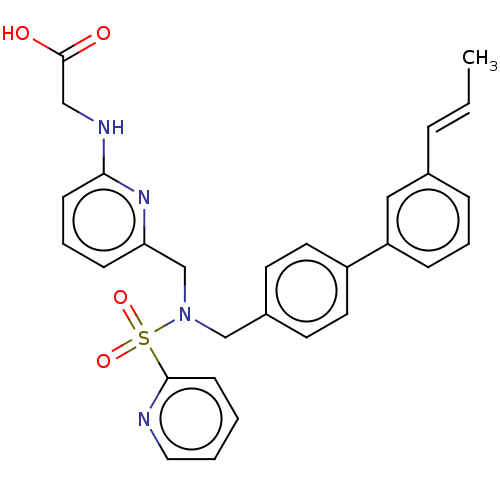
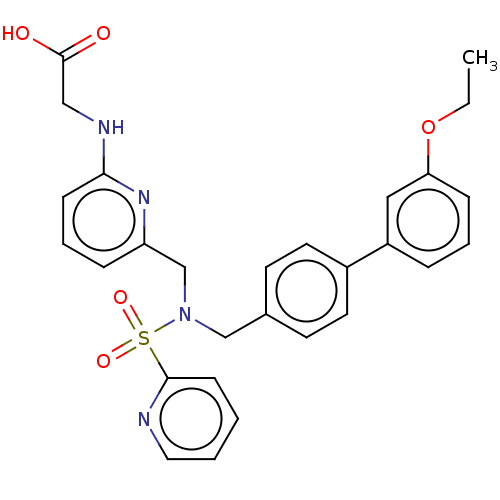
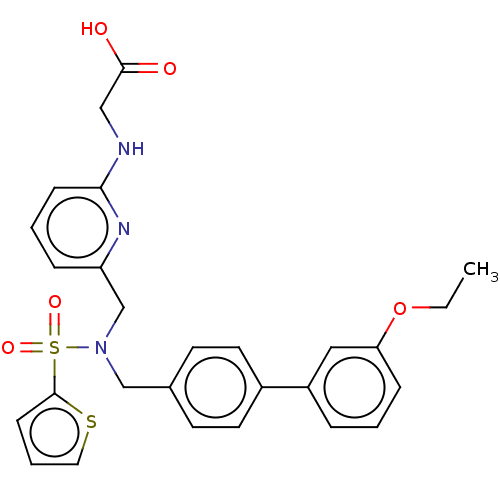
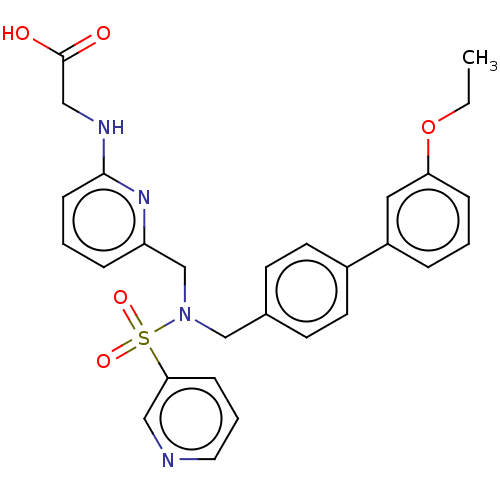
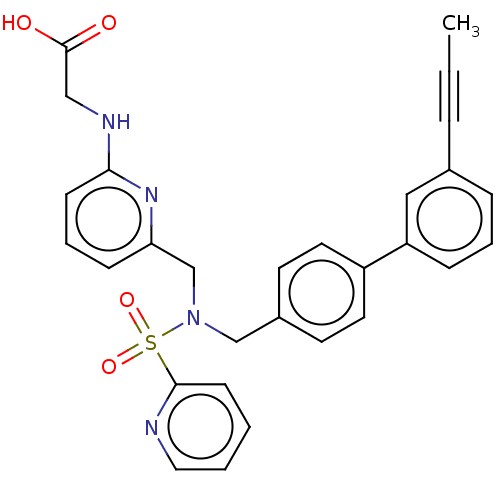
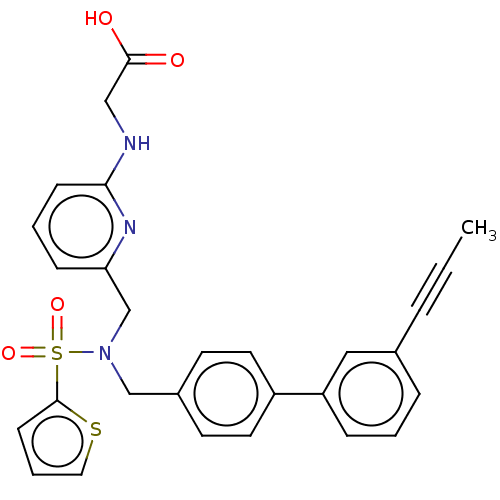
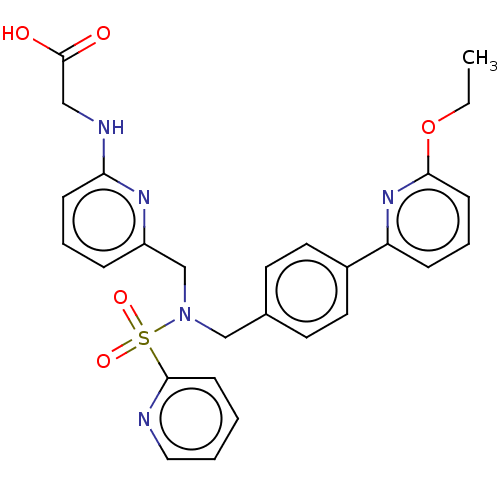
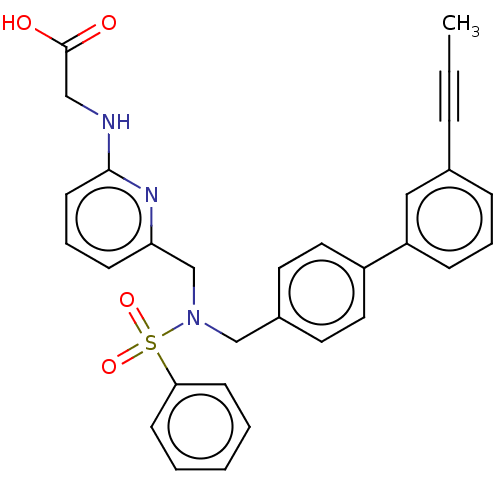
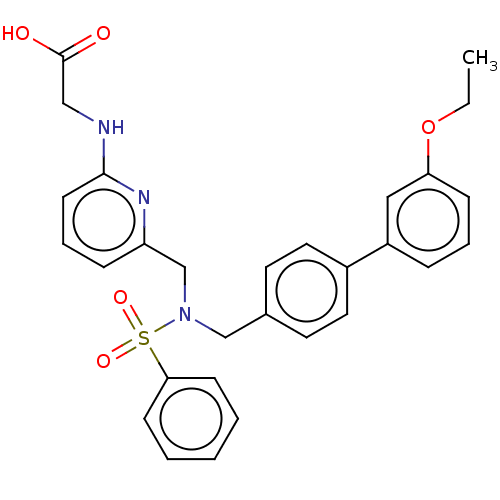
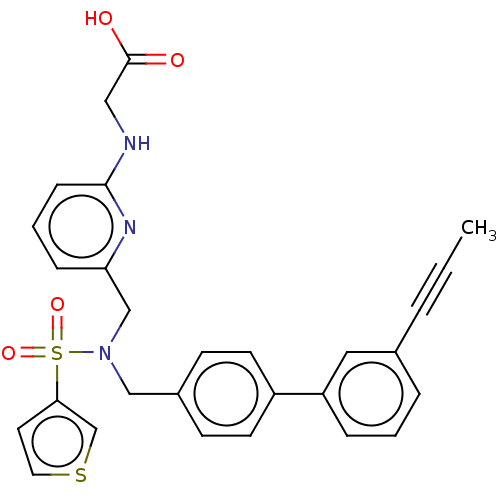
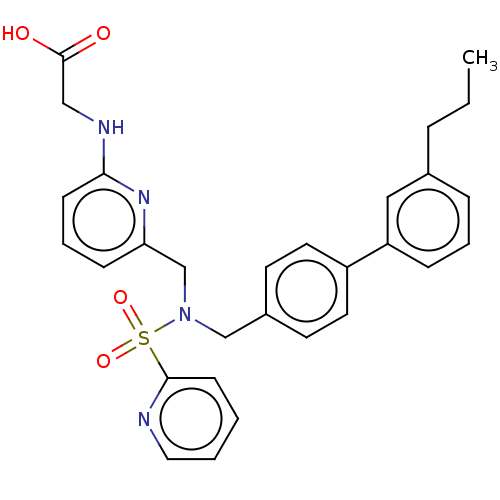
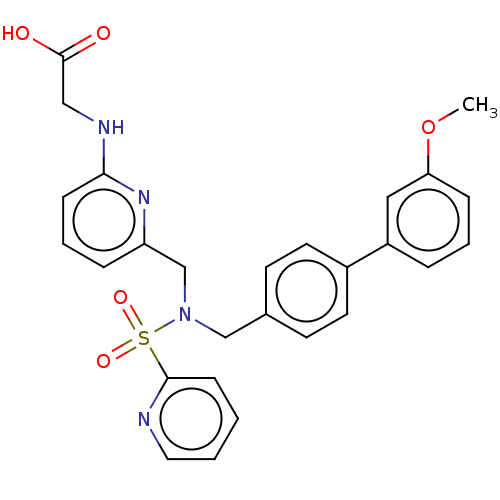
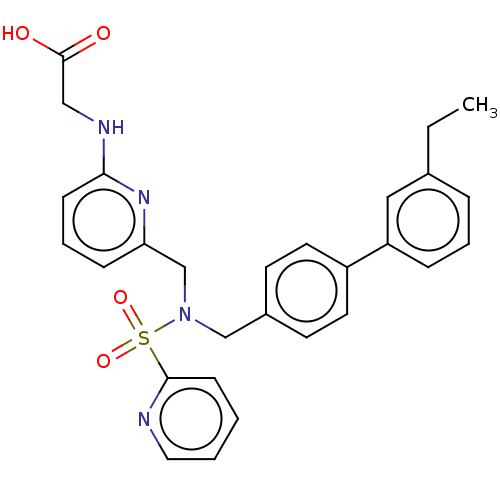
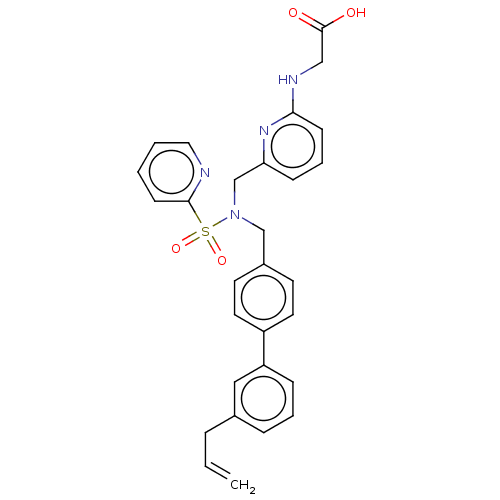
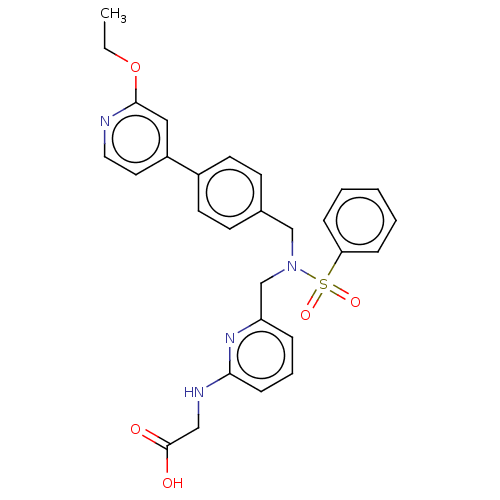
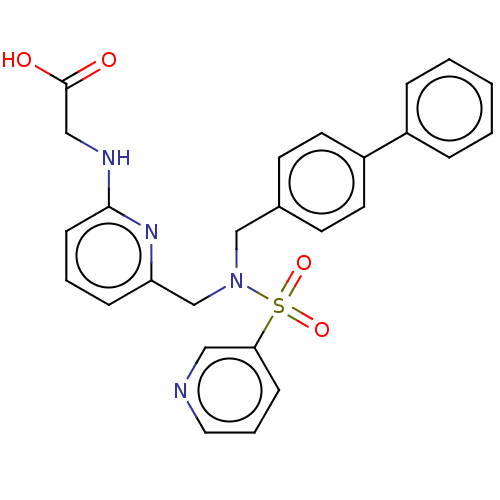
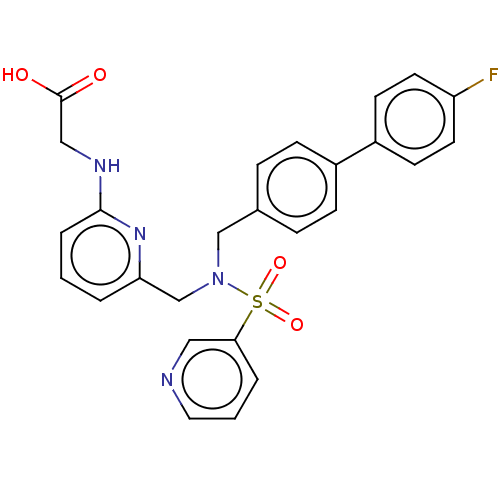
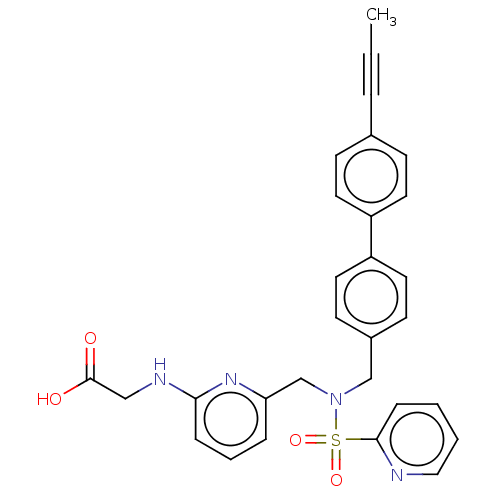
Affinity DataKi: 0.530nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
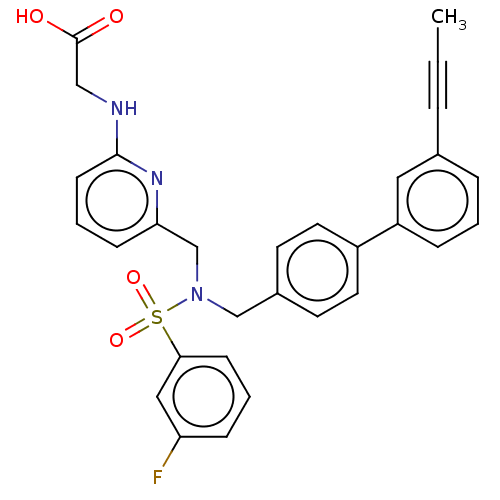
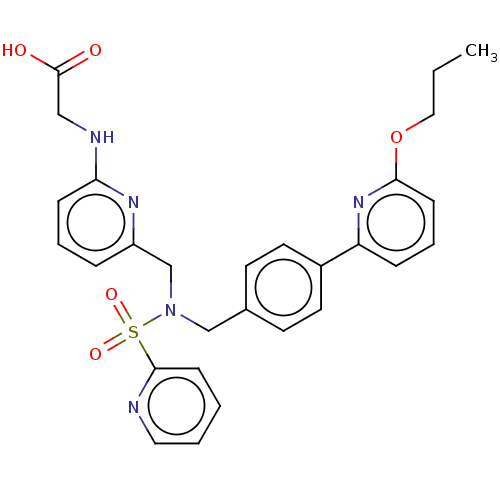
Affinity DataKi: 0.610nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
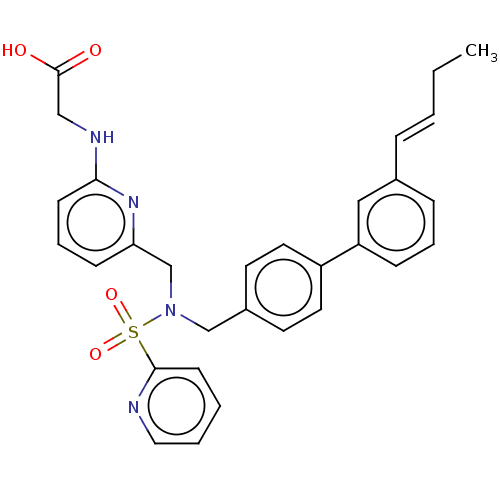
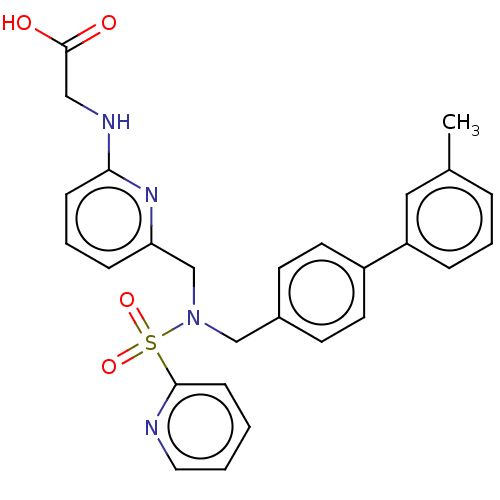
Affinity DataKi: 0.75nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
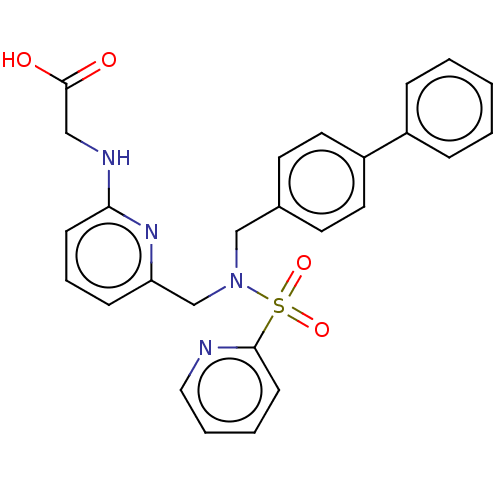
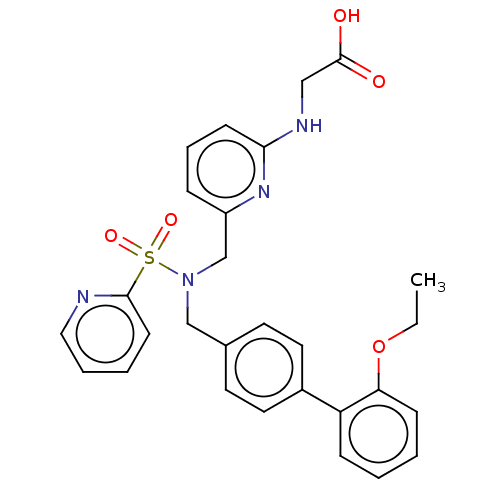
Affinity DataKi: 0.75nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 0.790nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 0.900nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 0.940nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 0.950nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 0.970nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 0.990nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 1.10nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 1.20nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 1.40nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 1.40nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 1.70nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 2nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 2.20nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 2.60nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 3.20nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 3.80nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 4.40nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 6.80nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 9.80nMAssay Description:Inhibition of [3H]-PDBu binding to PKCdelta C1B domain (unknown origin) by competitive binding assayMore data for this Ligand-Target Pair
Affinity DataKi: >10nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: >10nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: >10nMpH: 6.0Assay Description:Measurement of EP2 receptor binding action was performed in accordance with the method of Abramovitz et al. (Biochimica et Biophysica Acta, 1483, 285...More data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Inhibition of [3H]-PDBu binding to PKCdelta C1B domain (unknown origin) by competitive binding assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 5(Homo sapiens (Human))
Universit£
Curated by ChEMBL
Universit£
Curated by ChEMBL
Affinity DataKi: 77nMAssay Description:Inhibition of human recombinant ADAMTS5 using bovine nasal cartilage aggrecan as substrate assessed as inhibition of 1772-AGEG neopeptide formation i...More data for this Ligand-Target Pair
Affinity DataKi: 88nMAssay Description:Inhibition of [3H]-PDBu binding to PKCalpha C1A domain (unknown origin) by competitive binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 100nMAssay Description:Inhibition of [3H]-PDBu binding to PKCalpha C1A domain (unknown origin) by competitive binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 120nMAssay Description:Inhibition of [3H]-PDBu binding to PKCalpha C1A domain (unknown origin) by competitive binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 130nMAssay Description:Inhibition of [3H]-PDBu binding to PKCdelta C1B domain (unknown origin) by competitive binding assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 5(Homo sapiens (Human))
Universit£
Curated by ChEMBL
Universit£
Curated by ChEMBL
Affinity DataKi: 840nMAssay Description:Inhibition of human recombinant ADAMTS5 using bovine nasal cartilage aggrecan as substrate assessed as inhibition of 1772-AGEG neopeptide formation i...More data for this Ligand-Target Pair
Affinity DataKi: 6.80E+3nMAssay Description:Mechanism-based inhibition of recombinant human CYP2C9 expressed in microsomes isolated from Saccharomyces cerevisiae AH22 cells assessed as diclofen...More data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 5(Homo sapiens (Human))
Universit£
Curated by ChEMBL
Universit£
Curated by ChEMBL
Affinity DataKi: 2.10E+4nMAssay Description:Inhibition of human recombinant ADAMTS5 using bovine nasal cartilage aggrecan as substrate assessed as inhibition of 1772-AGEG neopeptide formation i...More data for this Ligand-Target Pair
Affinity DataKi: 4.20E+5nMpH: 4.0Assay Description:Binding affinity to Humicola insolens NCE5 assessed as dissociation constant at pH 4More data for this Ligand-Target Pair
Affinity DataKi: 1.60E+6nMpH: 3.0Assay Description:Binding affinity to Humicola insolens NCE5 assessed as dissociation constant at pH 3.0 and at 59 degC by differential scanning calorimetryMore data for this Ligand-Target Pair
Affinity DataKi: 3.90E+6nMpH: 3.0Assay Description:Binding affinity to Humicola insolens NCE5 assessed as dissociation constant at pH 3.0 and at 59 degC by differential scanning calorimetryMore data for this Ligand-Target Pair
Affinity DataKi: 2.50E+7nMpH: 3.0Assay Description:Binding affinity to Humicola insolens NCE5 assessed as dissociation constant at pH 3.0 and at 59 degC by differential scanning calorimetryMore data for this Ligand-Target Pair
Affinity DataKi: 2.90E+7nMpH: 3.0Assay Description:Binding affinity to Humicola insolens NCE5 assessed as dissociation constant at pH 3.0 and at 59 degC by differential scanning calorimetryMore data for this Ligand-Target Pair
Affinity DataKi: >3.00E+7nMpH: 3.0Assay Description:Binding affinity to Humicola insolens NCE5 assessed as dissociation constant at pH 3.0 and at 59 degC by differential scanning calorimetryMore data for this Ligand-Target Pair
Affinity DataKi: >3.00E+7nMpH: 3.0Assay Description:Binding affinity to Humicola insolens NCE5 assessed as dissociation constant at pH 3.0 and at 59 degC by differential scanning calorimetryMore data for this Ligand-Target Pair
Affinity DataIC50: 0.00200nMAssay Description:Inhibition of human activated TAFI using Hip-Arg as substrate incubated for 10 mins prior to substrate addition measured after 30 mins by spectrophot...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0250nMAssay Description:Inhibition of human activated TAFI using Hip-Arg as substrate incubated for 10 mins prior to substrate addition measured after 30 mins by spectrophot...More data for this Ligand-Target Pair
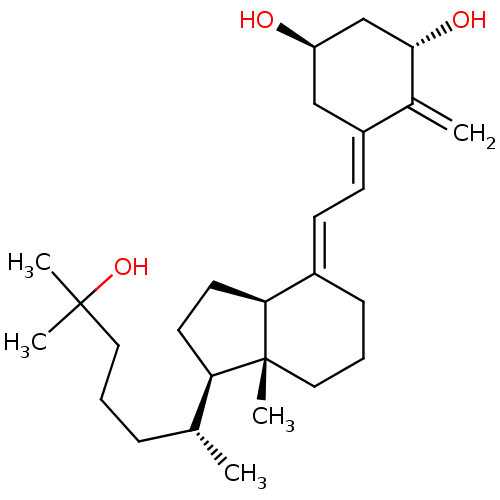
Affinity DataIC50: 0.0290nMAssay Description:Displacement of [3H]1alpha,25-dihydroxyvitamin D3 from human recombinant GST-tagged vitamin D3 receptor LBD expressed in Escherichia coli BL21 after ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0400nMAssay Description:Displacement of [3H]-1,25D3 from C-terminal GST-tagged human recombinant VDR LBD expressed in Escherichia coli BL21 (DE3) after 16 hrs by radioligand...More data for this Ligand-Target Pair