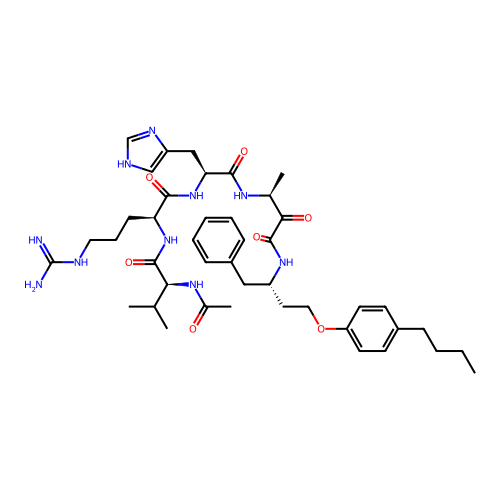
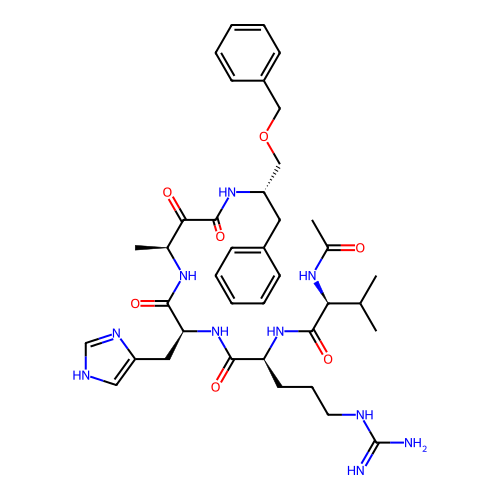
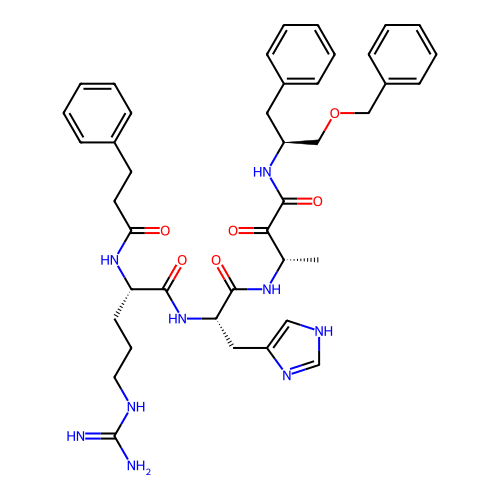
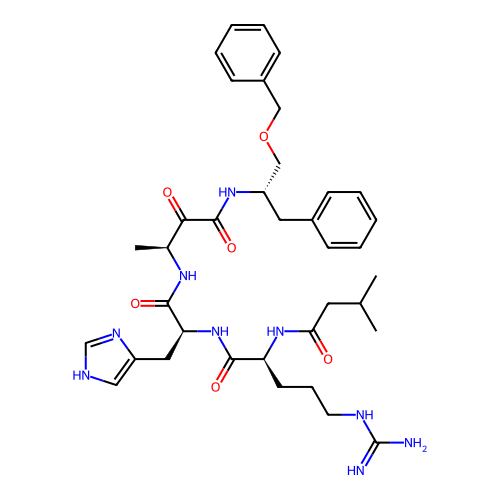
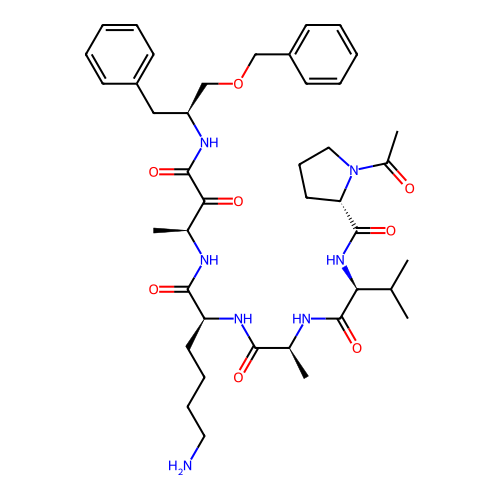
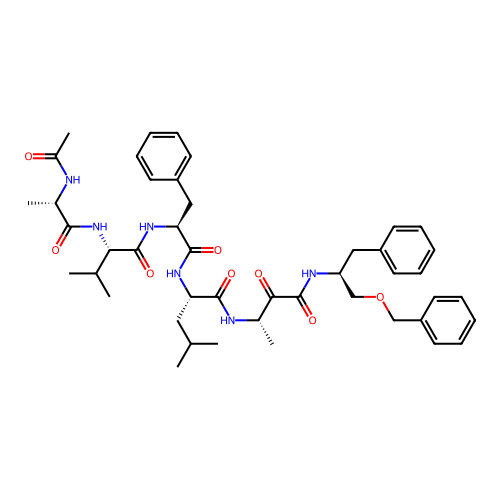
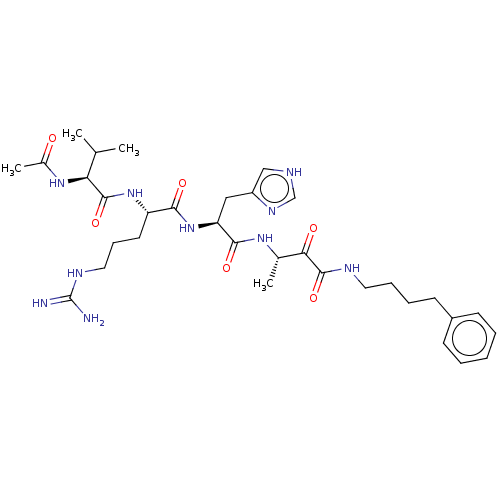
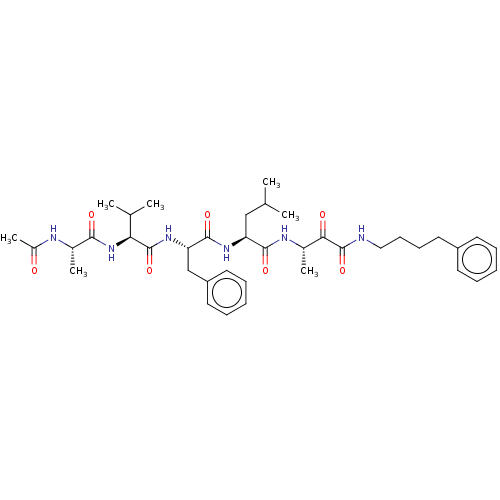
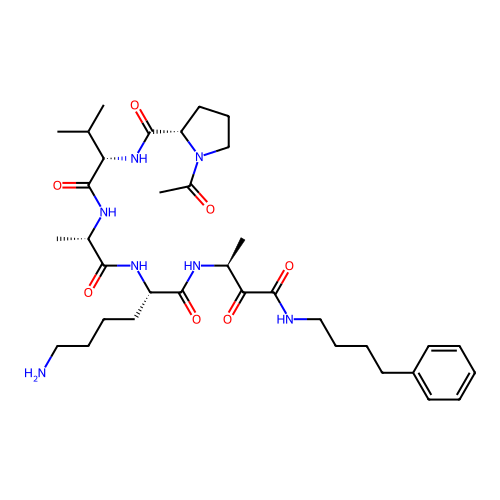
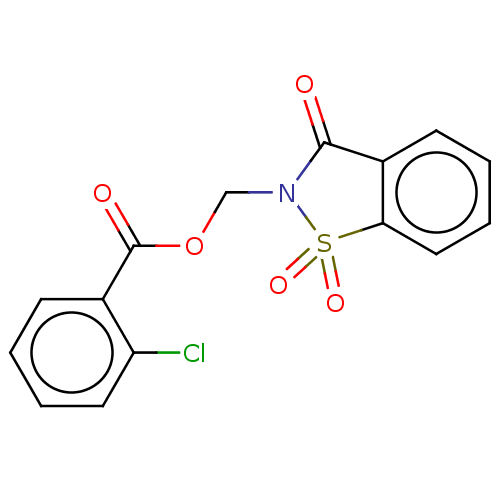
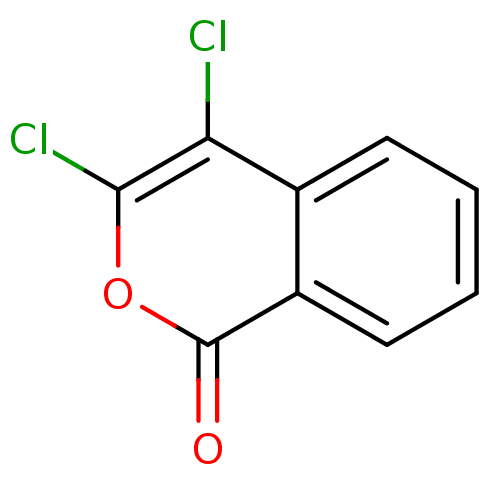
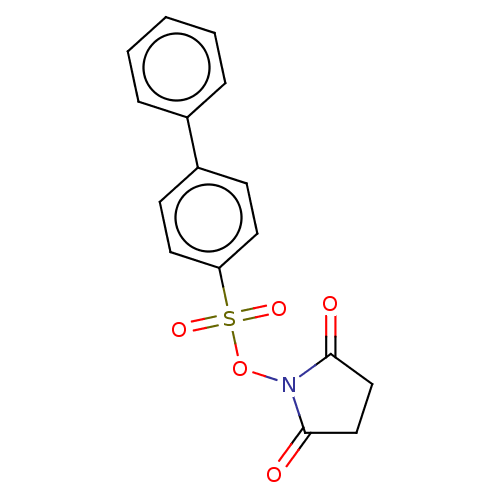
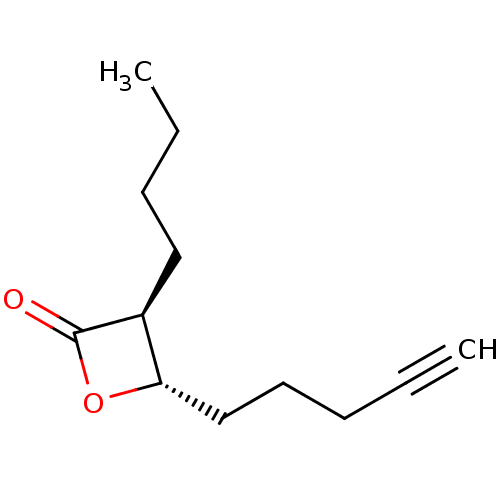
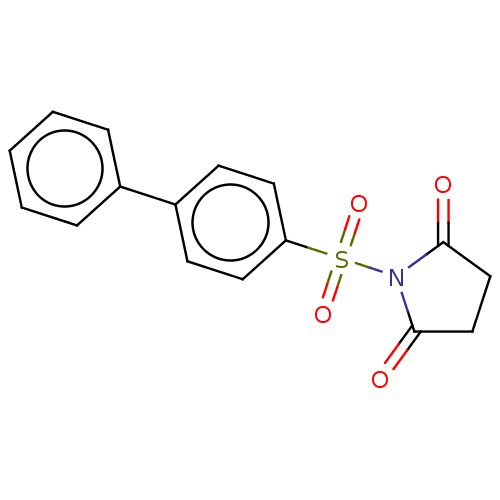
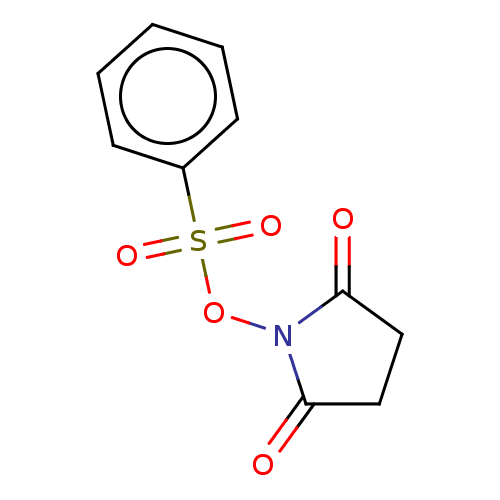
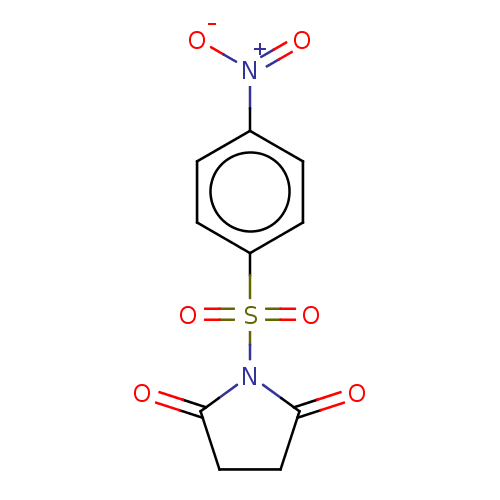
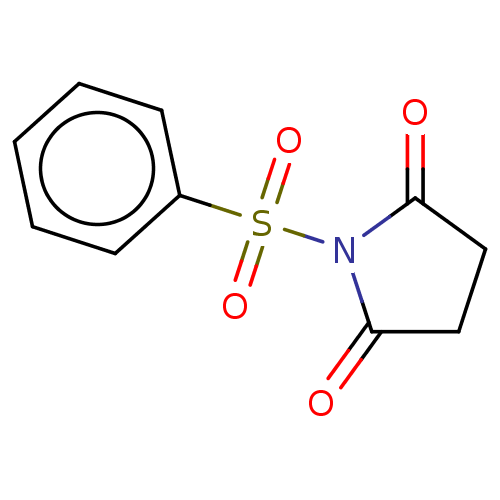
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataKi: 0.700nMAssay Description:Inhibition of Escherichia coli GlpG extracted from Escherichia coli C43(DE3) using KSp96 as substrate incubated for 1 hr by fluorescence based assayMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataKi: 1.30nMAssay Description:Inhibition of Escherichia coli GlpG extracted from Escherichia coli C43(DE3) using KSp96 as substrate incubated for 1 hr by fluorescence based assayMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataKi: 1.40nMAssay Description:Inhibition of Escherichia coli GlpG extracted from Escherichia coli C43(DE3) using KSp96 as substrate incubated for 1 hr by fluorescence based assayMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 1.90nMAssay Description:Inhibition of Escherichia coli GlpG extracted from Escherichia coli C43(DE3) using KSp96 as substrate incubated for 1 hr by fluorescence based assayMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 2nMAssay Description:Inhibition of Escherichia coli GlpG extracted from Escherichia coli C43(DE3) using KSp96 as substrate incubated for 1 hr by fluorescence based assayMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 2.30nMAssay Description:Inhibition of Escherichia coli GlpG extracted from Escherichia coli C43(DE3) using KSp96 as substrate incubated for 1 hr by fluorescence based assayMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 2.30nMAssay Description:Inhibition of Escherichia coli GlpG extracted from Escherichia coli C43(DE3) using KSp96 as substrate incubated for 1 hr by fluorescence based assayMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 2.5nMAssay Description:Inhibition of Escherichia coli GlpG extracted from Escherichia coli C43(DE3) using KSp96 as substrate incubated for 1 hr by fluorescence based assayMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 5.60nMAssay Description:Inhibition of Escherichia coli GlpG extracted from Escherichia coli C43(DE3) using KSp96 as substrate incubated for 1 hr by fluorescence based assayMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 15nMAssay Description:Inhibition of Escherichia coli GlpG extracted from Escherichia coli C43(DE3) using KSp96 as substrate incubated for 1 hr by fluorescence based assayMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 44nMAssay Description:Inhibition of Escherichia coli GlpG extracted from Escherichia coli C43(DE3) using KSp96 as substrate incubated for 1 hr by fluorescence based assayMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 56nMAssay Description:Inhibition of Escherichia coli GlpG extracted from Escherichia coli C43(DE3) using KSp96 as substrate incubated for 1 hr by fluorescence based assayMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataEC50: 123nMAssay Description:Inhibition of GlpG in wild-type Escherichia coli MC4100 using MBP-Flag-LacYTMD2-Trx as substrateMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataEC50: 131nMAssay Description:Inhibition of GlpG in wild-type Escherichia coli MC4100 using MBP-Flag-LacYTMD2-Trx as substrateMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 132nMAssay Description:Inhibition of C-terminal His-tagged full length Escherichia coli GIpG expressed in Escherichia coli C43 (DE3) using Ac-RVRHA-4mc as substrate preincu...More data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataEC50: 173nMAssay Description:Inhibition of GlpG in wild-type Escherichia coli MC4100 using MBP-Flag-LacYTMD2-Trx as substrateMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 300nMAssay Description:Inhibition of Escherichia coli GlpG extracted from Escherichia coli C43(DE3) using KSp96 as substrate incubated for 1 hr by fluorescence based assayMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 330nMAssay Description:Inhibition of Escherichia coli GlpG extracted from Escherichia coli C43(DE3) using KSp96 as substrate incubated for 1 hr by fluorescence based assayMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataEC50: 337nMAssay Description:Inhibition of GlpG in wild-type Escherichia coli MC4100 using MBP-Flag-LacYTMD2-Trx as substrateMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataEC50: 424nMAssay Description:Inhibition of GlpG in wild-type Escherichia coli MC4100 using MBP-Flag-LacYTMD2-Trx as substrateMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataEC50: 1.47E+3nMAssay Description:Inhibition of GlpG in wild-type Escherichia coli MC4100 using MBP-Flag-LacYTMD2-Trx as substrateMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataEC50: 1.61E+3nMAssay Description:Inhibition of GlpG in wild-type Escherichia coli MC4100 using MBP-Flag-LacYTMD2-Trx as substrateMore data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 2.80E+3nMAssay Description:Inhibition of GlpG in Escherichia coli BL21 assessed as inhibition of FP-Rh labeling treated with compound for 30 mins prior to FP-Rh addition by com...More data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 1.90E+4nMpH: 7.4 T: 2°CAssay Description:We selected several inhibitor structures and determined their apparent IC50 by competitive ABPP.More data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 2.60E+4nMAssay Description:Inhibition of GlpG in Escherichia coli BL21 assessed as inhibition of FP-Rh labeling treated with compound for 30 mins prior to FP-Rh addition by com...More data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 4.40E+4nMpH: 7.4 T: 2°CAssay Description:We selected several inhibitor structures and determined their apparent IC50 by competitive ABPP.More data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 6.80E+4nMAssay Description:Inhibition of GlpG in Escherichia coli BL21 assessed as inhibition of FP-Rh labeling treated with compound for 30 mins prior to FP-Rh addition by com...More data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 1.20E+5nMAssay Description:Inhibition of GlpG in Escherichia coli BL21 assessed as inhibition of FP-Rh labeling treated with compound for 30 mins prior to FP-Rh addition by com...More data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 3.10E+5nMAssay Description:Inhibition of GlpG in Escherichia coli BL21 assessed as inhibition of FP-Rh labeling treated with compound for 30 mins prior to FP-Rh addition by com...More data for this Ligand-Target Pair
TargetRhomboid protease GlpG(Escherichia coli (strain K12))
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Science
Curated by ChEMBL
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of GlpG in Escherichia coli BL21 assessed as inhibition of FP-Rh labeling treated with compound for 30 mins prior to FP-Rh addition by com...More data for this Ligand-Target Pair