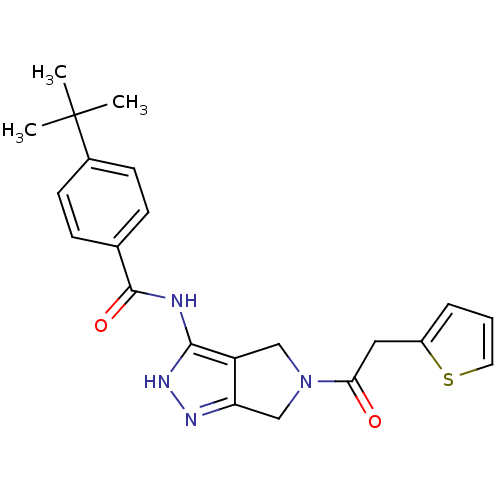
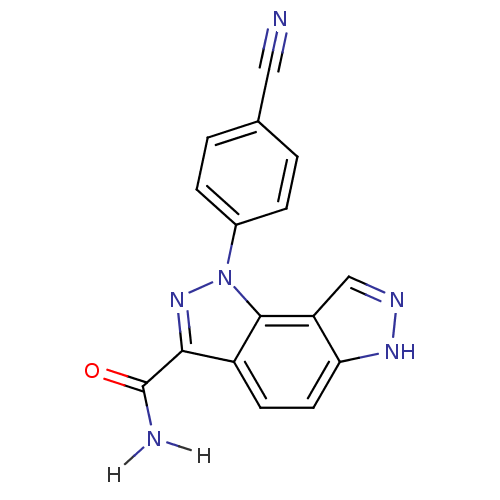
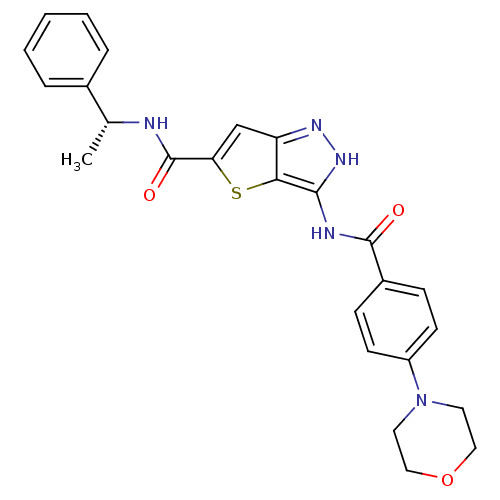
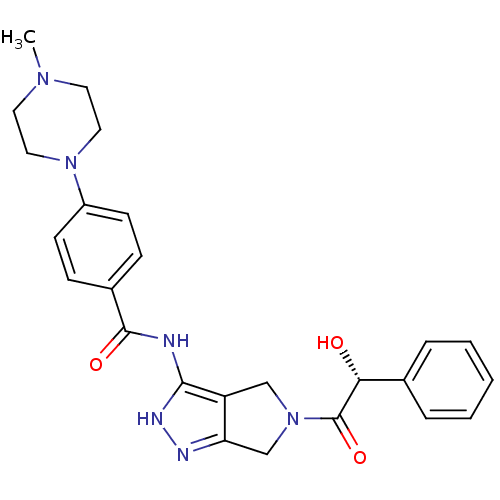
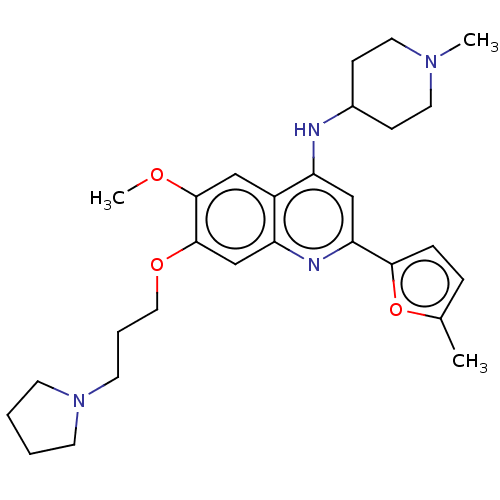
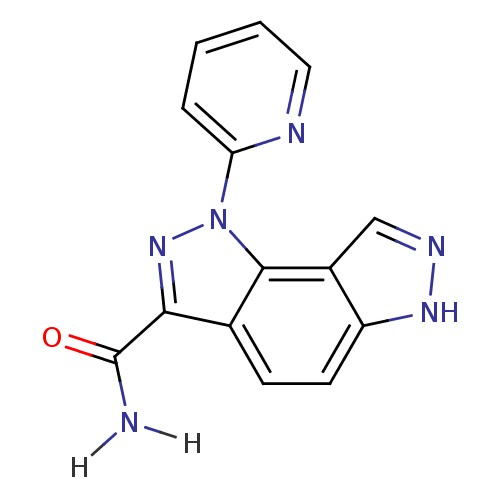
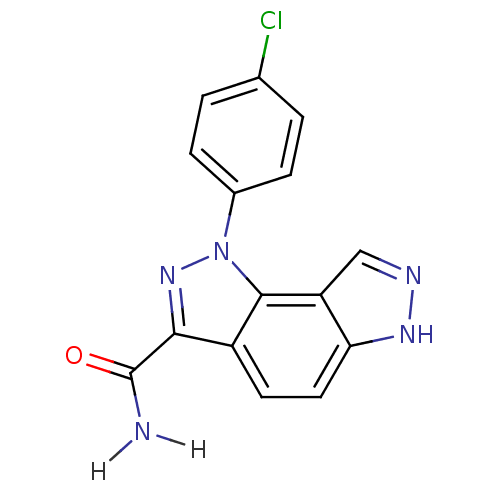
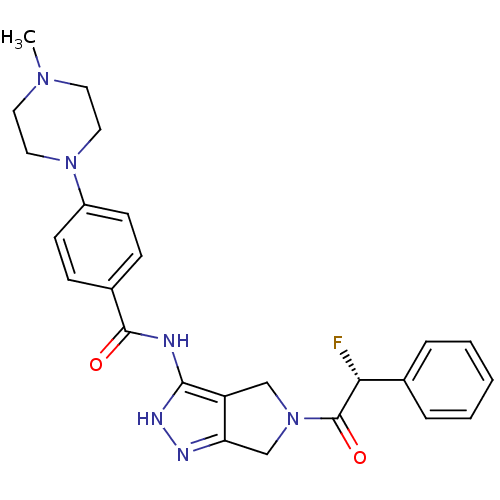
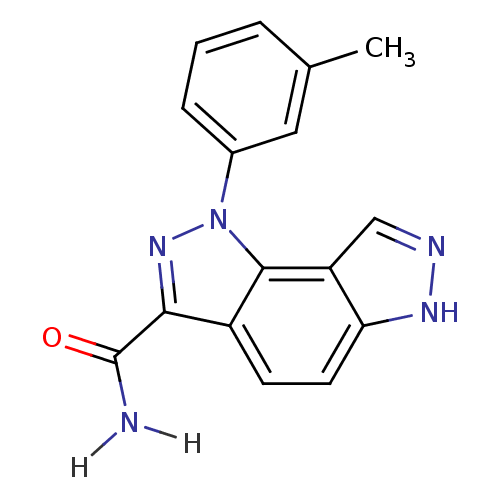
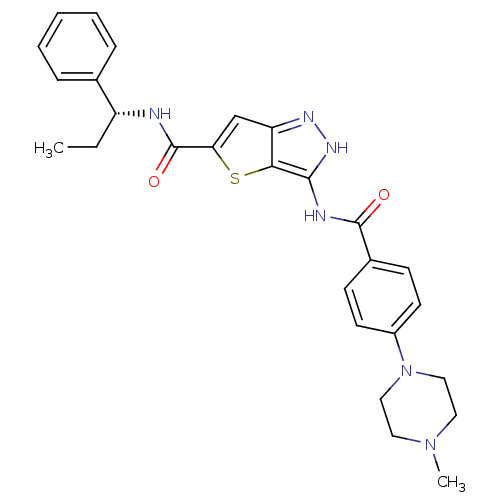
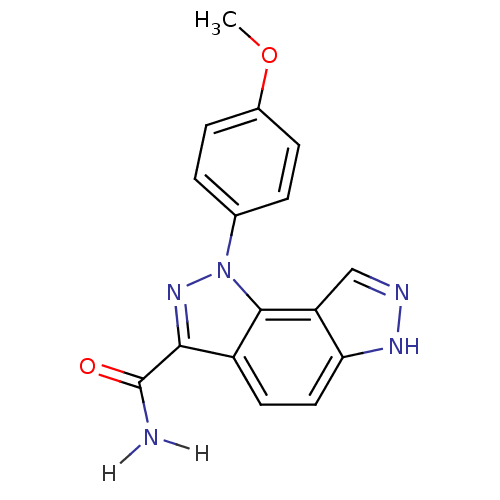
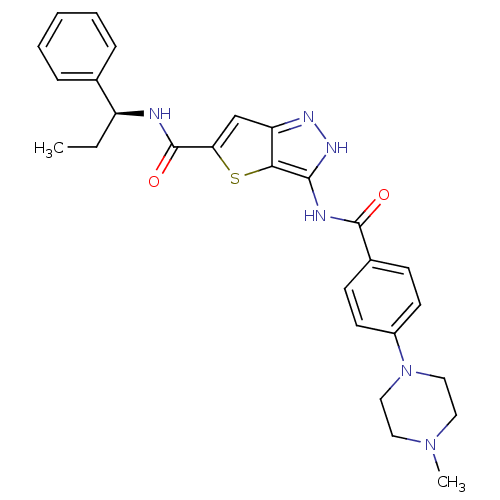
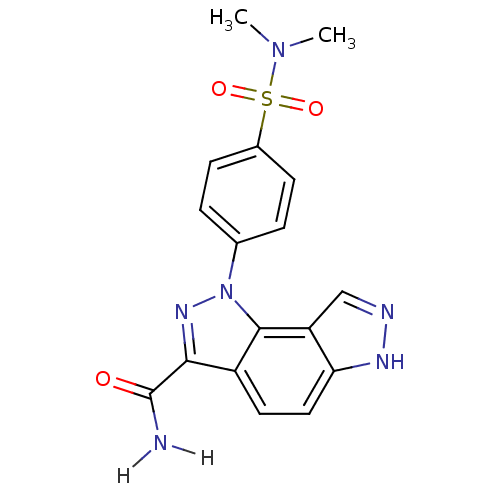
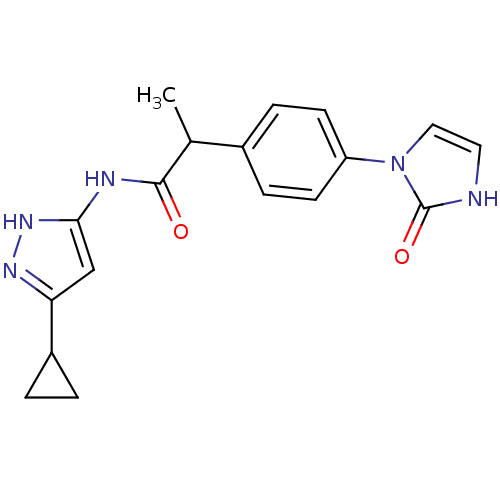
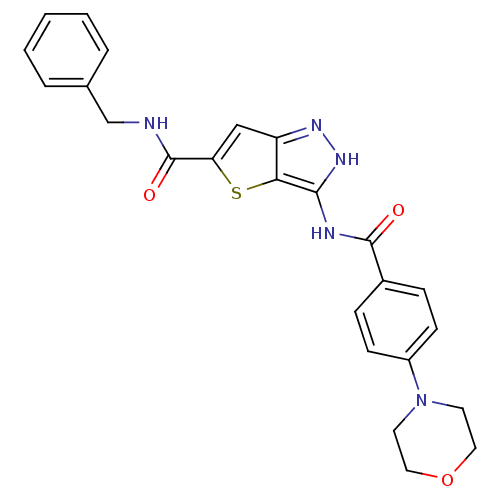
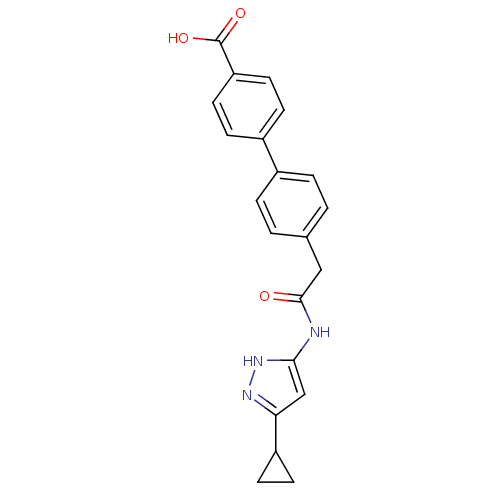
Target5'-methylthioadenosine/S-adenosylhomocysteine nucleosidase(Escherichia coli)
Victoria University Of Wellington
Curated by ChEMBL
Victoria University Of Wellington
Curated by ChEMBL
Affinity DataKi: 0.00200nMAssay Description:Inhibition of recombinant Escherichia coli MTAN expressed in Escherichia coli BL-21 DE3 using methylthioadenosine as substrate assessed as inhibition...More data for this Ligand-Target Pair
Target5'-methylthioadenosine/S-adenosylhomocysteine nucleosidase(Escherichia coli)
Victoria University Of Wellington
Curated by ChEMBL
Victoria University Of Wellington
Curated by ChEMBL
Affinity DataKi: 0.0480nMAssay Description:Inhibition of recombinant Escherichia coli MTAN expressed in Escherichia coli BL-21 DE3 using methylthioadenosine as substrate by xanthine oxidase co...More data for this Ligand-Target Pair
TargetPurine nucleoside phosphorylase(Homo sapiens (Human))
Victoria University Of Wellington
Curated by ChEMBL
Victoria University Of Wellington
Curated by ChEMBL
Affinity DataKi: 0.0700nMAssay Description:Inhibition of recombinant human PNP using inosine as substrate assessed as inhibition constant for slow onset inhibition of enzyme-inhibitor complex ...More data for this Ligand-Target Pair
TargetS-methyl-5'-thioadenosine phosphorylase(Homo sapiens (Human))
Victoria University Of Wellington
Curated by ChEMBL
Victoria University Of Wellington
Curated by ChEMBL
Affinity DataKi: 0.0780nMAssay Description:Inhibition of human MTAP using methylthioadenosine as substrate assessed as inhibition constant for slow onset inhibition of enzyme-inhibitor complex...More data for this Ligand-Target Pair
TargetPurine nucleoside phosphorylase(Homo sapiens (Human))
Victoria University Of Wellington
Curated by ChEMBL
Victoria University Of Wellington
Curated by ChEMBL
Affinity DataKi: 0.300nMAssay Description:Inhibition of recombinant human PNP using inosine as substrate by xanthine oxidase coupling enzyme assayMore data for this Ligand-Target Pair
TargetS-methyl-5'-thioadenosine phosphorylase(Homo sapiens (Human))
Victoria University Of Wellington
Curated by ChEMBL
Victoria University Of Wellington
Curated by ChEMBL
Affinity DataKi: 0.530nMAssay Description:Inhibition of human MTAP using methylthioadenosine as substrate by xanthine oxidase coupling enzyme assayMore data for this Ligand-Target Pair
TargetPurine nucleoside phosphorylase(Plasmodium falciparum)
Victoria University Of Wellington
Curated by ChEMBL
Victoria University Of Wellington
Curated by ChEMBL
Affinity DataKi: 0.900nMAssay Description:Inhibition of recombinant Plasmodium falciparum PNP using inosine as substrate assessed as inhibition constant for slow onset inhibition of enzyme-in...More data for this Ligand-Target Pair
Target5'-methylthioadenosine/S-adenosylhomocysteine nucleosidase(Escherichia coli)
Victoria University Of Wellington
Curated by ChEMBL
Victoria University Of Wellington
Curated by ChEMBL
Affinity DataKi: 4.5nMAssay Description:Inhibition of recombinant Escherichia coli MTAN expressed in Escherichia coli BL-21 DE3 using methylthioadenosine as substrate by xanthine oxidase co...More data for this Ligand-Target Pair
TargetPurine nucleoside phosphorylase(Plasmodium falciparum)
Victoria University Of Wellington
Curated by ChEMBL
Victoria University Of Wellington
Curated by ChEMBL
Affinity DataKi: 11nMAssay Description:Inhibition of recombinant Plasmodium falciparum PNP using inosine as substrate by xanthine oxidase coupling enzyme assayMore data for this Ligand-Target Pair
TargetPurine nucleoside phosphorylase(Homo sapiens (Human))
Victoria University Of Wellington
Curated by ChEMBL
Victoria University Of Wellington
Curated by ChEMBL
Affinity DataKi: 84nMAssay Description:Inhibition of recombinant human PNP using inosine as substrate assessed as inhibition constant for slow onset inhibition of enzyme-inhibitor complex ...More data for this Ligand-Target Pair
TargetPurine nucleoside phosphorylase(Plasmodium falciparum)
Victoria University Of Wellington
Curated by ChEMBL
Victoria University Of Wellington
Curated by ChEMBL
Affinity DataKi: 93nMAssay Description:Inhibition of recombinant Plasmodium falciparum PNP using inosine as substrate assessed as inhibition constant for slow onset inhibition of enzyme-in...More data for this Ligand-Target Pair
TargetPurine nucleoside phosphorylase(Homo sapiens (Human))
Victoria University Of Wellington
Curated by ChEMBL
Victoria University Of Wellington
Curated by ChEMBL
Affinity DataKi: 110nMAssay Description:Inhibition of recombinant human PNP using inosine as substrate by xanthine oxidase coupling enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 210nM ΔG°: -38.1kJ/mole IC50: 650nMpH: 7.5 T: 2°CAssay Description:An LDH enzymatic assay developed for high throughput format was used. The dehydrogenase reaction was run in the lactate to pyruvate direction and cou...More data for this Ligand-Target Pair
Affinity DataKi: 290nM ΔG°: -37.3kJ/mole IC50: 140nMpH: 7.5 T: 2°CAssay Description:An LDH enzymatic assay developed for high throughput format was used. The dehydrogenase reaction was run in the lactate to pyruvate direction and cou...More data for this Ligand-Target Pair
TargetPurine nucleoside phosphorylase(Plasmodium falciparum)
Victoria University Of Wellington
Curated by ChEMBL
Victoria University Of Wellington
Curated by ChEMBL
Affinity DataKi: 298nMAssay Description:Inhibition of recombinant Plasmodium falciparum PNP using inosine as substrate by xanthine oxidase coupling enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 470nM ΔG°: -36.1kJ/mole IC50: 1.10E+3nMpH: 7.5 T: 2°CAssay Description:An LDH enzymatic assay developed for high throughput format was used. The dehydrogenase reaction was run in the lactate to pyruvate direction and cou...More data for this Ligand-Target Pair
TargetS-methyl-5'-thioadenosine phosphorylase(Homo sapiens (Human))
Victoria University Of Wellington
Curated by ChEMBL
Victoria University Of Wellington
Curated by ChEMBL
Affinity DataKi: 750nMAssay Description:Inhibition of human MTAP using methylthioadenosine as substrate by xanthine oxidase coupling enzyme assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMpH: 7.4 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMpH: 7.4 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates, followed by quantitation of the phosphorylat...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMpH: 7.4 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 2nMpH: 7.4 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 2nMpH: 7.4 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates, followed by quantitation of the phosphorylat...More data for this Ligand-Target Pair
Affinity DataIC50: 2nMpH: 7.4 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates, followed by quantitation of the phosphorylat...More data for this Ligand-Target Pair
Affinity DataIC50: 2nMpH: 7.4 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMpH: 7.4 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMpH: 7.5 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMpH: 7.5 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Victoria University Of Wellington
Curated by ChEMBL
Victoria University Of Wellington
Curated by ChEMBL
Affinity DataIC50: 8nMAssay Description:Inhibition of G9a (unknown origin) using biotinylated histone monomethyl-H3K9 peptide by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 9nMpH: 7.5 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMpH: 7.4 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMpH: 7.5 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzymes and substrates in the presence ATP/[gamma-33P] ATP. After in...More data for this Ligand-Target Pair

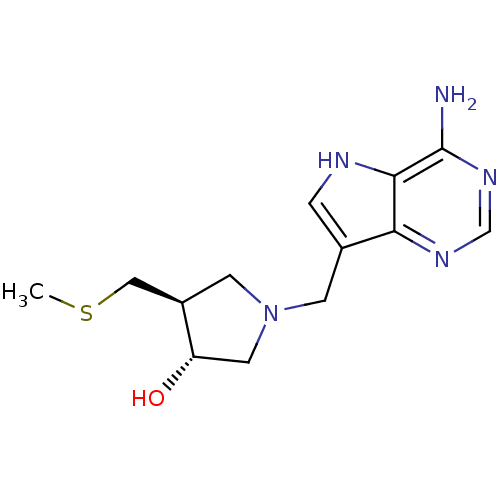
 3D Structure (crystal)
3D Structure (crystal)