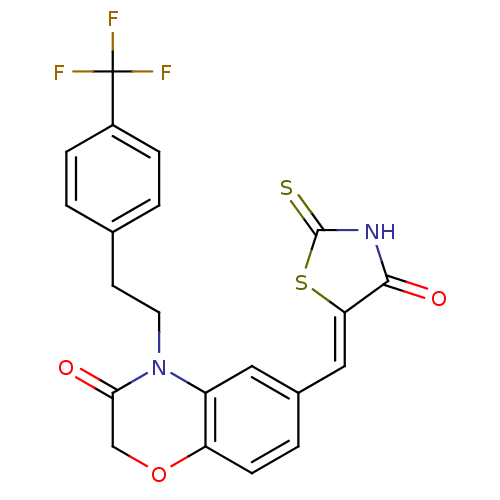
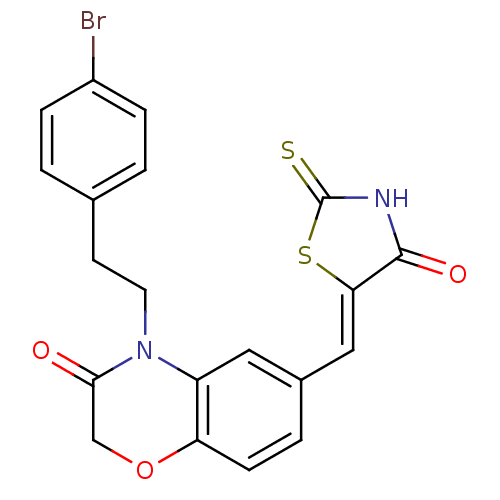
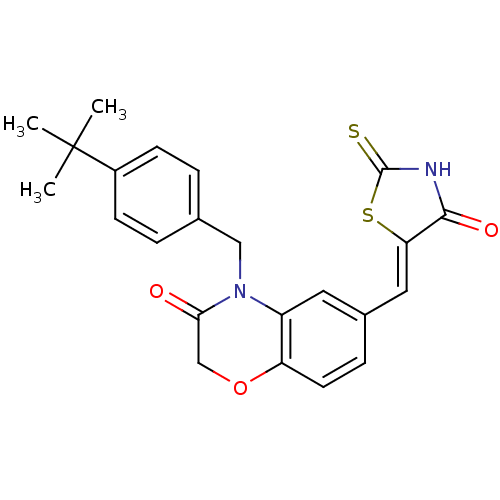
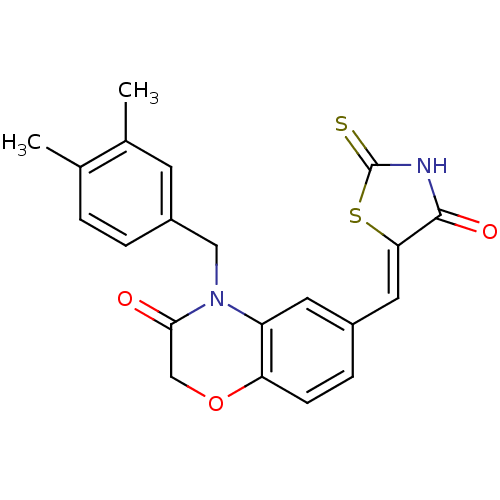
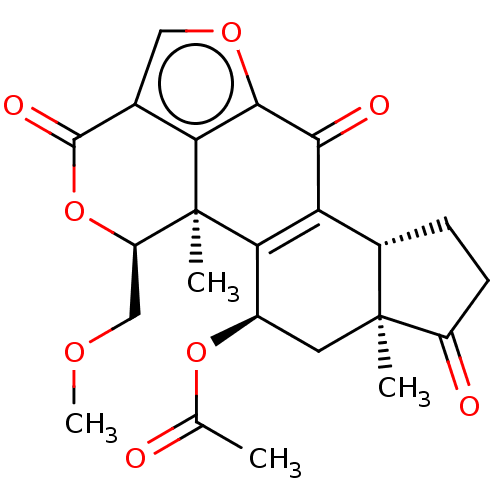
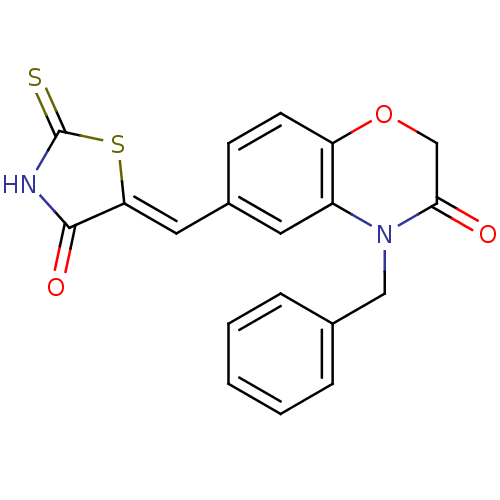
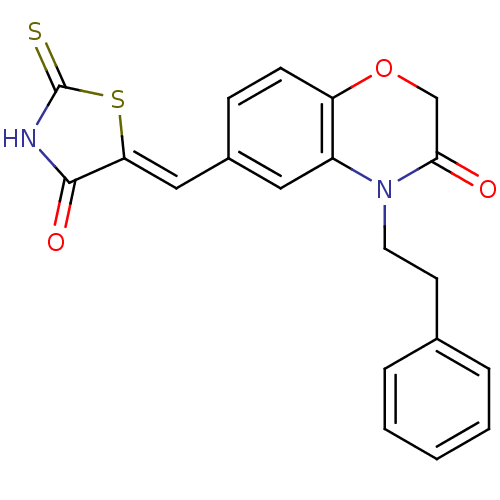
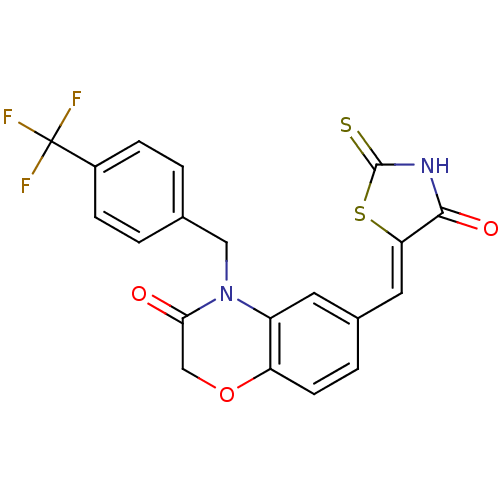
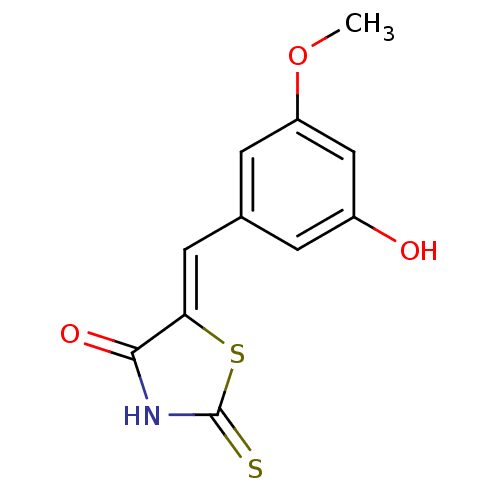
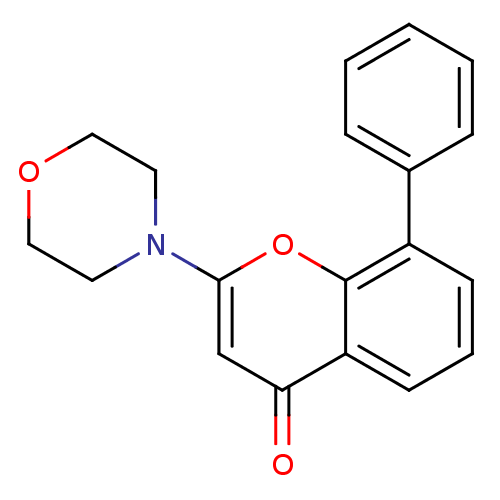
Report error Found 26 Enz. Inhib. hit(s) with all data for entry = 2825
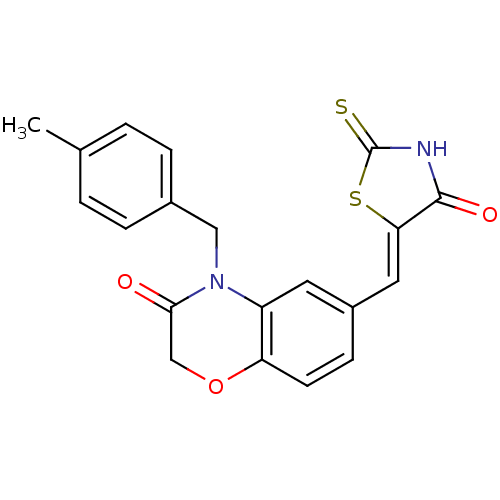
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
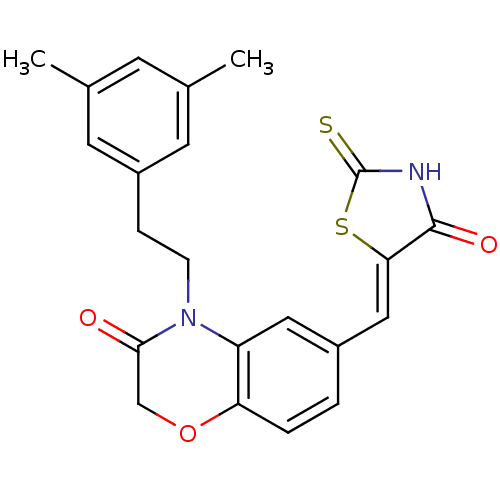
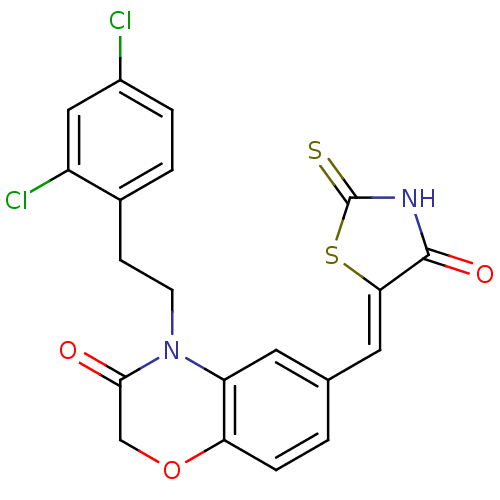
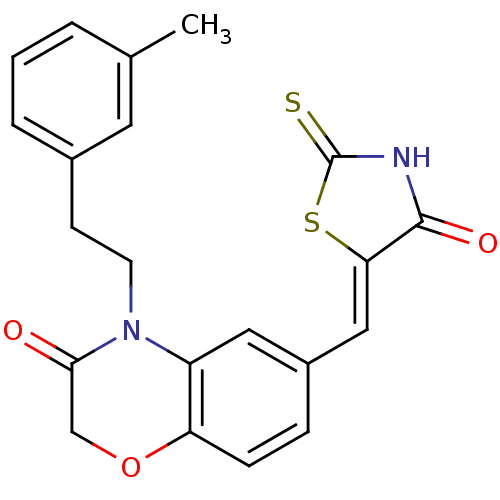
Affinity DataIC50: 1.92nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
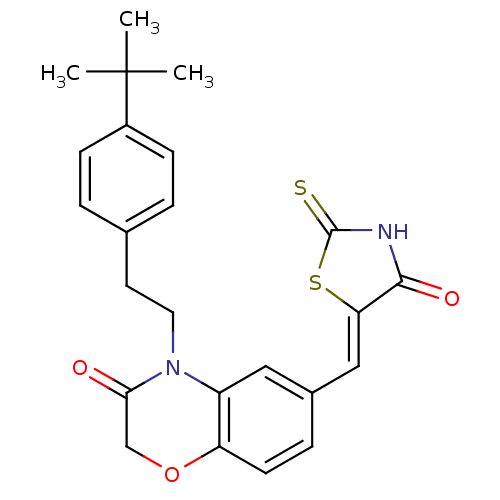
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
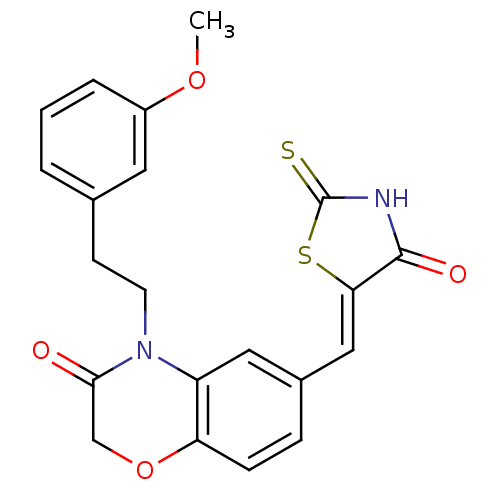
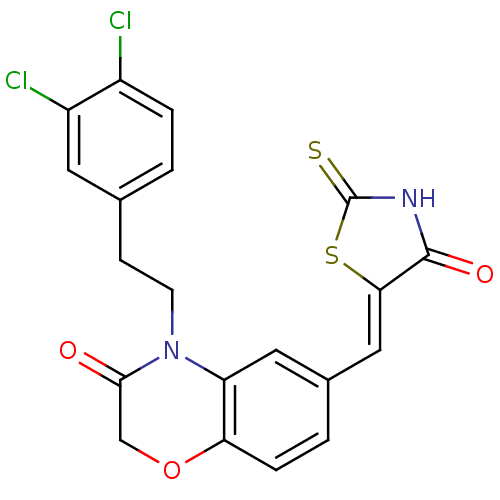
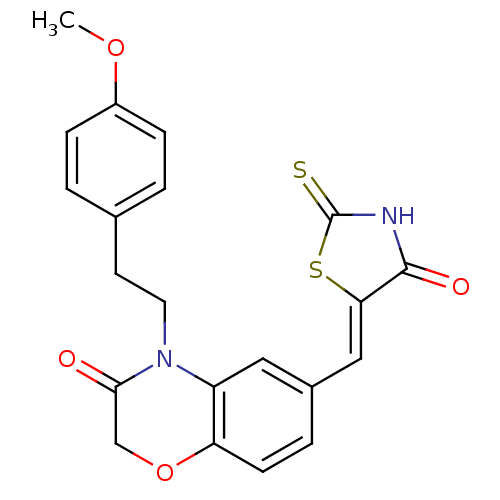
Affinity DataIC50: 2.34nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
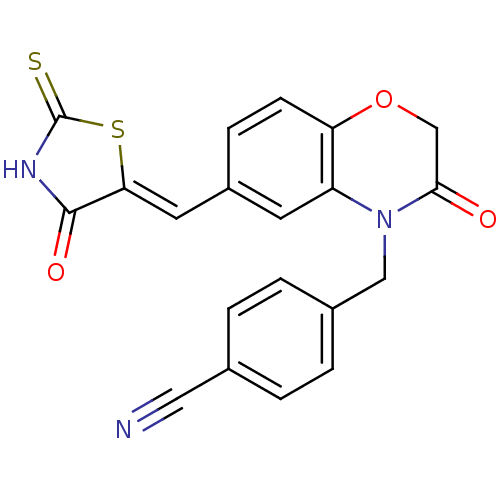
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
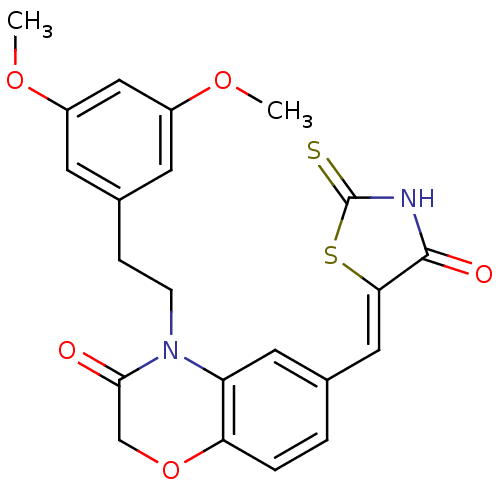
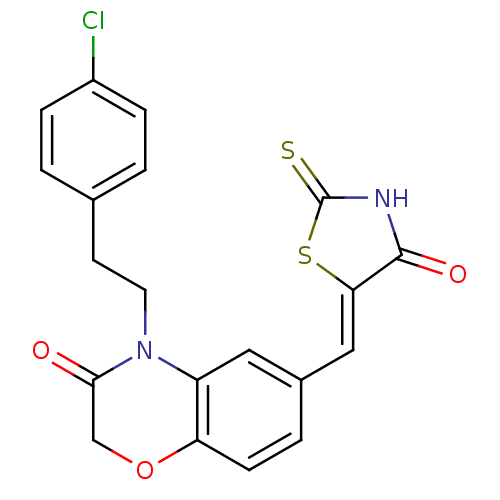
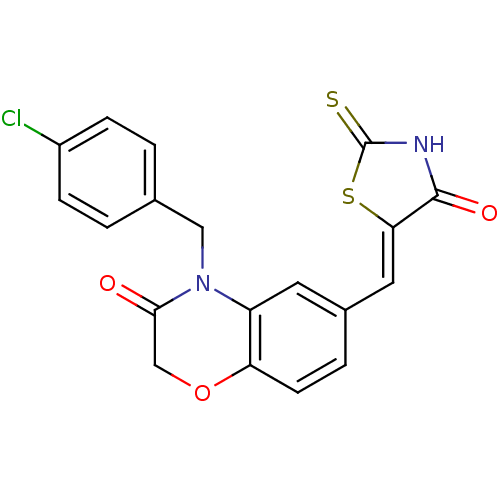
Affinity DataIC50: 2.42nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
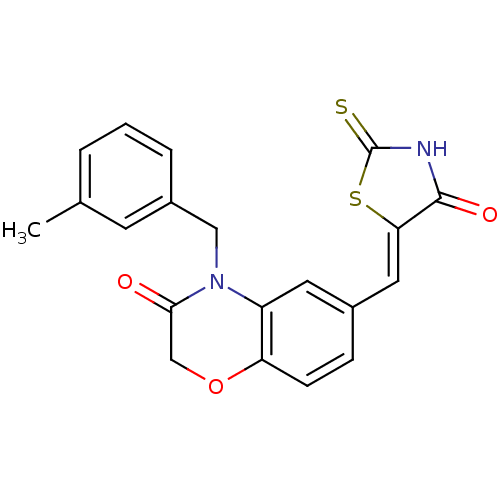
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
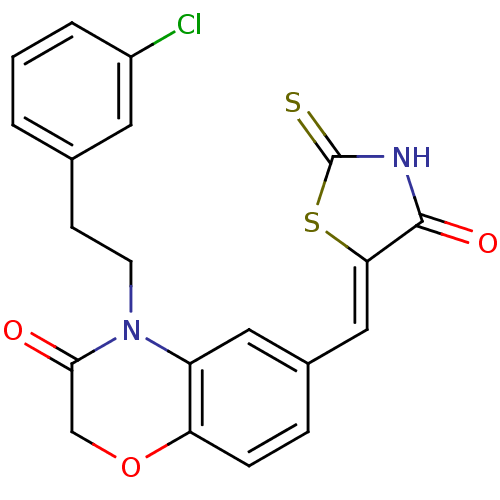
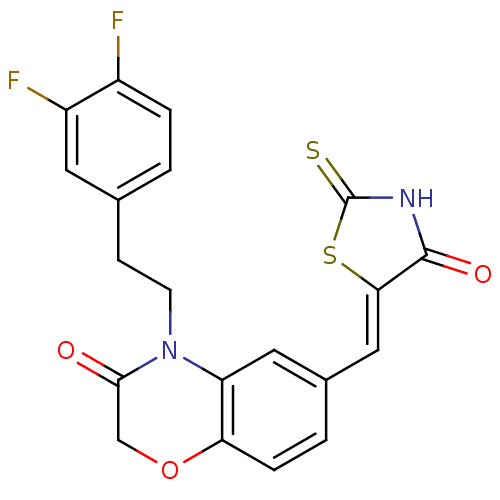
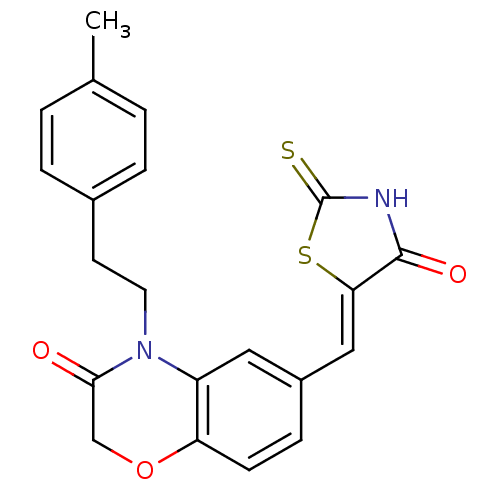
Affinity DataIC50: 2.59nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 2.65nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 2.66nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 2.79nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 4.27nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 4.28nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 4.36nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 4.76nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 4.84nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 4.96nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 6.57nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 7.16nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 8.83nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 19nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 23nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 24.5nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 27.5nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 29.5nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 33.2nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 36.1nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 43.7nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 77.7nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform/Phosphoinositide 3-kinase regulatory subunit 5(Human)
Pfizer
Pfizer
Affinity DataIC50: 1.72E+3nMpH: 7.4 T: 2°CAssay Description:PI3Kgamma activity was assessed by incubation of baculoviral co-expressed regulatory and catalytic subunits (p101 and p110) with lipid micelles prepa...More data for this Ligand-Target Pair