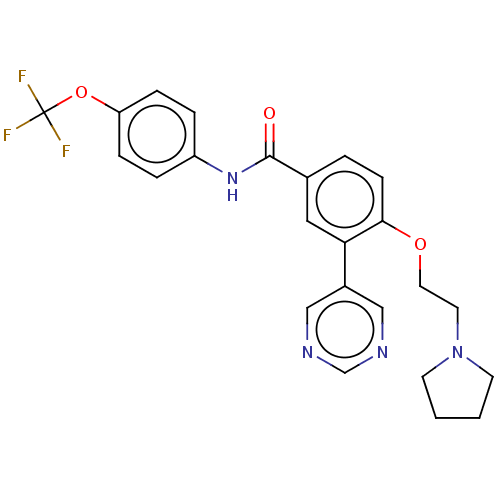
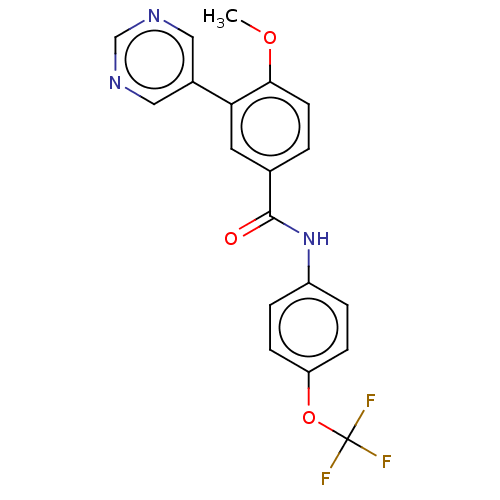
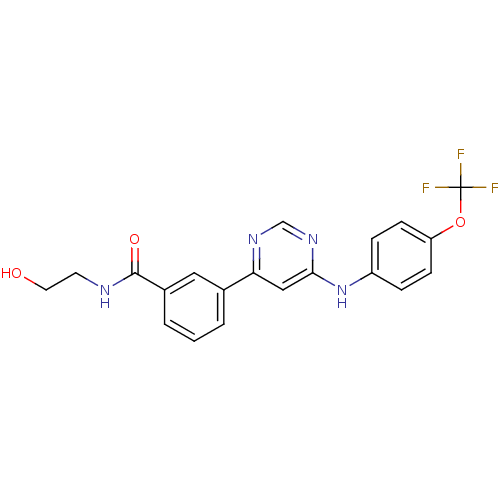
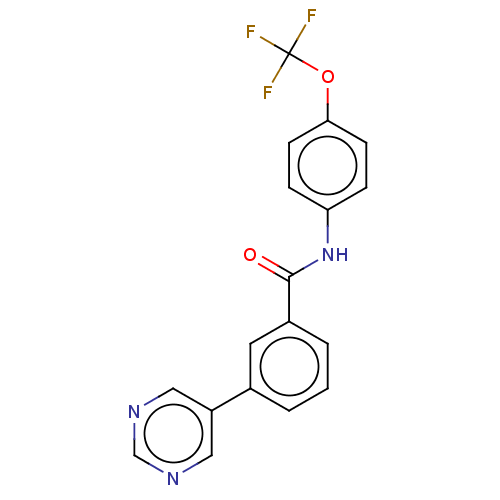
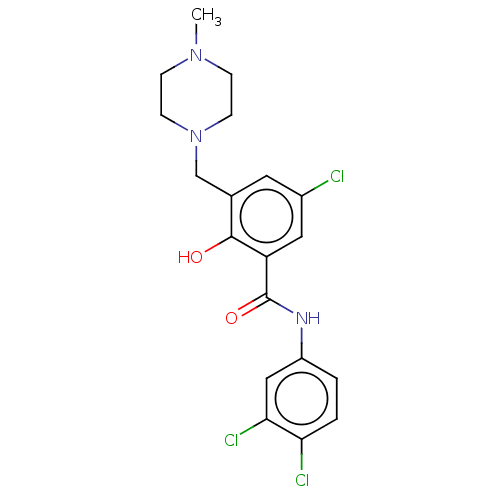
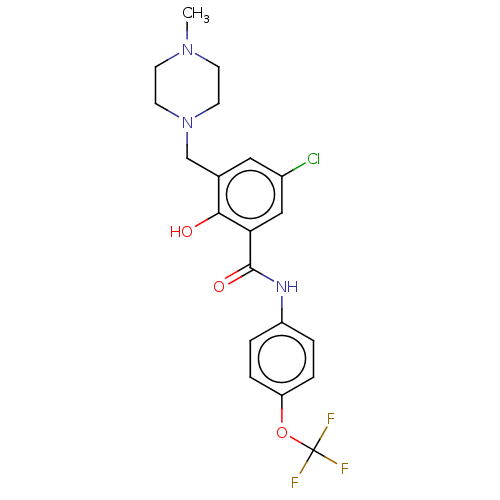
Report error Found 53 Enz. Inhib. hit(s) with all data for entry = 50002484
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 0.300nMAssay Description:Inhibition of ABL1 (64 to 515 residues)(unknown origin) expressed in Escherichia coli using FITC-Ahx-EAIYAAPFAKKK-NH2 peptide as substrate after 60 m...More data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 0.400nMAssay Description:Inhibition of ABL1 (64 to 515 residues)(unknown origin) expressed in Escherichia coli using FITC-Ahx-EAIYAAPFAKKK-NH2 peptide as substrate after 60 m...More data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataKd: 0.5nMAssay Description:Binding affinity to ABL1 (64 to 515 residues) (unknown origin) expressed in Escherichia coli by ITC analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 0.5nMAssay Description:Inhibition of ABL1 (64 to 515 residues)(unknown origin) expressed in Escherichia coli using FITC-Ahx-EAIYAAPFAKKK-NH2 peptide as substrate after 60 m...More data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 0.5nMAssay Description:Inhibition of ABL1 (64 to 515 residues)(unknown origin) expressed in Escherichia coli using FITC-Ahx-EAIYAAPFAKKK-NH2 peptide as substrate after 60 m...More data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 0.700nMAssay Description:Inhibition of ABL1 (64 to 515 residues)(unknown origin) expressed in Escherichia coli using FITC-Ahx-EAIYAAPFAKKK-NH2 peptide as substrate after 60 m...More data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 1.10nMAssay Description:Inhibition of ABL1 (64 to 515 residues)(unknown origin) expressed in Escherichia coli using FITC-Ahx-EAIYAAPFAKKK-NH2 peptide as substrate after 60 m...More data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 1.60nMAssay Description:Inhibition of ABL1 (64 to 515 residues)(unknown origin) expressed in Escherichia coli using FITC-Ahx-EAIYAAPFAKKK-NH2 peptide as substrate after 60 m...More data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 2.30nMAssay Description:Inhibition of ABL1 (64 to 515 residues)(unknown origin) expressed in Escherichia coli using FITC-Ahx-EAIYAAPFAKKK-NH2 peptide as substrate after 60 m...More data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of ABL1 (64 to 515 residues)(unknown origin) expressed in Escherichia coli using FITC-Ahx-EAIYAAPFAKKK-NH2 peptide as substrate after 60 m...More data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 7nMAssay Description:Inhibition of ABL1 (64 to 515 residues)(unknown origin) expressed in Escherichia coli using FITC-Ahx-EAIYAAPFAKKK-NH2 peptide as substrate after 60 m...More data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 9nMAssay Description:Inhibition of recombinant human c-ABL SH3/SH2/SH1 domain (46 to 531 residues) expressed in sf9 insect cells after 30 mins in presence of [gamma-32P]A...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 9nMAssay Description:Displacement of [3H]dofetilide from human ERG by high throughput assayMore data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of ABL1 (64 to 515 residues)(unknown origin) expressed in Escherichia coli using FITC-Ahx-EAIYAAPFAKKK-NH2 peptide as substrate after 60 m...More data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 17nMAssay Description:Inhibition of recombinant human c-ABL SH3/SH2/SH1 domain (46 to 515 residues) expressed in bacterial expression system using EAIYAAPFAKKK as substrat...More data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 18nMAssay Description:Inhibition of ABL1 (64 to 515 residues)(unknown origin) expressed in Escherichia coli using FITC-Ahx-EAIYAAPFAKKK-NH2 peptide as substrate after 60 m...More data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 18nMAssay Description:Inhibition of ABL1 (64 to 515 residues)(unknown origin) expressed in Escherichia coli using FITC-Ahx-EAIYAAPFAKKK-NH2 peptide as substrate after 60 m...More data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 19nMAssay Description:Inhibition of ABL1 (64 to 515 residues)(unknown origin) expressed in Escherichia coli using FITC-Ahx-EAIYAAPFAKKK-NH2 peptide as substrate after 60 m...More data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 24nMAssay Description:Inhibition of ABL1 (64 to 515 residues)(unknown origin) expressed in Escherichia coli using FITC-Ahx-EAIYAAPFAKKK-NH2 peptide as substrate after 60 m...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 310nMAssay Description:Displacement of [3H]dofetilide from human ERG by high throughput assayMore data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 550nMAssay Description:Inhibition of ABL1 (64 to 515 residues)(unknown origin) expressed in Escherichia coli using FITC-Ahx-EAIYAAPFAKKK-NH2 peptide as substrate after 60 m...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 1.10E+3nMAssay Description:Displacement of [3H]dofetilide from human ERG by high throughput assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 1.50E+3nMAssay Description:Displacement of [3H]dofetilide from human ERG by high throughput assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of human ERG by automated patch clamp assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibition of human ERG by automated patch clamp assayMore data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataKd: 2.00E+3nMAssay Description:Binding affinity to ABL1 (64 to 515 residues) (unknown origin) expressed in Escherichia coli by NMR analysisMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 3.60E+3nMAssay Description:Displacement of [3H]dofetilide from human ERG by high throughput assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 3.70E+3nMAssay Description:Displacement of [3H]dofetilide from human ERG by high throughput assayMore data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataKd: 4.00E+3nMAssay Description:Binding affinity to ABL1 (64 to 515 residues) (unknown origin) expressed in Escherichia coli by NMR analysisMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 4.20E+3nMAssay Description:Inhibition of human ERG by automated patch clamp assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 4.50E+3nMAssay Description:Inhibition of human ERG by manual patch clamp assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of human ERG by manual patch clamp assayMore data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataKd: 6.00E+3nMAssay Description:Binding affinity to ABL1 (64 to 515 residues) (unknown origin) expressed in Escherichia coli by NMR analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataKd: 6.00E+3nMAssay Description:Binding affinity to ABL1 (64 to 515 residues) (unknown origin) expressed in Escherichia coli by NMR analysisMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 9.60E+3nMAssay Description:Displacement of [3H]dofetilide from human ERG by high throughput assayMore data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataKd: 1.00E+4nMAssay Description:Binding affinity to ABL1 (64 to 515 residues) (unknown origin) expressed in Escherichia coli by NMR analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of recombinant human c-ABL SH1 domain expressed in sf9 insect cells after 30 mins in presence of [gamma-32P]ATP by autoradiographyMore data for this Ligand-Target Pair
TargetTyrosine-protein kinase ABL1(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of recombinant human c-ABL SH1 domain (46 to 515 residues) expressed in bacterial expression system using EAIYAAPFAKKK as substrate measur...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Displacement of [3H]dofetilide from human ERG by high throughput assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 1.05E+4nMAssay Description:Displacement of [3H]dofetilide from human ERG by high throughput assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 1.70E+4nMAssay Description:Displacement of [3H]dofetilide from human ERG by high throughput assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of CYP2C9 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of CYP2D6 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of CYP3A4 (unknown origin)More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 2.60E+4nMAssay Description:Inhibition of human ERG by automated patch clamp assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of human ERG by automated patch clamp assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of human ERG by automated patch clamp assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Displacement of [3H]dofetilide from human ERG by high throughput assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Displacement of [3H]dofetilide from human ERG by high throughput assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Novartis Institutes For Biomedical Research
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Displacement of [3H]dofetilide from human ERG by high throughput assayMore data for this Ligand-Target Pair
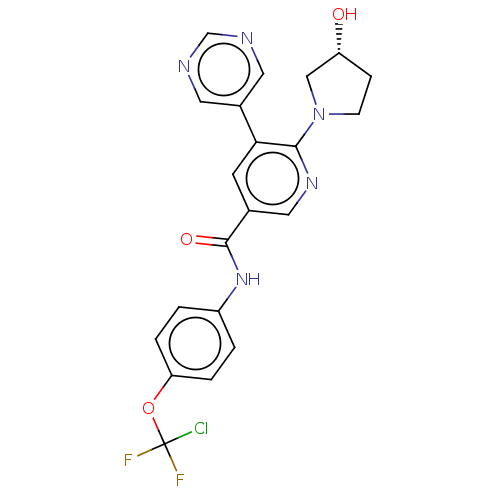
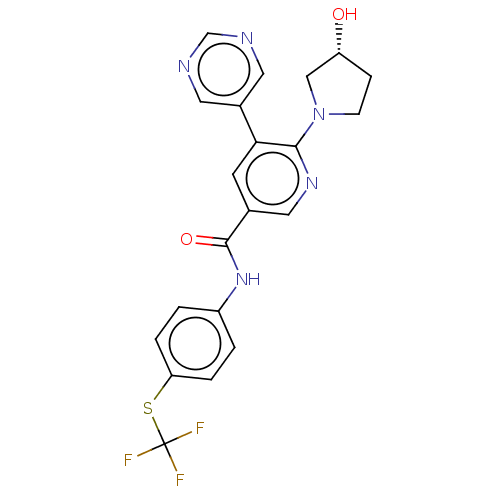
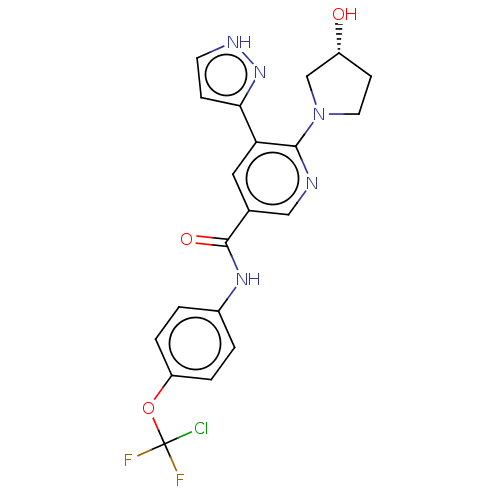
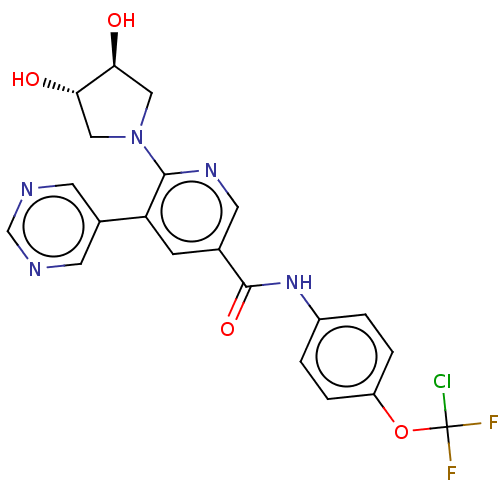


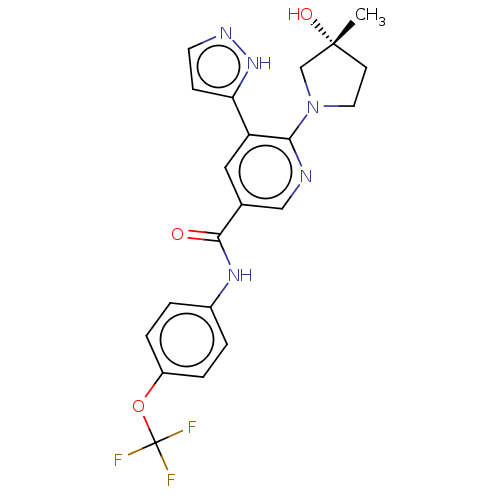

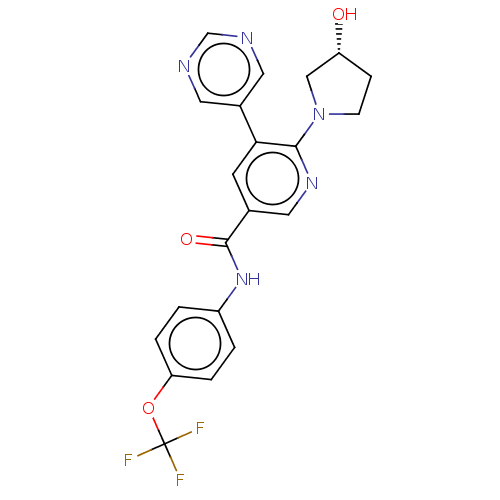
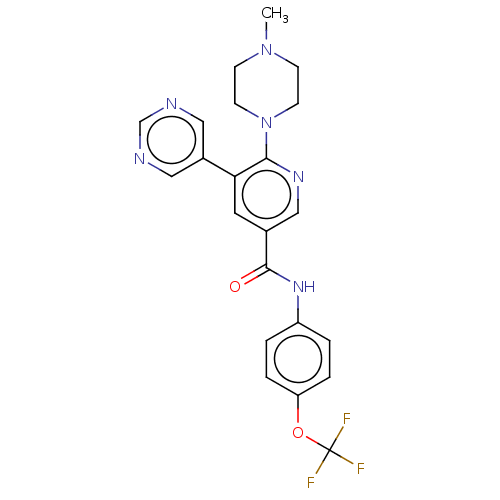
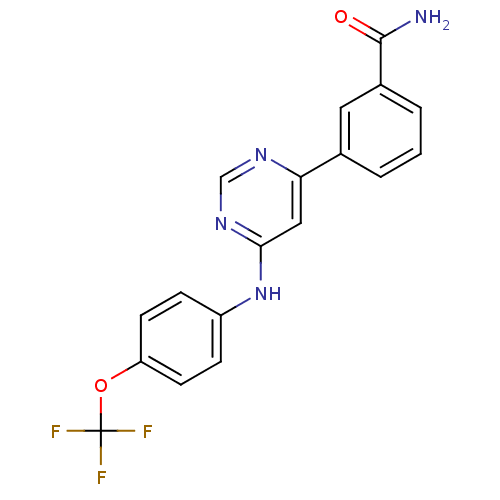
 3D Structure (crystal)
3D Structure (crystal)