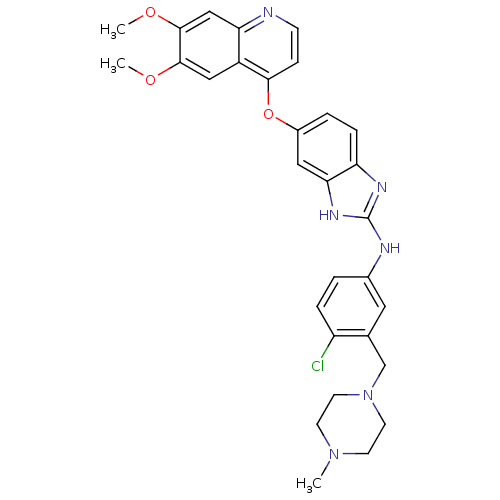
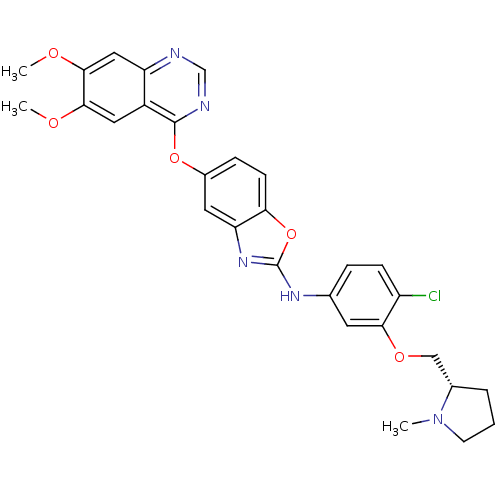
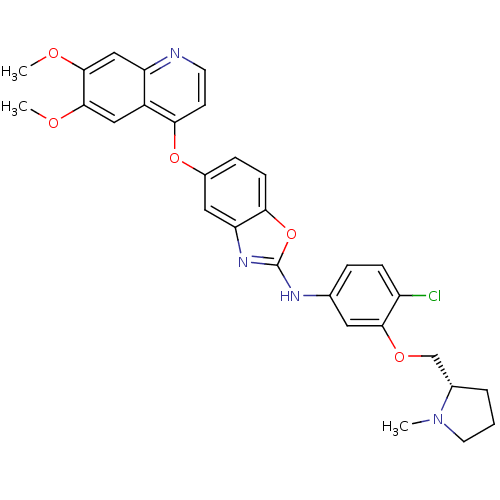
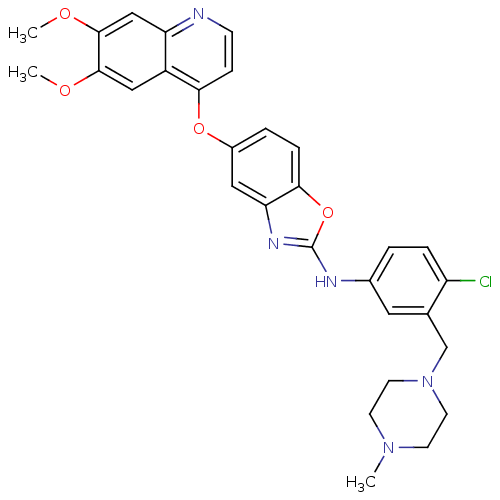
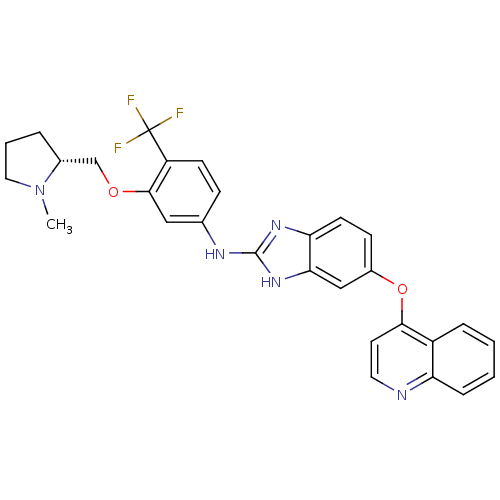
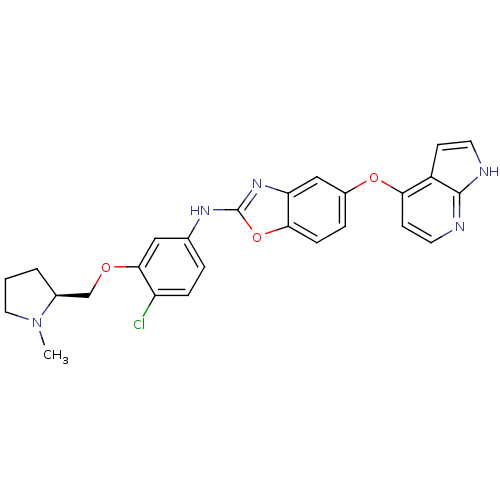
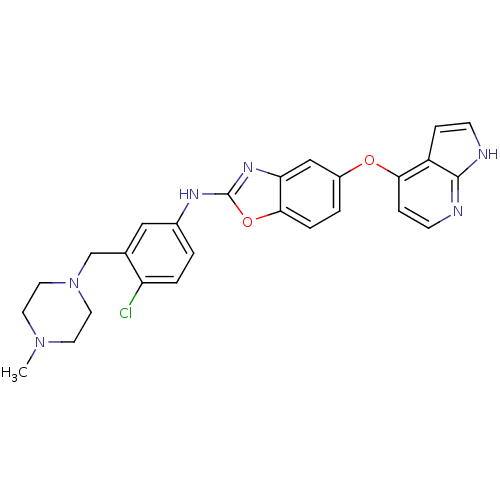
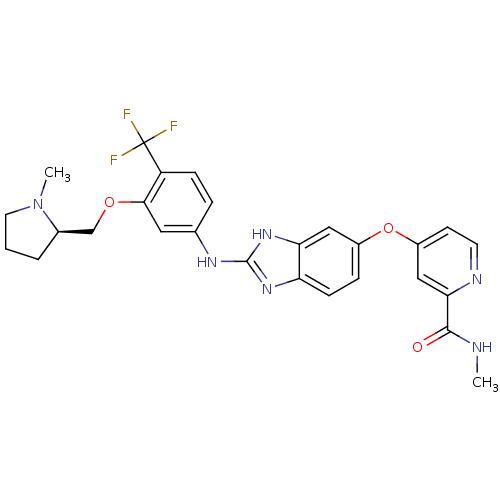
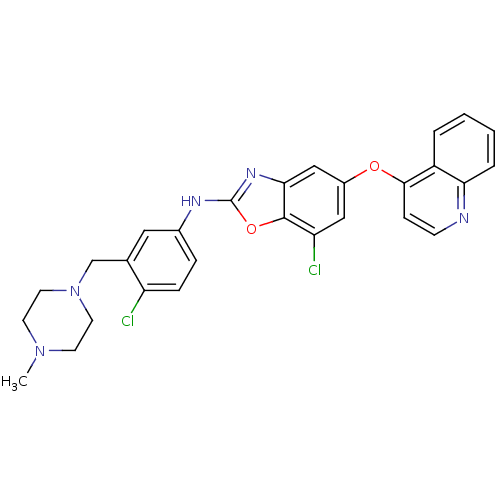
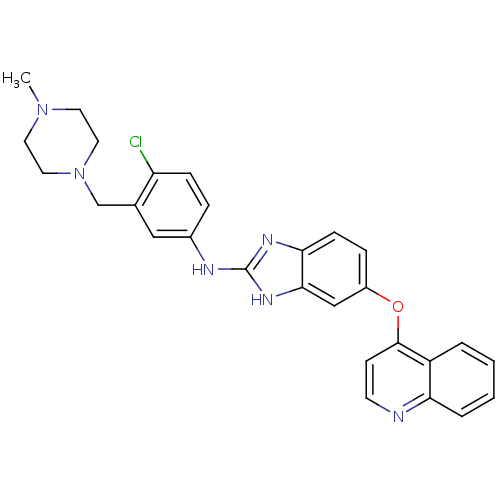
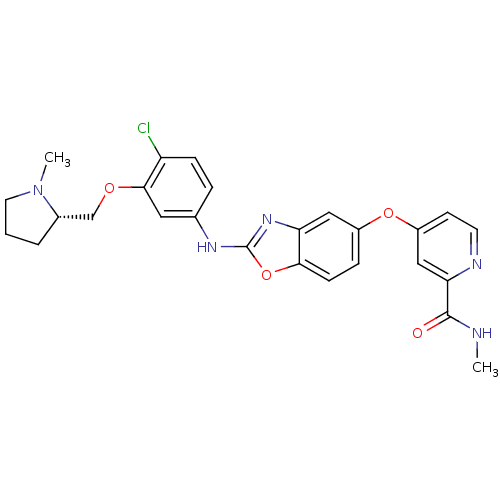
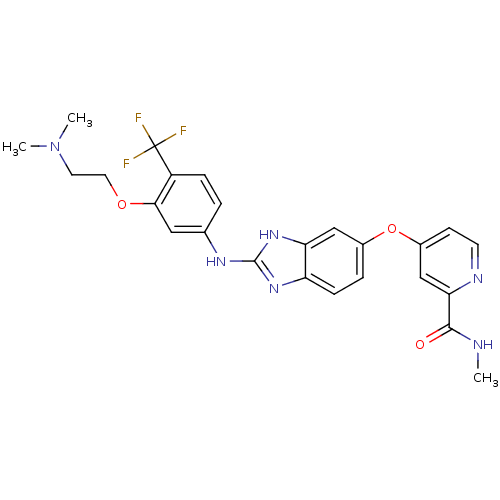
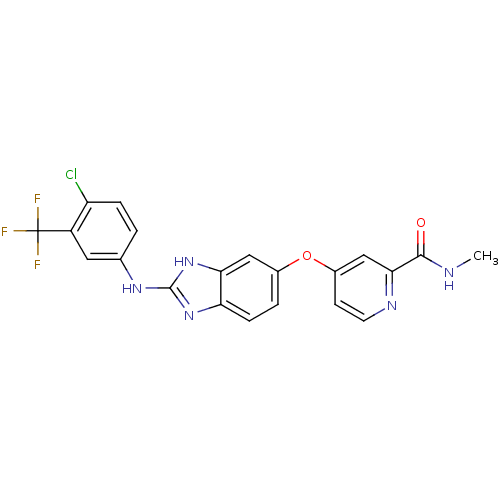
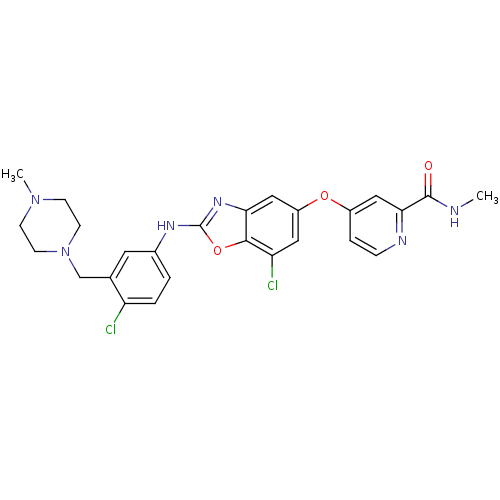
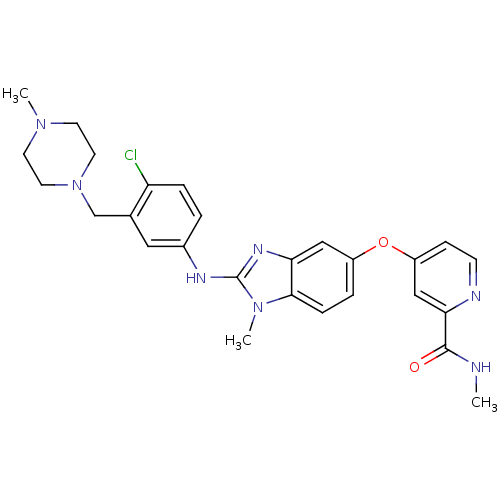
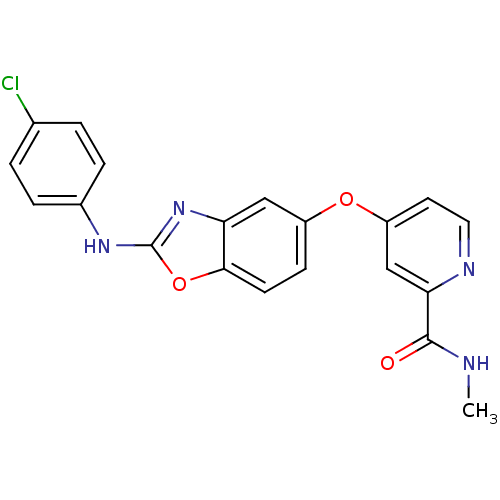
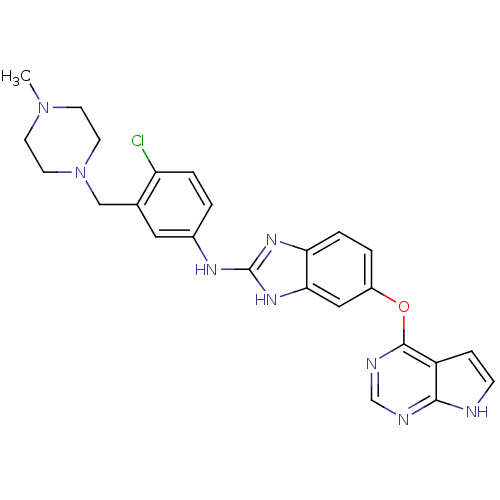
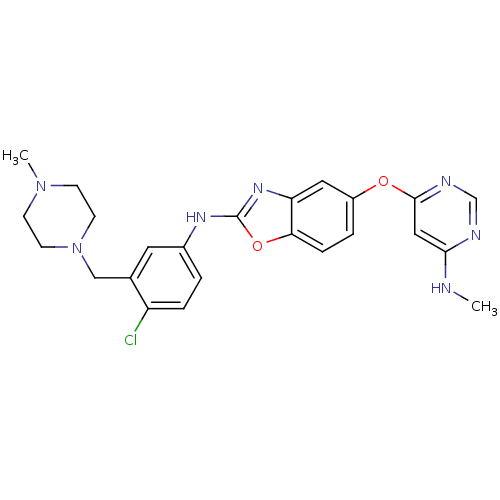
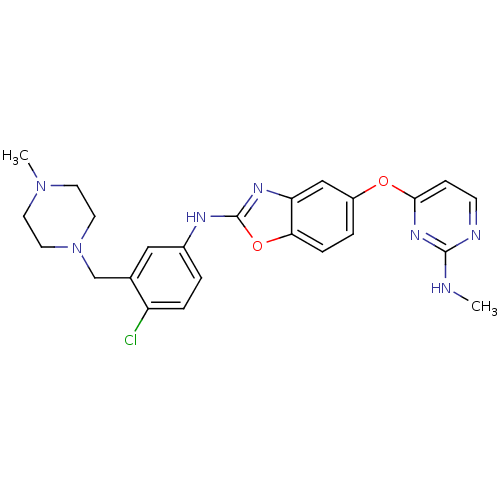
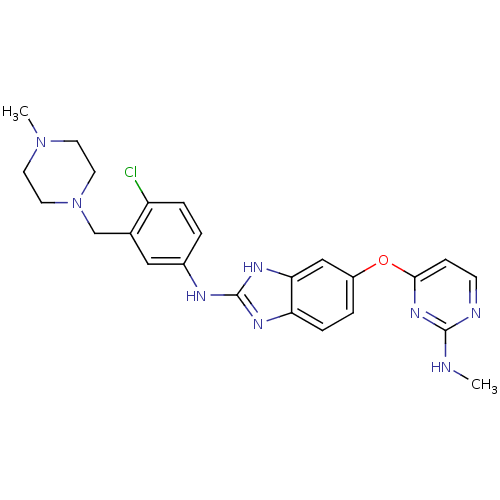
Report error Found 45 Enz. Inhib. hit(s) with all data for entry = 2165
Affinity DataKi: 0.200nM ΔG°: -54.8kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -50.9kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 2nM ΔG°: -49.2kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 2nM ΔG°: -49.2kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 2.80nM ΔG°: -48.3kJ/mole IC50: 4.60nMpH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -48.2kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -48.2kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 3.60nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 3.90nM ΔG°: -47.5kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 4nM ΔG°: -47.5kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 8nM ΔG°: -45.7kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 9nM ΔG°: -45.5kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 9nM ΔG°: -45.5kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 10nM ΔG°: -45.2kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 12nM ΔG°: -44.8kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 17nM ΔG°: -43.9kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMpH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 40nM ΔG°: -41.8kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 48nM ΔG°: -41.4kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataIC50: 80nMpH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 128nM ΔG°: -38.9kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 160nM ΔG°: -38.4kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataKi: 287nM ΔG°: -37.0kJ/molepH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataIC50: 720nMpH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataIC50: 1.14E+3nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataIC50: 3.18E+3nMpH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataIC50: 4.01E+3nMpH: 7.5 T: 2°CAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataIC50: 4.37E+3nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+4nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+4nMAssay Description:The assay involves the phosphorylation of a biotinylated substrate and the detection of this phosphorylation after the addition of a streptavidin-all...More data for this Ligand-Target Pair