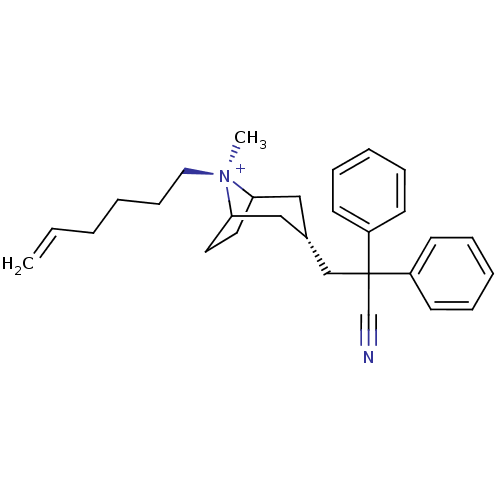
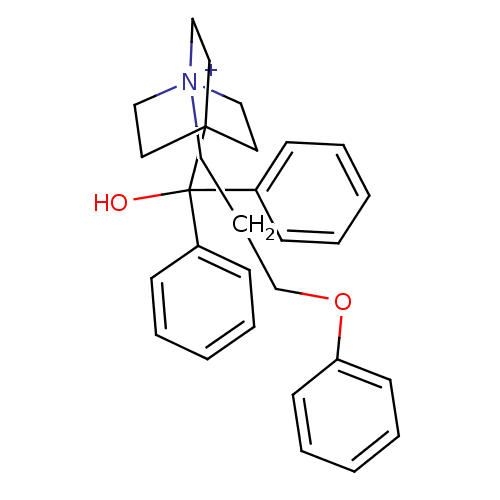
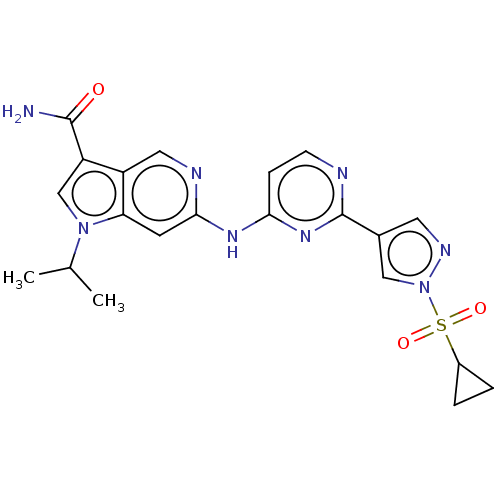
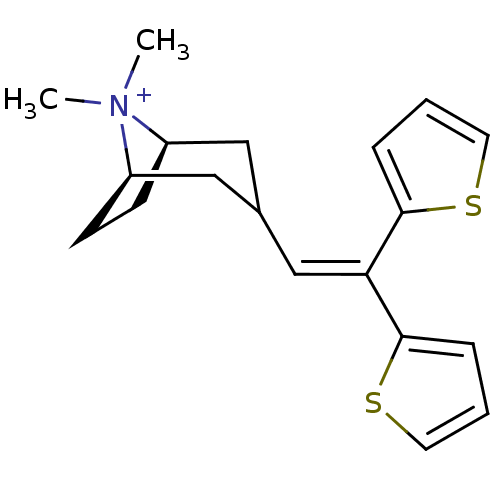
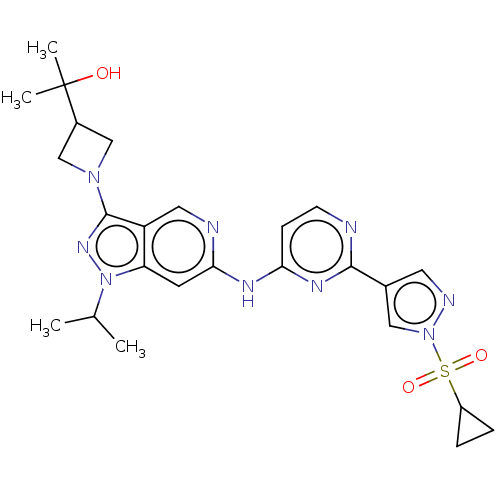
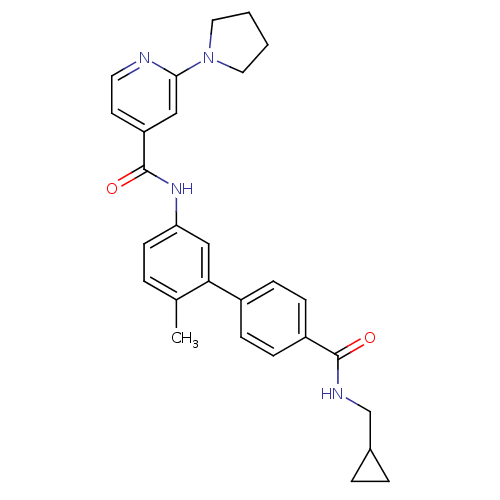
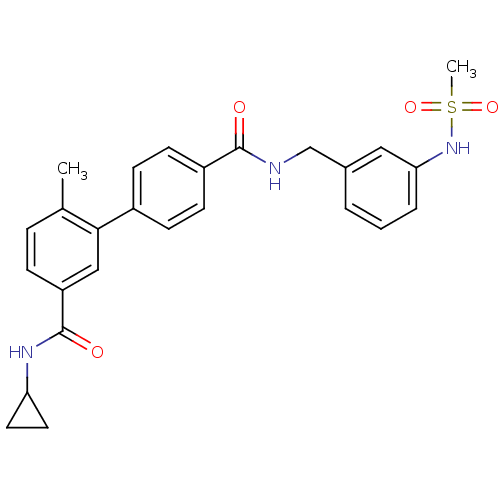
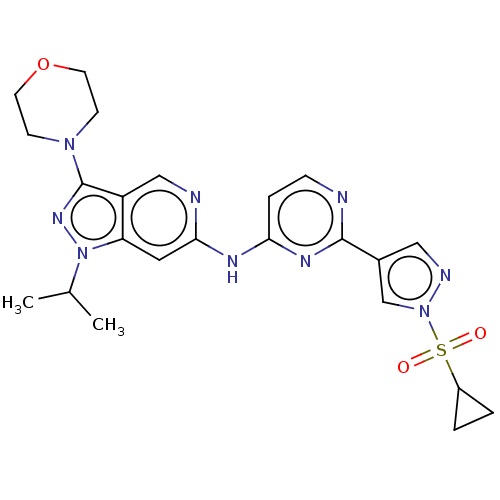
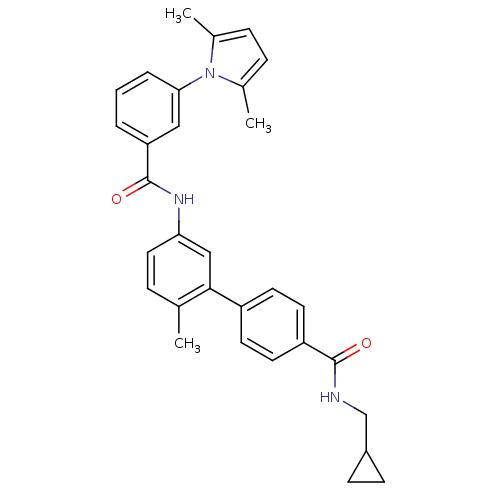
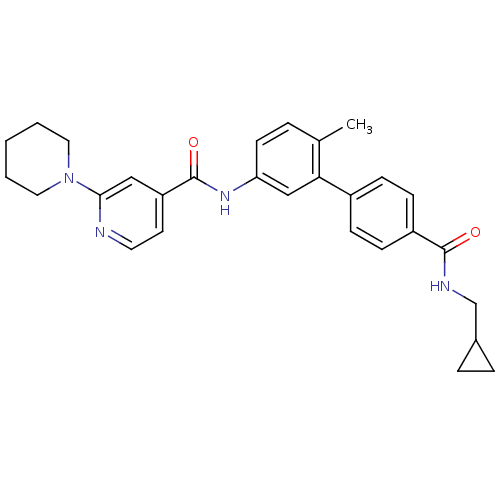
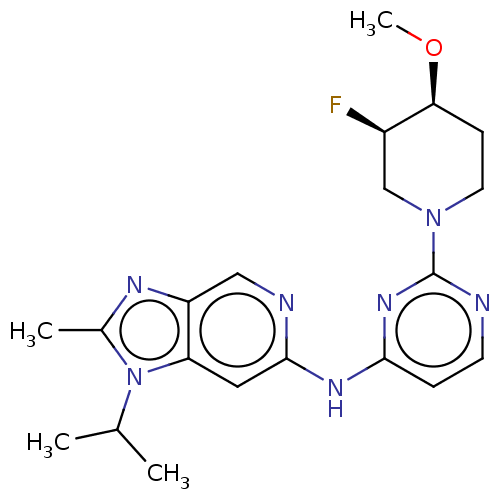
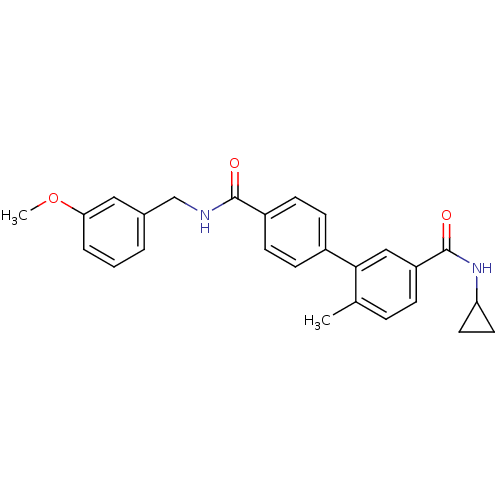
Affinity DataKi: 0.0316nMAssay Description:Displacement of [3H]N-methyl Scopolamine from human cloned muscarinic M3 receptor expressed in CHO cells by scintillation proximity assayMore data for this Ligand-Target Pair
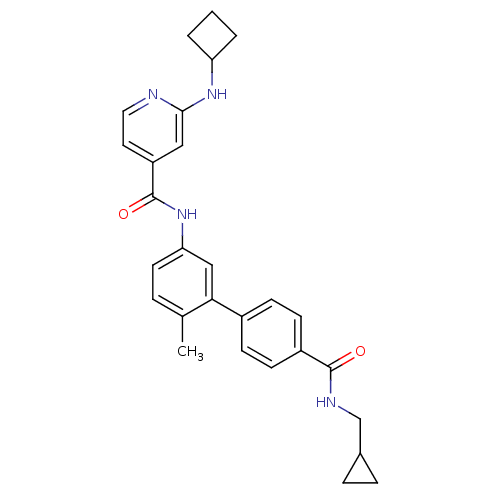
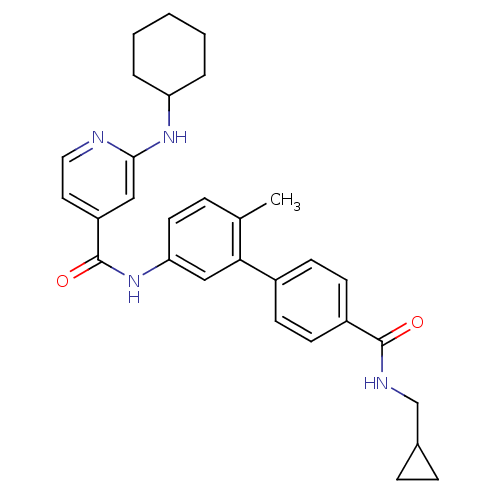
Affinity DataKi: 0.0500nMAssay Description:Displacement of [3H]N-methyl Scopolamine from human muscarinic M1 receptor expressed in CHO cells by scintillation proximity assayMore data for this Ligand-Target Pair
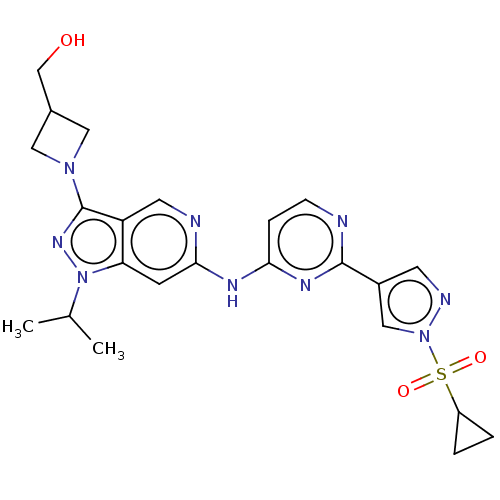
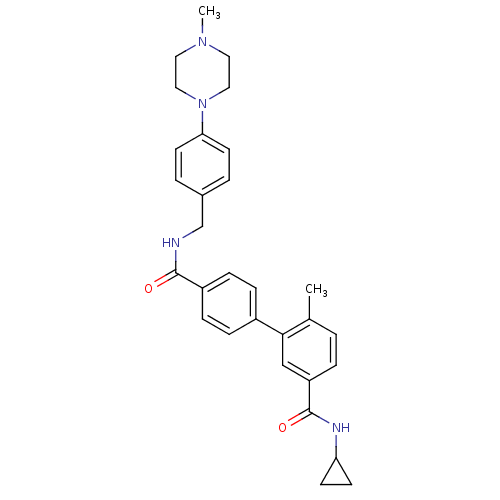
Affinity DataKi: 0.0600nMAssay Description:Displacement of [3H]N-methyl scopalamine from human cloned muscarinic M3 receptor expressed in CHO cells coexpressing Gqi5 by scintillation proximity...More data for this Ligand-Target Pair
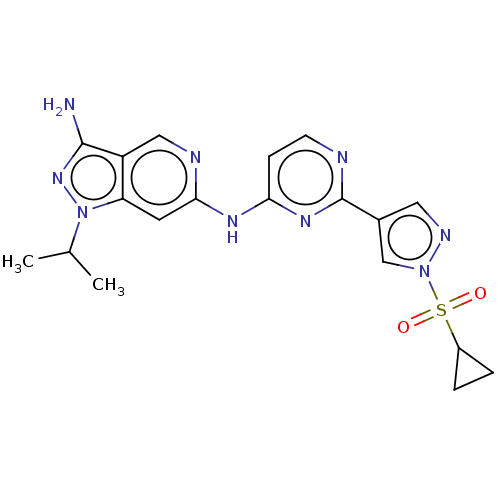
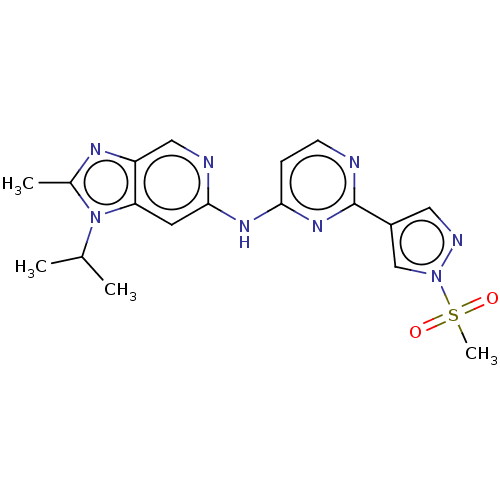
Affinity DataKi: 0.0600nMAssay Description:Displacement of [3H]N-methyl Scopolamine from human muscarinic M3 receptor expressed in CHO cells by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.100nMAssay Description:Inhibition of wild-type EGFR (unknown origin) by high-throughput biochemical screeningMore data for this Ligand-Target Pair
Affinity DataKi: 0.100nMAssay Description:Inhibition of EGFR T790M/L858R mutant (unknown origin) by high-throughput biochemical screeningMore data for this Ligand-Target Pair
Affinity DataKi: 0.100nMAssay Description:Displacement of [3H]-N-methyl scopolamine from muscarinic acetylcholine M2 receptor expressed in CHO cell membraneMore data for this Ligand-Target Pair
Affinity DataKi: 0.120nMAssay Description:Displacement of [3H]-N-methyl scopolamine from muscarinic acetylcholine M1 receptor expressed in CHO cell membraneMore data for this Ligand-Target Pair
Affinity DataKi: 0.130nMAssay Description:Displacement of [3H]N-methyl scopalamine from human cloned muscarinic M3 receptor expressed in CHO cells coexpressing Gqi5 by scintillation proximity...More data for this Ligand-Target Pair
Affinity DataKi: 0.150nMAssay Description:Displacement of [3H]N-methyl scopalamine from human cloned muscarinic M2 receptor expressed in CHO cells coexpressing Gqi5 by scintillation proximity...More data for this Ligand-Target Pair
Affinity DataKi: 0.160nMAssay Description:Displacement of [3H]-N-methyl scopolamine from muscarinic acetylcholine M3 receptor expressed in CHO cell membraneMore data for this Ligand-Target Pair
Affinity DataKi: 0.160nMAssay Description:Displacement of [3H]N-methyl scopalamine from human cloned muscarinic M1 receptor expressed in CHO cells coexpressing Gqi5 by scintillation proximity...More data for this Ligand-Target Pair
Affinity DataKi: 0.170nMAssay Description:Displacement of [3H]N-methyl Scopolamine from human muscarinic M2 receptor expressed in CHO cells by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Inhibition of EGFR T790M/del746 to 750 mutant (unknown origin) by high-throughput biochemical screeningMore data for this Ligand-Target Pair
Affinity DataKi: 0.350nMAssay Description:Inhibition of human EGFR T790M/del (746 to 750 residues) mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 peptide as subs...More data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 0.380nMAssay Description:Displacement of [3H]N-methyl Scopolamine from human muscarinic M1 receptor expressed in CHO cells by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.450nMAssay Description:Displacement of [3H]N-methyl Scopolamine from human muscarinic M3 receptor expressed in CHO cells by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataKi: <0.600nMAssay Description:Inhibition of human EGFR T790M/L858R double mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 as peptide substrate preincu...More data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 0.600nMAssay Description:Displacement of rhodamine-green labeled (2-(6-amino-3-imino-3H-xanthen-9-yl)-5-{[({4-[4-(4-Cl-3-hydroxyphenyl)-5-(4-pyridinyl)-1H-imidazol-2yl]phenyl...More data for this Ligand-Target Pair
Affinity DataKi: <0.600nMAssay Description:Inhibition of human EGFR T790M/L858R double mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 as peptide substrate preincu...More data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 0.640nMAssay Description:Displacement of [3H]N-methyl scopalamine from human cloned muscarinic M1 receptor expressed in CHO cells coexpressing Gqi5 by scintillation proximity...More data for this Ligand-Target Pair
Affinity DataKi: 0.730nMAssay Description:Displacement of [3H]N-methyl Scopolamine from human muscarinic M2 receptor expressed in CHO cells by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.790nMAssay Description:Inhibition of human EGFR T790M/L858R double mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 as peptide substrate preincu...More data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 0.940nMAssay Description:Displacement of [3H]N-methyl scopalamine from human cloned muscarinic M2 receptor expressed in CHO cells coexpressing Gqi5 by scintillation proximity...More data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Inhibition of human EGFR T790M/L858R double mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 as peptide substrate preincu...More data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 1.10nMAssay Description:Inhibition of human EGFR T790M/L858R double mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 as peptide substrate preincu...More data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 1.20nMAssay Description:Inhibition of human EGFR T790M/L858R double mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 as peptide substrate preincu...More data for this Ligand-Target Pair
Ligand InfoPDB
Affinity DataKi: 1.20nMAssay Description:Inhibition of human EGFR del (746 to 750) mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 as substrate preincubated for ...More data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 1.20nMAssay Description:Inhibition of human EGFR T790M/L858R double mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 as peptide substrate preincu...More data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 1.40nMAssay Description:Inhibition of human EGFR del (746 to 750) mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 as substrate preincubated for ...More data for this Ligand-Target Pair
Ligand InfoPDB
Affinity DataKi: 1.60nMAssay Description:Displacement of rhodamine-green labeled (2-(6-amino-3-imino-3H-xanthen-9-yl)-5-{[({4-[4-(4-Cl-3-hydroxyphenyl)-5-(4-pyridinyl)-1H-imidazol-2yl]phenyl...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Inhibition of human EGFR T790M/del (746 to 750 residues) mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 peptide as subs...More data for this Ligand-Target Pair
Ligand InfoPDB
Affinity DataKi: 1.80nMAssay Description:Inhibition of human EGFR T790M/del (746 to 750 residues) mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 peptide as subs...More data for this Ligand-Target Pair
Ligand InfoPDB
Affinity DataKi: 1.90nMAssay Description:Displacement of rhodamine-green labeled (2-(6-amino-3-imino-3H-xanthen-9-yl)-5-{[({4-[4-(4-Cl-3-hydroxyphenyl)-5-(4-pyridinyl)-1H-imidazol-2yl]phenyl...More data for this Ligand-Target Pair
Affinity DataKi: 1.90nMAssay Description:Displacement of rhodamine-green labeled (2-(6-amino-3-imino-3H-xanthen-9-yl)-5-{[({4-[4-(4-Cl-3-hydroxyphenyl)-5-(4-pyridinyl)-1H-imidazol-2yl]phenyl...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of human EGFR T790M/del (746 to 750 residues) mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 peptide as subs...More data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 2nMAssay Description:Inhibition of wild type human EGFR assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 as substrate preincubated for 30 mins follow...More data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 2nMAssay Description:Displacement of rhodamine-green labelled (2-(6-amino-3-imino-3H-xanthen-9-yl)-5-{[({4-[4-(4-Cl-3-hydroxyphenyl)-5-(4-pyridinyl)-1H-imidazol-2yl]pheny...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of human EGFR del (746 to 750) mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 as substrate preincubated for ...More data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 2.10nMAssay Description:Inhibition of human EGFR T790M/L858R double mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 as peptide substrate preincu...More data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 2.10nMAssay Description:Displacement of rhodamine-green labeled (2-(6-amino-3-imino-3H-xanthen-9-yl)-5-{[({4-[4-(4-Cl-3-hydroxyphenyl)-5-(4-pyridinyl)-1H-imidazol-2yl]phenyl...More data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:Displacement of rhodamine-green labeled (2-(6-amino-3-imino-3H-xanthen-9-yl)-5-{[({4-[4-(4-Cl-3-hydroxyphenyl)-5-(4-pyridinyl)-1H-imidazol-2yl]phenyl...More data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of rhodamine-green labeled (2-(6-amino-3-imino-3H-xanthen-9-yl)-5-{[({4-[4-(4-Cl-3-hydroxyphenyl)-5-(4-pyridinyl)-1H-imidazol-2yl]phenyl...More data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of rhodamine-green labelled (2-(6-amino-3-imino-3H-xanthen-9-yl)-5-{[({4-[4-(4-Cl-3-hydroxyphenyl)-5-(4-pyridinyl)-1H-imidazol-2yl]pheny...More data for this Ligand-Target Pair
Affinity DataKi: 3.40nMAssay Description:Inhibition of wild type human EGFR assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 as substrate preincubated for 30 mins follow...More data for this Ligand-Target Pair
Ligand InfoPDB
Affinity DataKi: 3.5nMAssay Description:Inhibition of human EGFR T790M/del (746 to 750 residues) mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 peptide as subs...More data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 3.60nMAssay Description:Inhibition of human EGFR T790M/L858R double mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 as peptide substrate preincu...More data for this Ligand-Target Pair
Affinity DataKi: 3.80nMAssay Description:Inhibition of human EGFR L858R mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 peptide as substrate preincubated for 30 ...More data for this Ligand-Target Pair
Ligand InfoPDB
Affinity DataKi: 4nMAssay Description:Displacement of rhodamine-green labelled (2-(6-amino-3-imino-3H-xanthen-9-yl)-5-{[({4-[4-(4-Cl-3-hydroxyphenyl)-5-(4-pyridinyl)-1H-imidazol-2yl]pheny...More data for this Ligand-Target Pair
Affinity DataKi: 4.10nMAssay Description:Inhibition of human EGFR T790M/L858R double mutant assessed as apparent inhibition constant using Fl-EEPLYWSFPAKKK-CONH2 as peptide substrate preincu...More data for this Ligand-Target Pair
Ligand InfoPDB

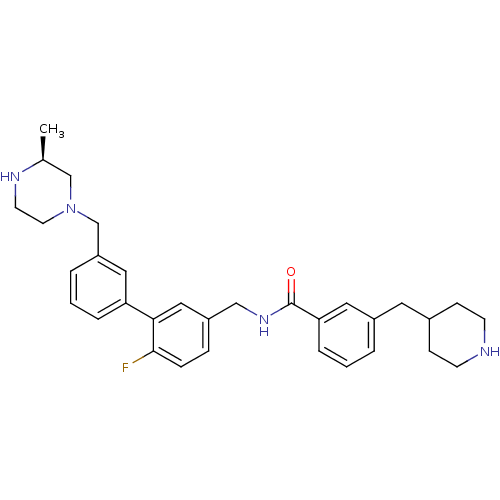
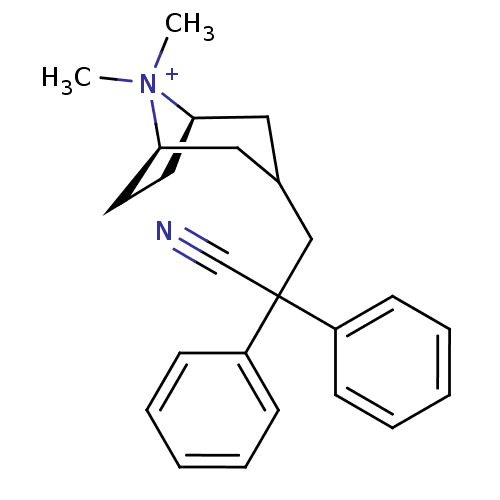
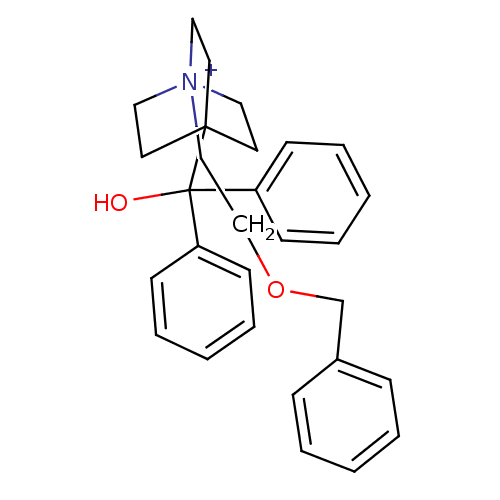
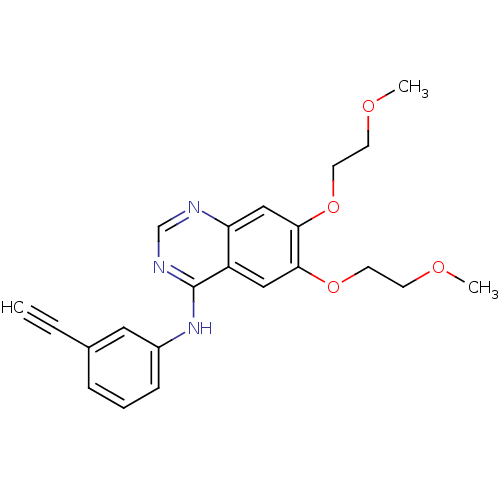
 3D Structure (crystal)
3D Structure (crystal)