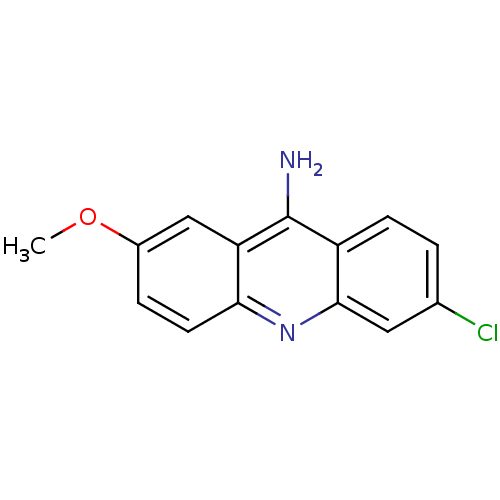
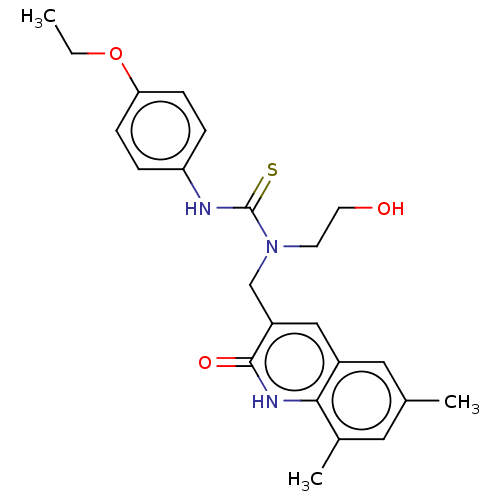
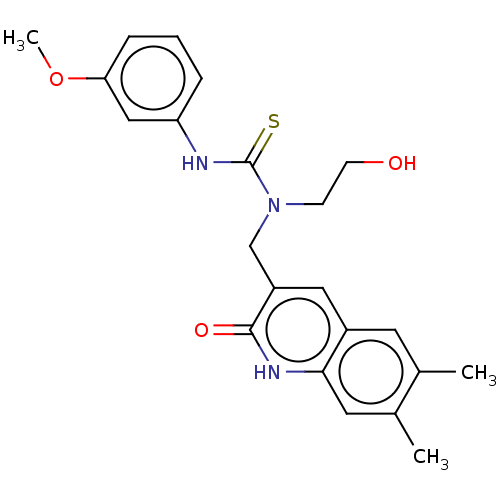
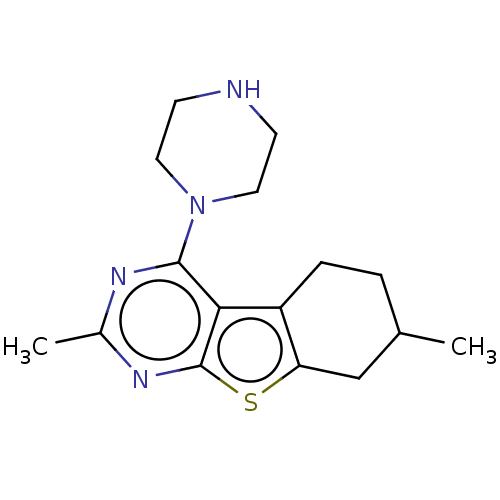
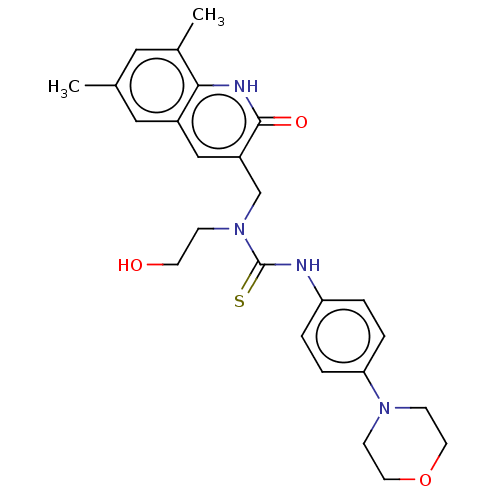
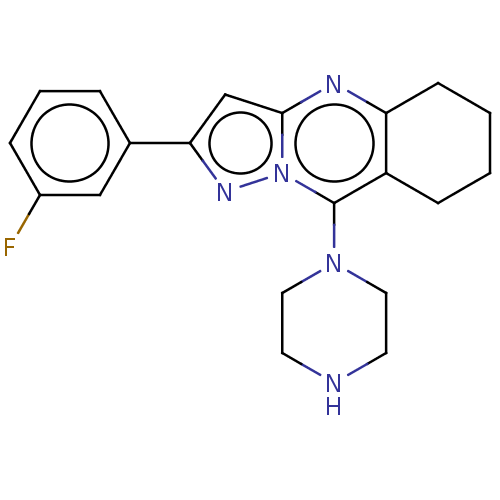
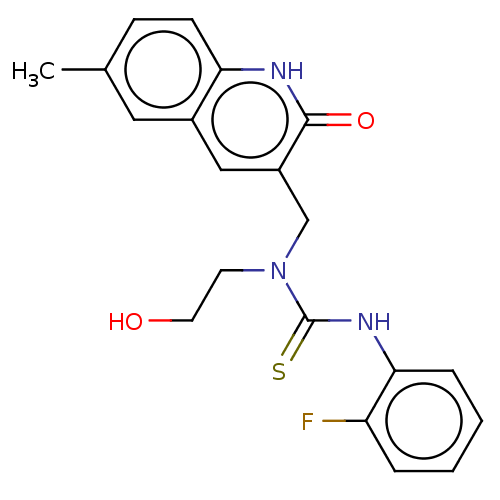
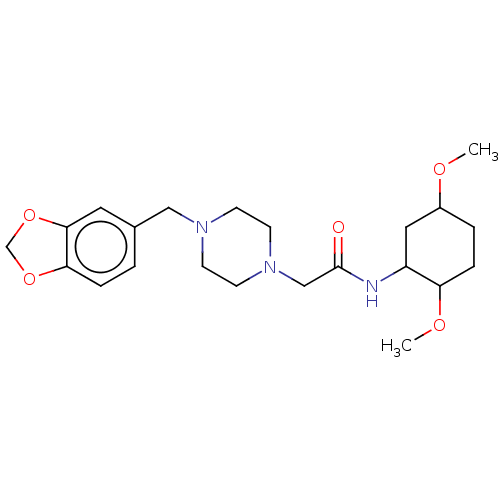
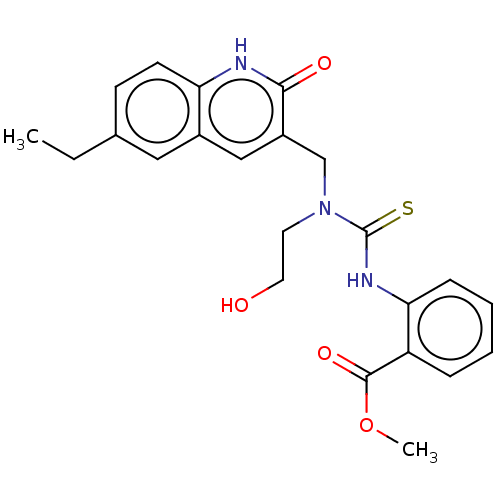
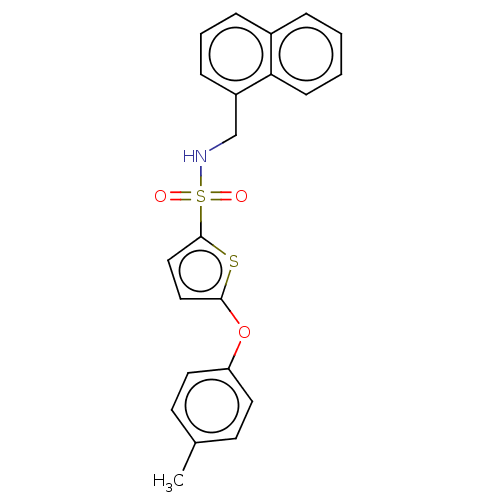
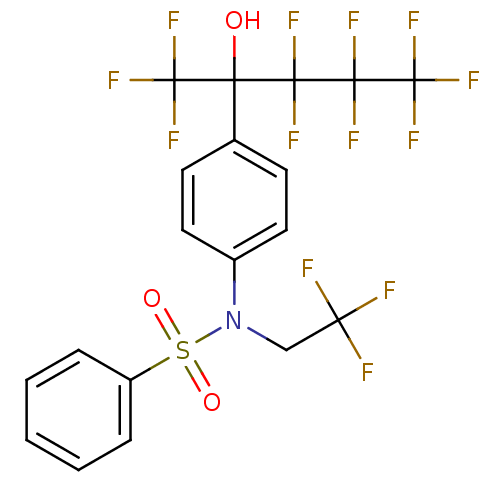
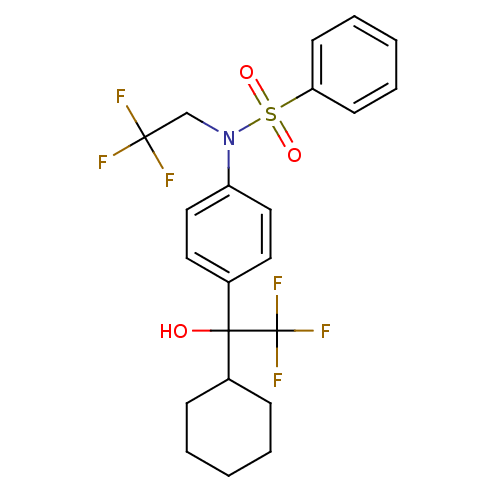
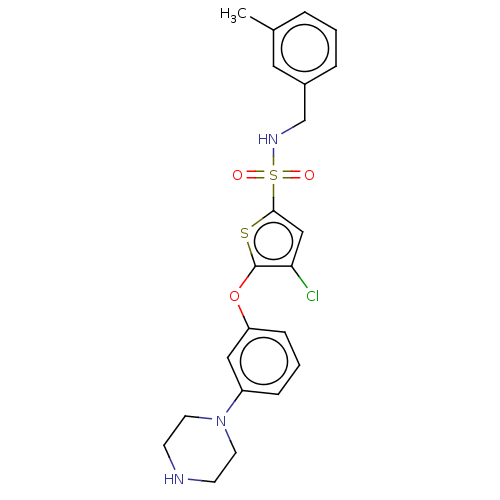
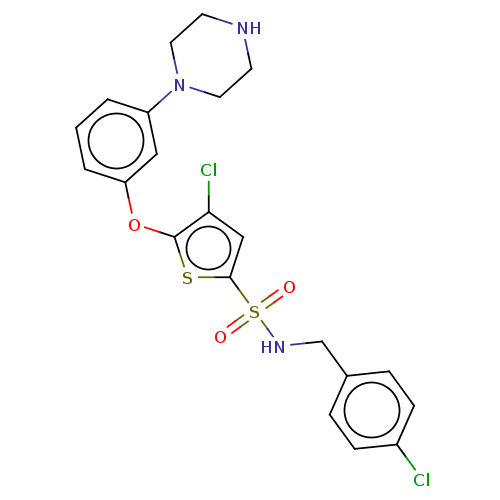
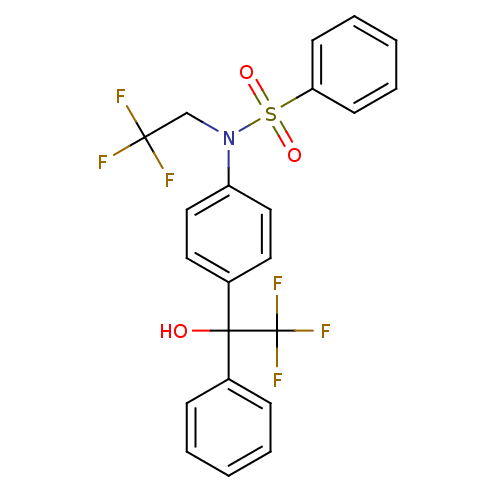
TargetType IV secretion-like conjugative transfer relaxase protein TraI(Escherichia coli (strain SMS-3-5 / SECEC))
University Of North Carolina
Curated by ChEMBL
University Of North Carolina
Curated by ChEMBL
Affinity DataKi: 2nMAssay Description:Inhibition of Escherichia coli F plasmid TraI relaxase Y16F mutant assessed as oriT ssDNA cleavage by competitive inhibition assayMore data for this Ligand-Target Pair
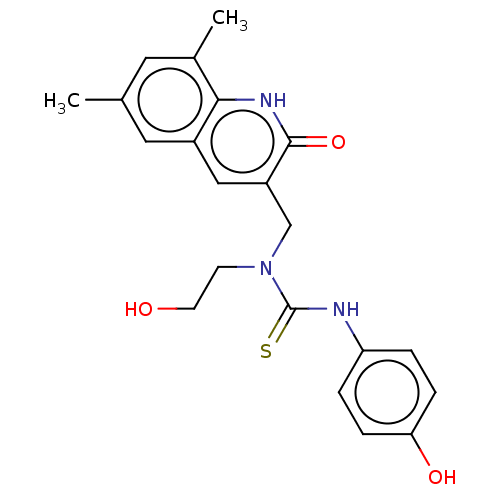
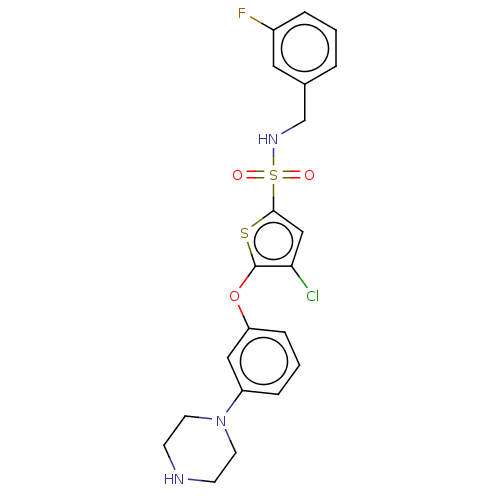
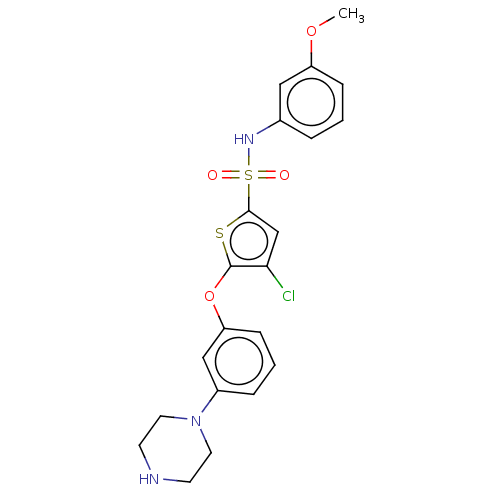
TargetType IV secretion-like conjugative transfer relaxase protein TraI(Escherichia coli (strain SMS-3-5 / SECEC))
University Of North Carolina
Curated by ChEMBL
University Of North Carolina
Curated by ChEMBL
Affinity DataKi: 2.70nMAssay Description:Inhibition of Escherichia coli F plasmid TraI relaxase Y16F mutant assessed as oriT ssDNA cleavage by uncompetitive inhibition assayMore data for this Ligand-Target Pair
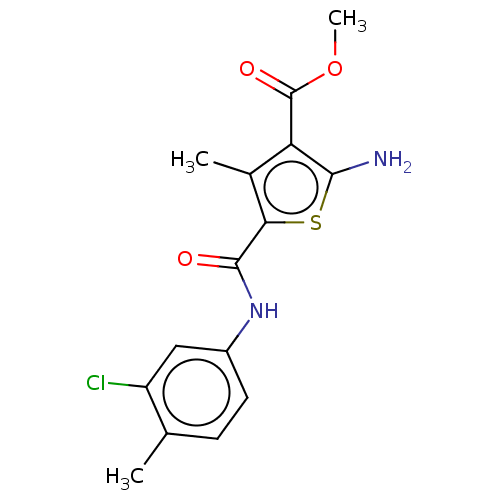
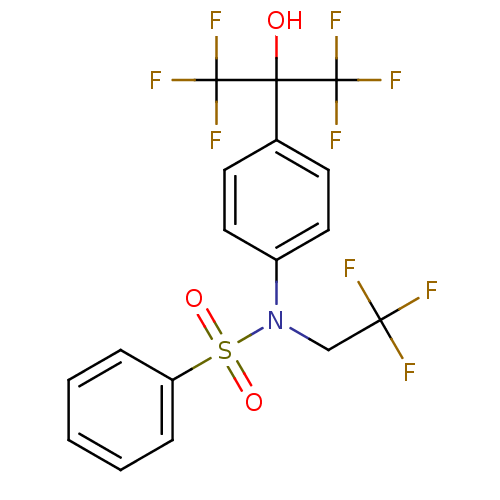
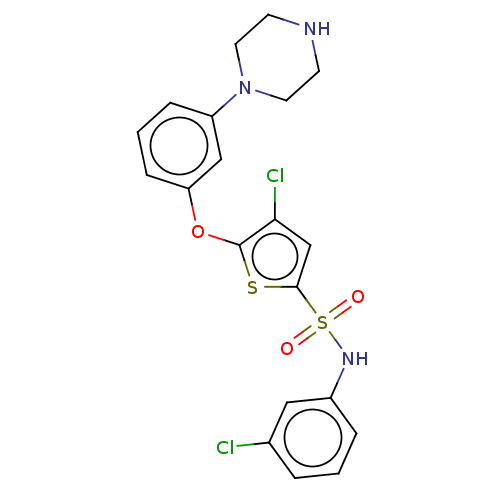
TargetType IV secretion-like conjugative transfer relaxase protein TraI(Escherichia coli (strain SMS-3-5 / SECEC))
University Of North Carolina
Curated by ChEMBL
University Of North Carolina
Curated by ChEMBL
Affinity DataKi: 3nMAssay Description:Inhibition of Escherichia coli F plasmid TraI relaxase Y16F mutant assessed as oriT ssDNA cleavage by competitive inhibition assayMore data for this Ligand-Target Pair
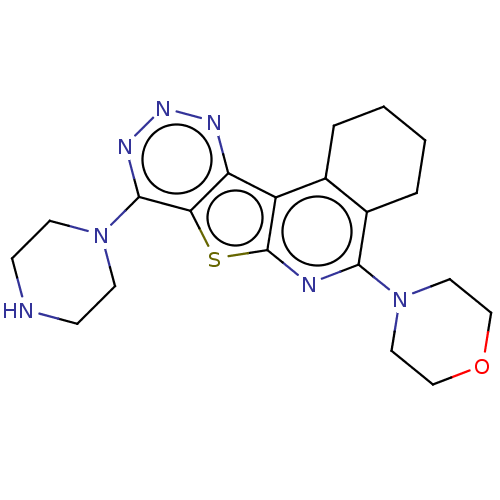
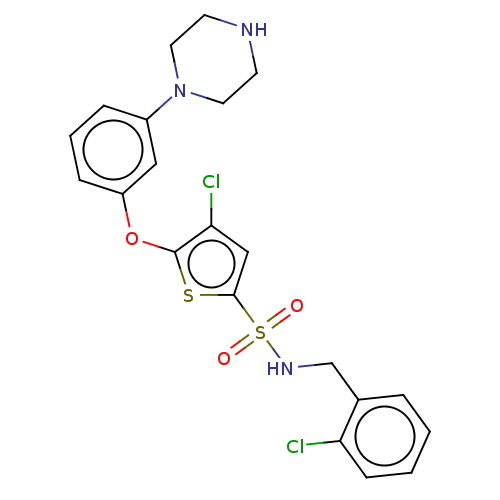
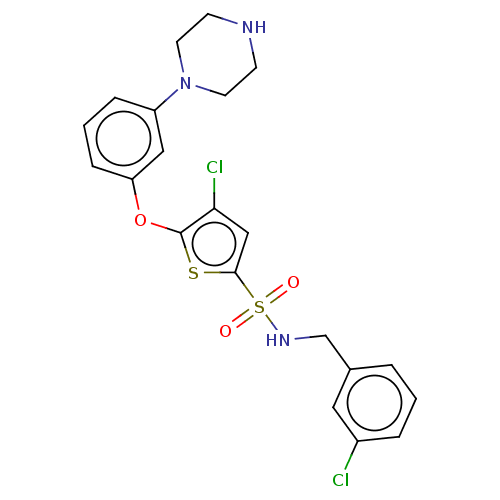
TargetType IV secretion-like conjugative transfer relaxase protein TraI(Escherichia coli (strain SMS-3-5 / SECEC))
University Of North Carolina
Curated by ChEMBL
University Of North Carolina
Curated by ChEMBL
Affinity DataKi: 3nMAssay Description:Inhibition of Escherichia coli F plasmid TraI relaxase Y16F mutant assessed as oriT ssDNA cleavage by competitive inhibition assayMore data for this Ligand-Target Pair
TargetType IV secretion-like conjugative transfer relaxase protein TraI(Escherichia coli (strain SMS-3-5 / SECEC))
University Of North Carolina
Curated by ChEMBL
University Of North Carolina
Curated by ChEMBL
Affinity DataKi: 3nMAssay Description:Inhibition of Escherichia coli F plasmid TraI relaxase Y16F mutant assessed as oriT ssDNA cleavage by competitive inhibition assayMore data for this Ligand-Target Pair
Affinity DataKi: 6.90nM ΔG°: -46.1kJ/molepH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Estimates of the competitive inhibition constants (Ki) were ob...More data for this Ligand-Target Pair
Affinity DataKi: 49nM ΔG°: -41.3kJ/molepH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Estimates of the competitive inhibition constants (Ki) were ob...More data for this Ligand-Target Pair
TargetBeta-glucuronidase(Escherichia coli (Enterobacteria))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 160nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-glucuronidase(Escherichia coli (Enterobacteria))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 210nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-glucuronidase(Escherichia coli (Enterobacteria))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 220nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
Affinity DataKi: 490nM ΔG°: -35.7kJ/molepH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Estimates of the competitive inhibition constants (Ki) were ob...More data for this Ligand-Target Pair
TargetBeta-galactosidase [V151I,I185V](Clostridium perfringens (Firmicutes))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 540nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-glucuronidase(Escherichia coli (Enterobacteria))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 610nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-glucuronidase(Escherichia coli (Enterobacteria))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 670nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-glucuronidase(Escherichia coli (Enterobacteria))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 680nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-galactosidase(Streptococcus agalactiae (Firmicutes))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 810nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-glucuronidase(Escherichia coli (Enterobacteria))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 960nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-galactosidase [V151I,I185V](Clostridium perfringens (Firmicutes))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 970nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-galactosidase [V151I,I185V](Clostridium perfringens (Firmicutes))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 1.10E+3nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-galactosidase(Streptococcus agalactiae (Firmicutes))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 1.40E+3nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-glucuronidase(Escherichia coli (Enterobacteria))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 1.40E+3nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-glucuronidase(Escherichia coli (Enterobacteria))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 1.90E+3nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-glucuronidase(Escherichia coli (Enterobacteria))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 1.90E+3nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-galactosidase(Streptococcus agalactiae (Firmicutes))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 2.80E+3nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-galactosidase(Streptococcus agalactiae (Firmicutes))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 3.00E+3nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
Affinity DataKi: 5.80E+3nM ΔG°: -29.6kJ/molepH: 7.4 T: 2°CAssay Description:Inhibition constants were calculated by assessment of the reduction in the formation of o-nitrophenol, as monitored by a spectrophotometric assay at ...More data for this Ligand-Target Pair
TargetBeta-galactosidase [V151I,I185V](Clostridium perfringens (Firmicutes))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 6.10E+3nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-galactosidase [V151I,I185V](Clostridium perfringens (Firmicutes))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 7.80E+3nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-galactosidase(Streptococcus agalactiae (Firmicutes))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 1.10E+4nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
Affinity DataKi: 1.71E+4nM ΔG°: -26.9kJ/molepH: 7.4 T: 2°CAssay Description:Inhibition constants were calculated by assessment of the reduction in the formation of o-nitrophenol, as monitored by a spectrophotometric assay at ...More data for this Ligand-Target Pair
TargetBeta-galactosidase [V151I,I185V](Clostridium perfringens (Firmicutes))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 2.40E+4nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-galactosidase(Streptococcus agalactiae (Firmicutes))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataKi: 3.60E+4nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 �L assay buffer, 5 �L inhibitor solution, 5 �L of 10 nM enz...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nM ΔG°: >-22.6kJ/molepH: 7.4 T: 2°CAssay Description:Inhibition constants were calculated by assessment of the reduction in the formation of o-nitrophenol, as monitored by a spectrophotometric assay at ...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nM ΔG°: >-22.6kJ/molepH: 7.4 T: 2°CAssay Description:Inhibition constants were calculated by assessment of the reduction in the formation of o-nitrophenol, as monitored by a spectrophotometric assay at ...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nM ΔG°: >-22.6kJ/molepH: 7.4 T: 2°CAssay Description:Inhibition constants were calculated by assessment of the reduction in the formation of o-nitrophenol, as monitored by a spectrophotometric assay at ...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nM ΔG°: >-22.6kJ/molepH: 7.4 T: 2°CAssay Description:Inhibition constants were calculated by assessment of the reduction in the formation of o-nitrophenol, as monitored by a spectrophotometric assay at ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.984nMAssay Description:PheG Assay: Two �-glucuronidase activity assays were employed to examine the potency of inhibitors both in vitro and in cell-based studies. An absorb...More data for this Ligand-Target Pair
Affinity DataIC50: <1nMAssay Description:The β-glucuronidase assay was performed by the addition of 0.5 μl of compound (or DMSO) to the well of a black 384-well plate followed by t...More data for this Ligand-Target Pair
Affinity DataIC50: 1.17nMAssay Description:PNPG Assay: Two �-glucuronidase activity assays were employed to examine the potency of inhibitors both in vitro and in cell-based studies. An absorb...More data for this Ligand-Target Pair
TargetNuclear receptor subfamily 1 group I member 2(Homo sapiens (Human))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataIC50: 20nM EC50: 10nMpH: 8.0 T: 2°CAssay Description:Polylysine YiO imaging beads (Amersham, GE Healthcare) were coated with histidine-tagged WT PXR LBD. Nonspecific binding sites were blocked with BSA....More data for this Ligand-Target Pair
TargetNuclear receptor subfamily 1 group I member 2(Homo sapiens (Human))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataIC50: 25nM EC50: 3nMpH: 8.0 T: 2°CAssay Description:Polylysine YiO imaging beads (Amersham, GE Healthcare) were coated with histidine-tagged WT PXR LBD. Nonspecific binding sites were blocked with BSA....More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:The β-glucuronidase assay was performed by the addition of 0.5 μl of compound (or DMSO) to the well of a black 384-well plate followed by t...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:The β-glucuronidase assay was performed by the addition of 0.5 μl of compound (or DMSO) to the well of a black 384-well plate followed by t...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:The β-glucuronidase assay was performed by the addition of 0.5 μl of compound (or DMSO) to the well of a black 384-well plate followed by t...More data for this Ligand-Target Pair
TargetNuclear receptor subfamily 1 group I member 2(Homo sapiens (Human))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataIC50: 40nM EC50: 13nMpH: 8.0 T: 2°CAssay Description:Polylysine YiO imaging beads (Amersham, GE Healthcare) were coated with histidine-tagged WT PXR LBD. Nonspecific binding sites were blocked with BSA....More data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:The β-glucuronidase assay was performed by the addition of 0.5 μl of compound (or DMSO) to the well of a black 384-well plate followed by t...More data for this Ligand-Target Pair
TargetNuclear receptor subfamily 1 group I member 2(Homo sapiens (Human))
University of North Carolina At Chapel Hill
University of North Carolina At Chapel Hill
Affinity DataIC50: 63nM EC50: 16nMpH: 8.0 T: 2°CAssay Description:Polylysine YiO imaging beads (Amersham, GE Healthcare) were coated with histidine-tagged WT PXR LBD. Nonspecific binding sites were blocked with BSA....More data for this Ligand-Target Pair
Affinity DataIC50: 70nMAssay Description:The β-glucuronidase assay was performed by the addition of 0.5 μl of compound (or DMSO) to the well of a black 384-well plate followed by t...More data for this Ligand-Target Pair
Affinity DataIC50: 70nMAssay Description:The β-glucuronidase assay was performed by the addition of 0.5 μl of compound (or DMSO) to the well of a black 384-well plate followed by t...More data for this Ligand-Target Pair
Affinity DataIC50: 90nMAssay Description:The β-glucuronidase assay was performed by the addition of 0.5 μl of compound (or DMSO) to the well of a black 384-well plate followed by t...More data for this Ligand-Target Pair

 3D Structure (crystal)
3D Structure (crystal)