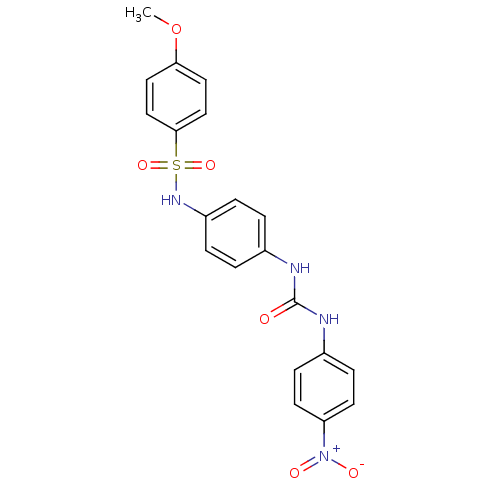
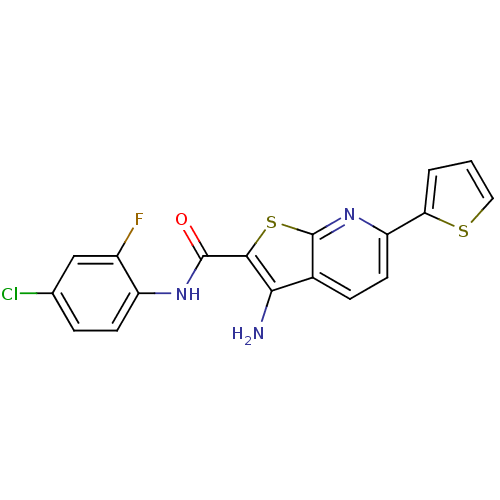
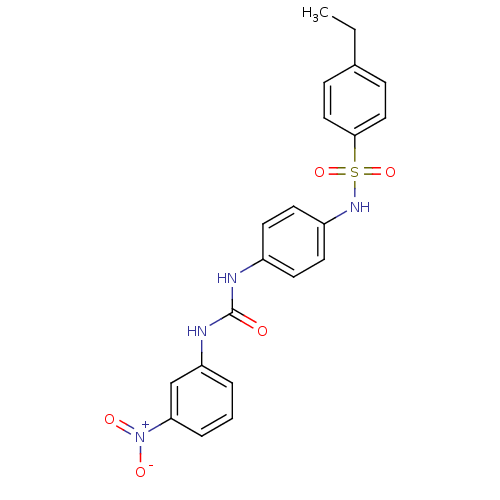
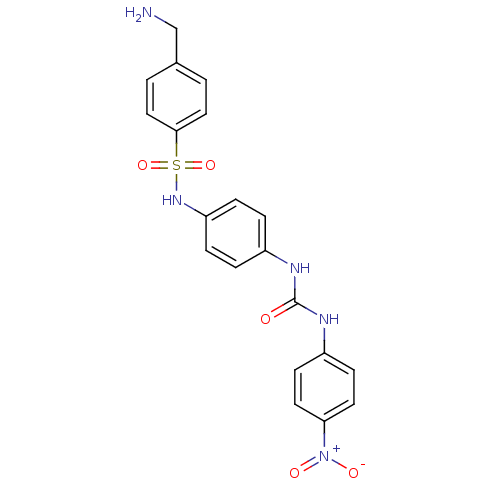
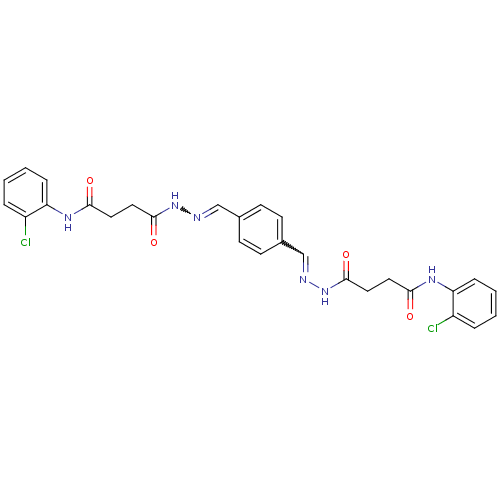
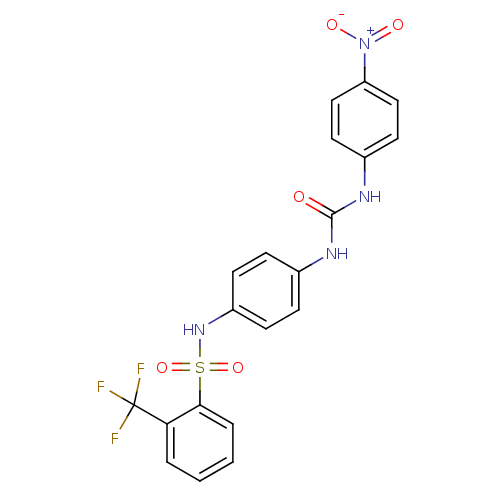
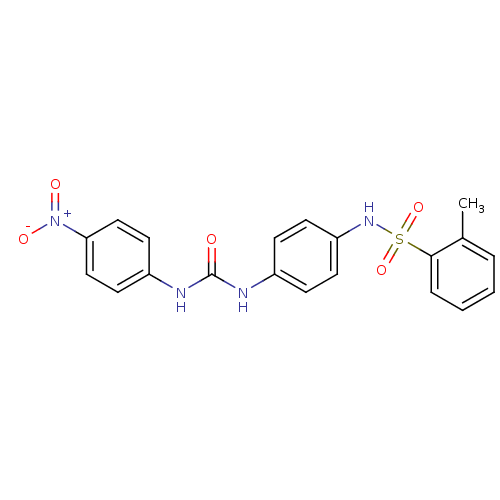
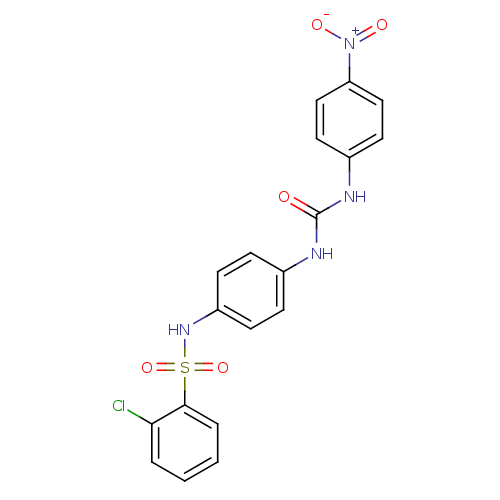
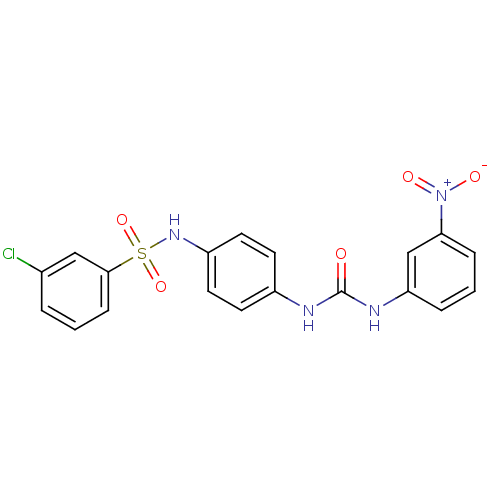
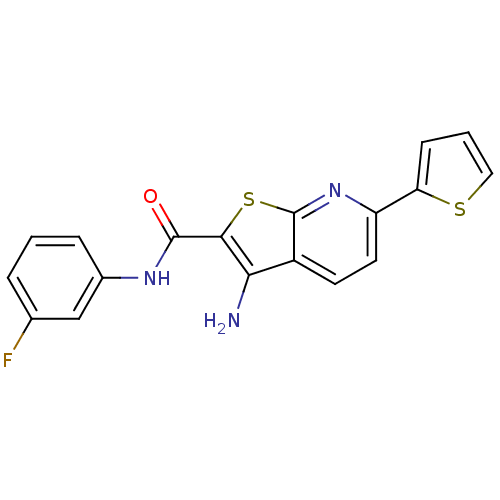
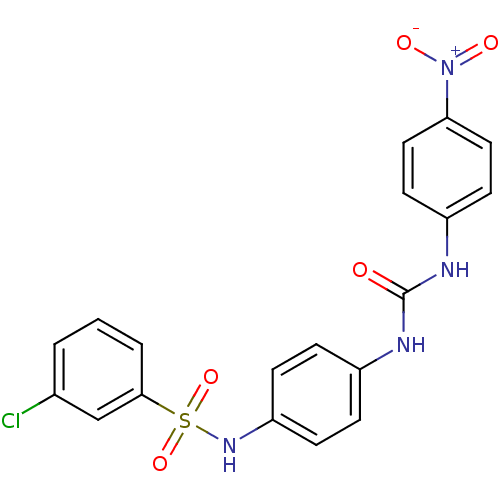
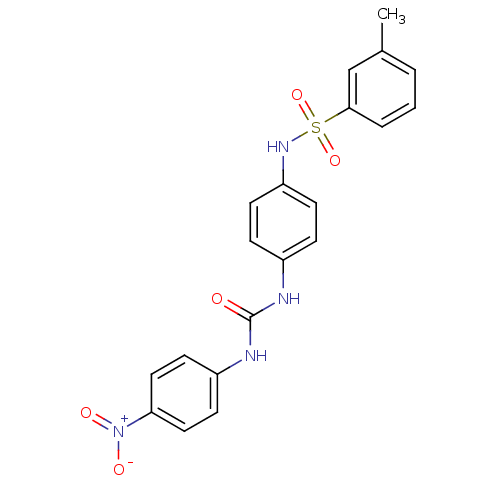
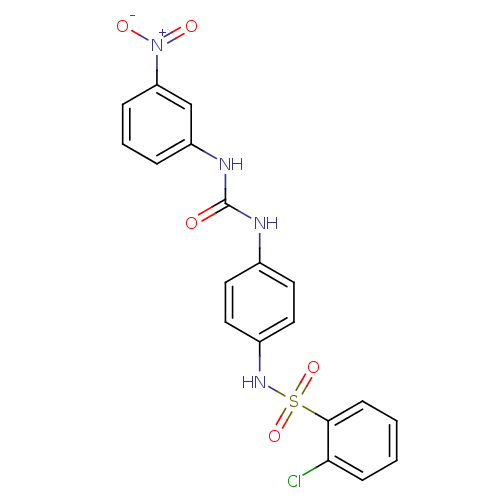
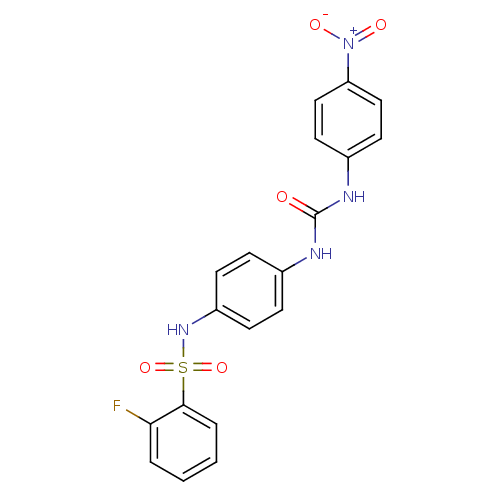
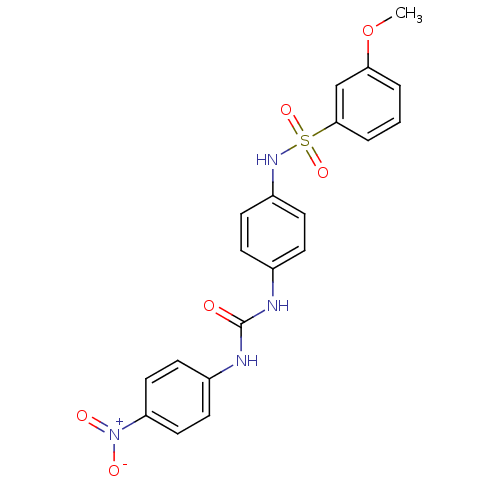
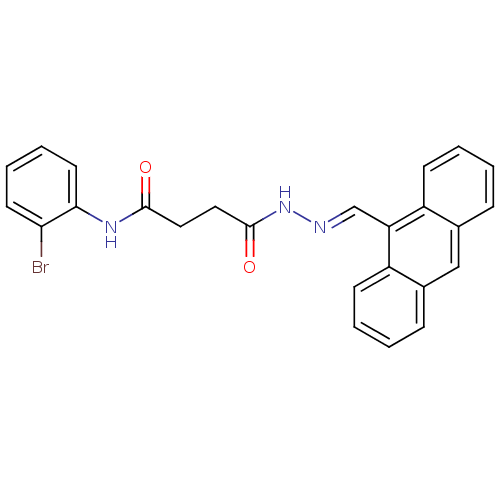
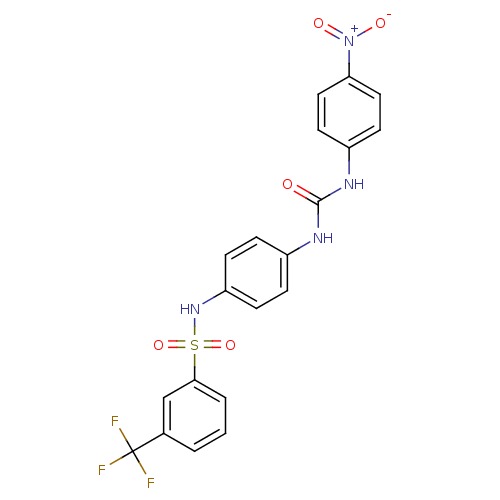
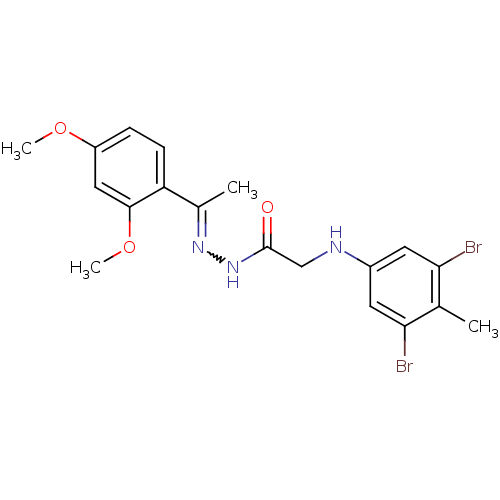
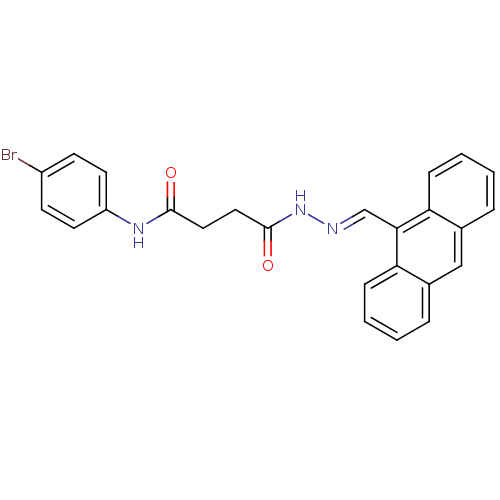
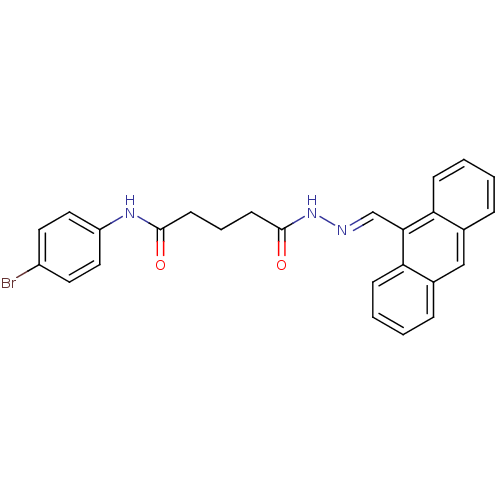
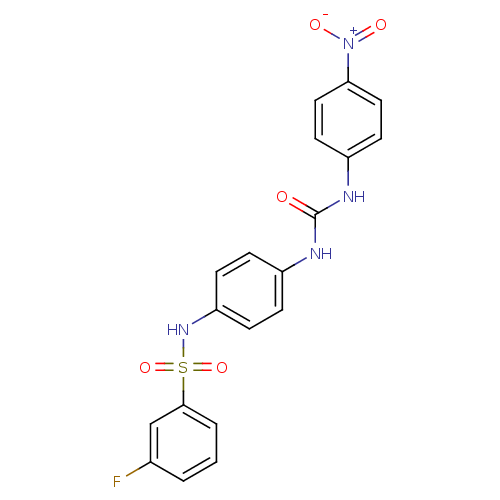
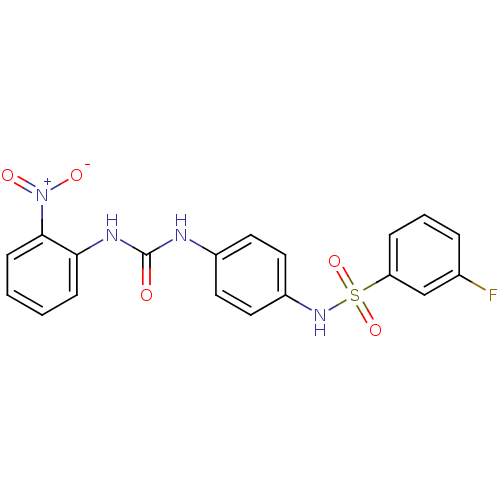
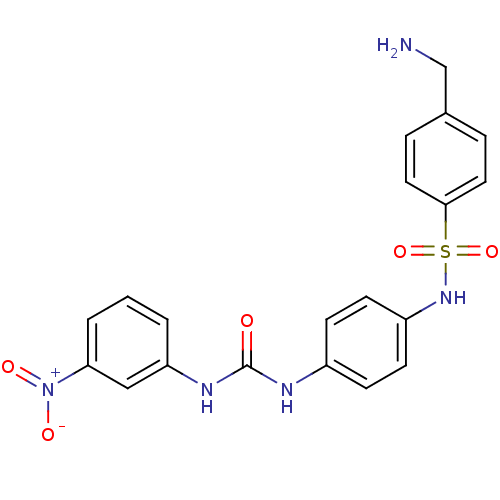
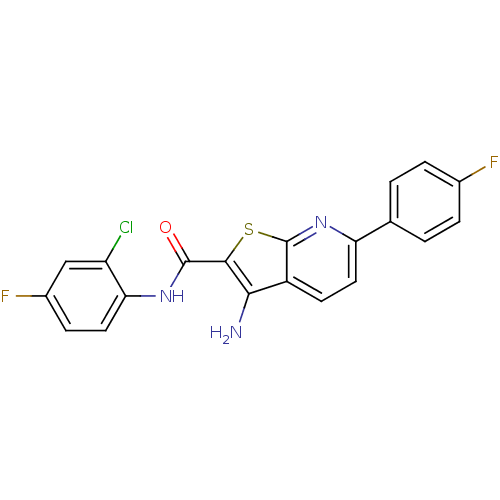
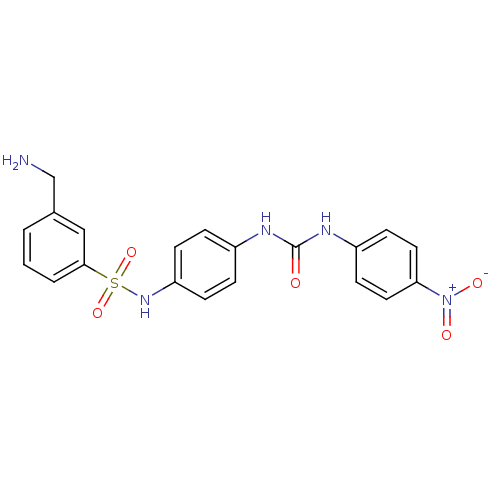
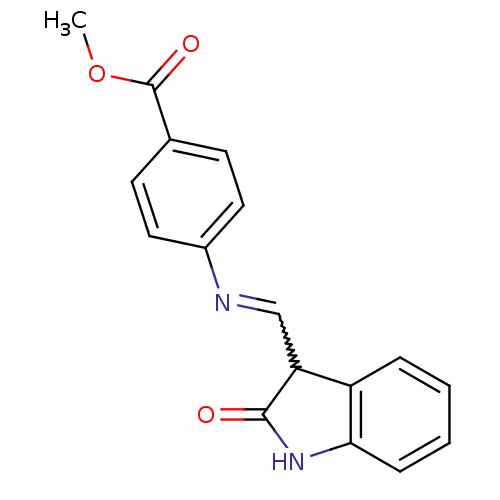
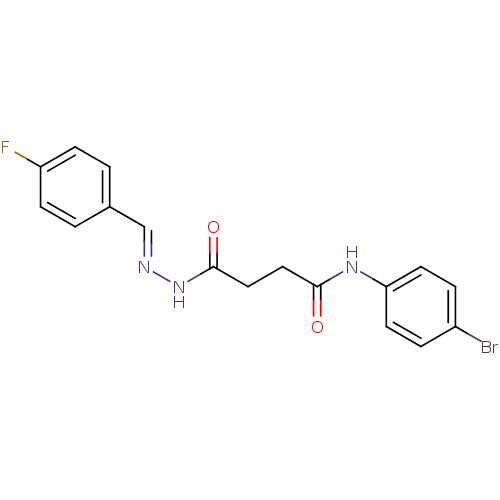
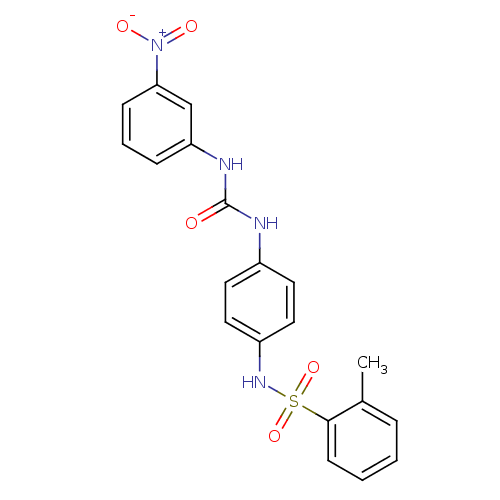
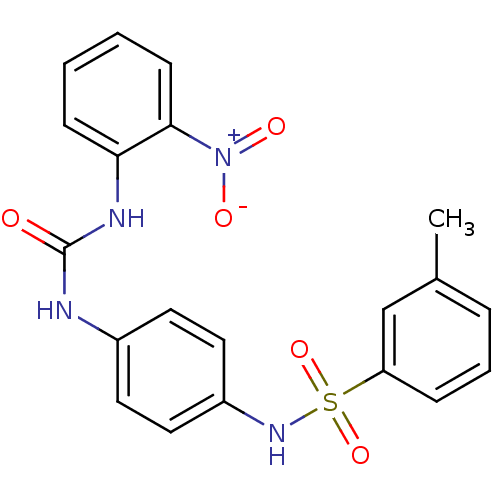
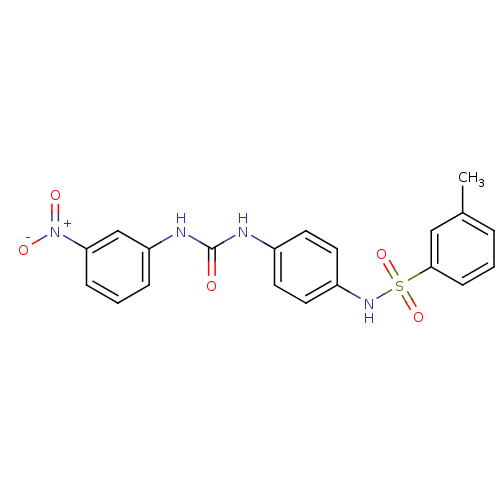
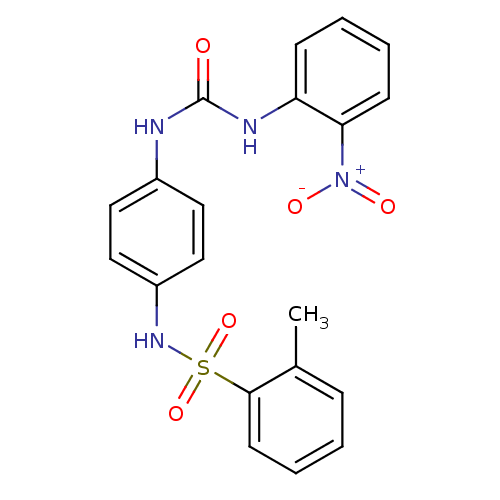
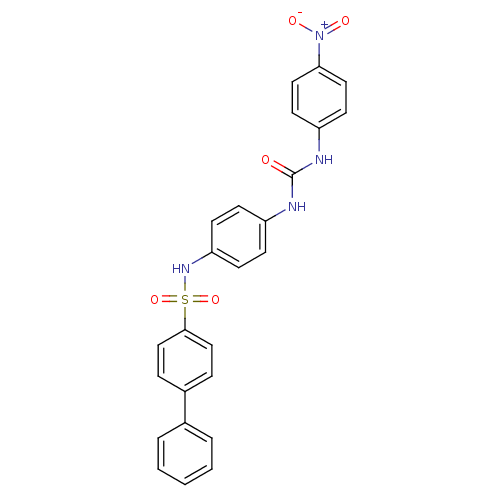
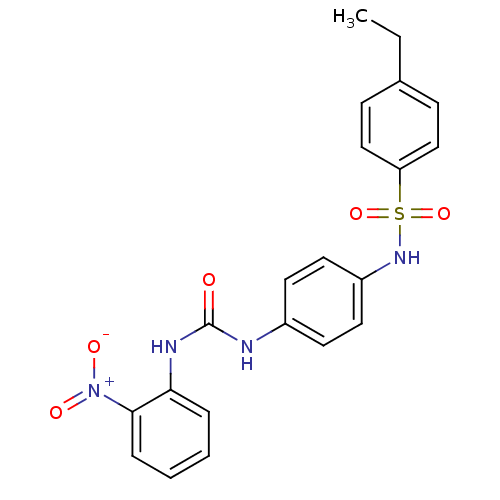
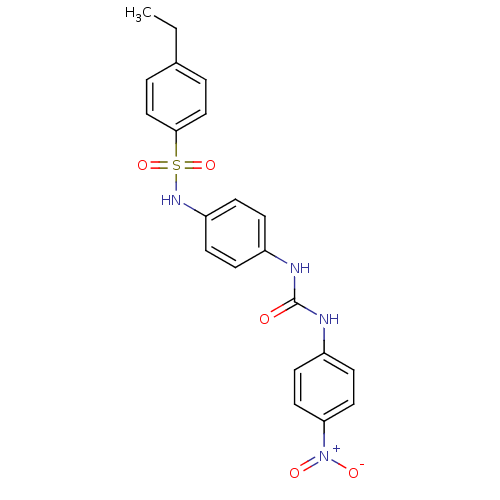
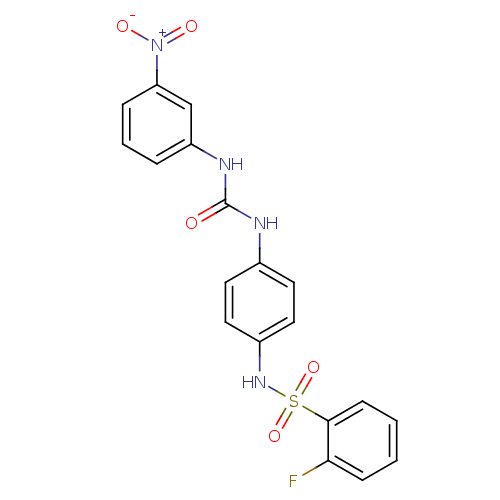
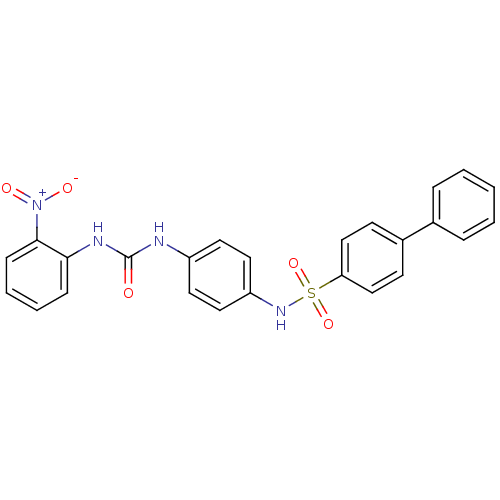
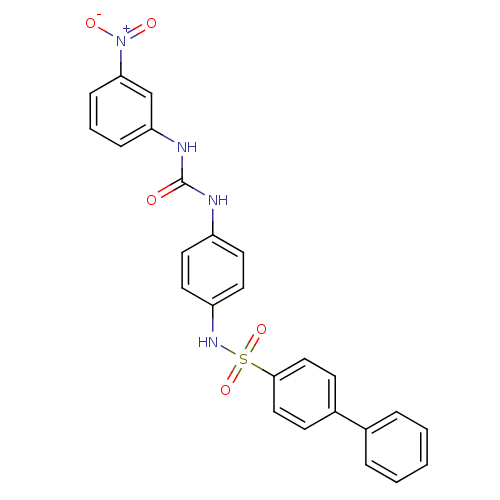
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 2.00E+3nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 4.00E+3nMAssay Description:Inhibition of Bacillus anthracis nicotinate-nucleotide adenosyltransferase preincubated for 5 mins before addition of substrate by malachite green re...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 7.30E+3nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 9.00E+3nMAssay Description:The enzymatic assay was masured at fixed NaMN and ATP concentration (equal to two-fold km values)with various concentration of inhibitory compounds.More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataKi: 9.00E+3nM IC50: 2.50E+4nMAssay Description:The enzymatic assay was masured at fixed NaMN and ATP concentration (equal to two-fold km values)with various concentration of inhibitory compounds.More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataKi: 9.00E+3nMAssay Description:Inhibition of Bacillus anthracis nicotinate-nucleotide adenosyltransferase preincubated for 5 mins before addition of NaMN substrate by malachite gre...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataKi: 1.00E+4nMAssay Description:Inhibition of Bacillus anthracis nicotinate-nucleotide adenosyltransferase preincubated for 5 mins before addition of ATP substrate by malachite gree...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataKi: 1.00E+4nMAssay Description:The enzymatic assay was masured at fixed NaMN and ATP concentration (equal to two-fold km values)with various concentration of inhibitory compounds.More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.01E+4nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.20E+4nMAssay Description:The enzymatic assay was masured at fixed NaMN and ATP concentration (equal to two-fold km values)with various concentration of inhibitory compounds.More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.20E+4nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.21E+4nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.30E+4nMAssay Description:Inhibition of Bacillus anthracis nicotinate-nucleotide adenosyltransferase preincubated for 5 mins before addition of substrate by malachite green re...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.30E+4nMAssay Description:Inhibition of Bacillus anthracis nicotinate-nucleotide adenosyltransferase preincubated for 5 mins before addition of substrate by malachite green re...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.33E+4nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.63E+4nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.73E+4nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.75E+4nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataKi: 1.80E+4nM IC50: 3.60E+4nMAssay Description:The enzymatic assay was masured at fixed NaMN and ATP concentration (equal to two-fold km values)with various concentration of inhibitory compounds.More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataKi: 1.80E+4nMAssay Description:Inhibition of Bacillus anthracis nicotinate-nucleotide adenosyltransferase preincubated for 5 mins before addition of NaMN substrate by malachite gre...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.93E+4nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.99E+4nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 2.04E+4nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 2.22E+4nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 2.50E+4nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibition of Bacillus anthracis nicotinate-nucleotide adenosyltransferase preincubated for 5 mins before addition of substrate by malachite green re...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibition of Bacillus anthracis nicotinate-nucleotide adenosyltransferase preincubated for 5 mins before addition of substrate by malachite green re...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 2.67E+4nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Bacillus anthracis nicotinate-nucleotide adenosyltransferase preincubated for 5 mins before addition of substrate by malachite green re...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataKi: 3.20E+4nMAssay Description:The enzymatic assay was masured at fixed NaMN and ATP concentration (equal to two-fold km values)with various concentration of inhibitory compounds.More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataKi: 3.20E+4nMAssay Description:Inhibition of Bacillus anthracis nicotinate-nucleotide adenosyltransferase preincubated for 5 mins before addition of ATP substrate by malachite gree...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 3.30E+4nMAssay Description:Inhibition of Bacillus anthracis nicotinate-nucleotide adenosyltransferase preincubated for 5 mins before addition of substrate by malachite green re...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 3.30E+4nMAssay Description:The enzymatic assay was masured at fixed NaMN and ATP concentration (equal to two-fold km values)with various concentration of inhibitory compounds.More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 3.58E+4nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 3.61E+4nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 5.12E+4nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 6.30E+4nMAssay Description:The enzymatic assay was masured at fixed NaMN and ATP concentration (equal to two-fold km values)with various concentration of inhibitory compounds.More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 7.46E+4nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 9.80E+4nMAssay Description:The enzymatic assay was masured at fixed NaMN and ATP concentration (equal to two-fold km values)with various concentration of inhibitory compounds.More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.22E+5nMAssay Description:Inhibition of Bacillus anthracis nicotinate-nucleotide adenosyltransferase preincubated for 5 mins before addition of substrate by malachite green re...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.50E+5nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.50E+5nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.50E+5nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.50E+5nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.50E+5nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.50E+5nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.50E+5nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.50E+5nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.50E+5nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
TargetProbable nicotinate-nucleotide adenylyltransferase(Bacillus anthracis)
University of Alabama At Birmingham
University of Alabama At Birmingham
Affinity DataIC50: 1.50E+5nMpH: 7.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinic acid mononucleotide adenylyltransferase (NaMNAT)using HPLC Enzyme As...More data for this Ligand-Target Pair
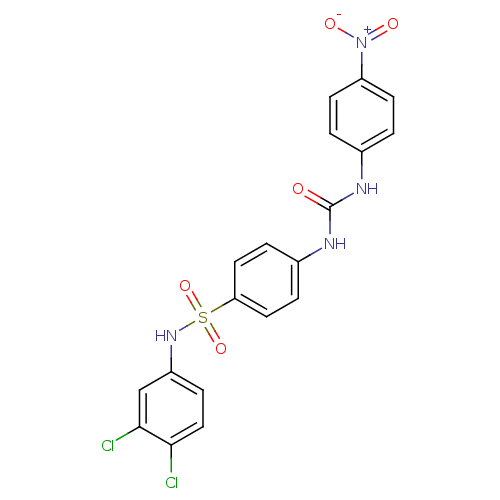
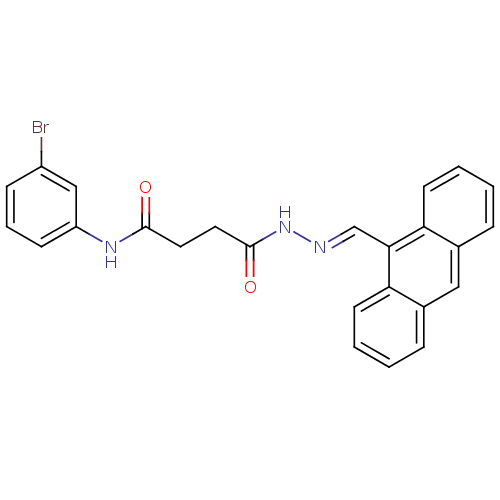
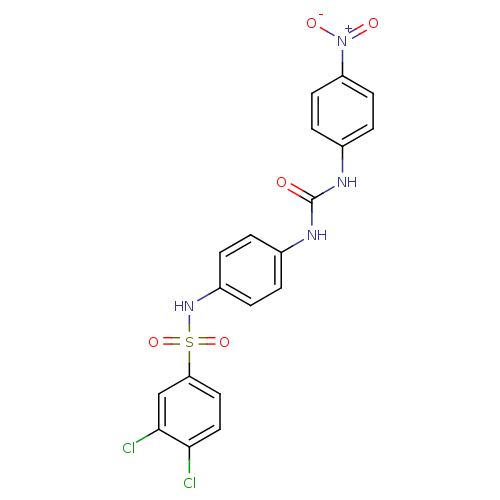
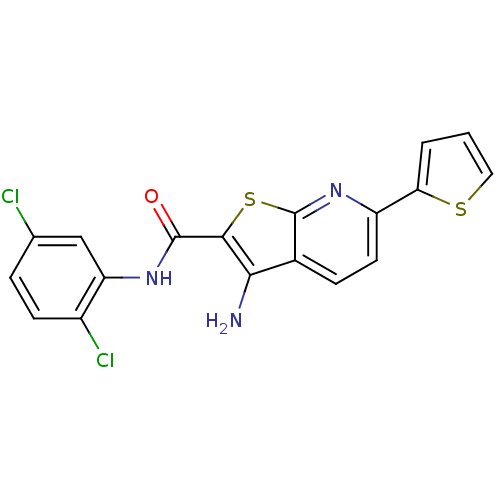
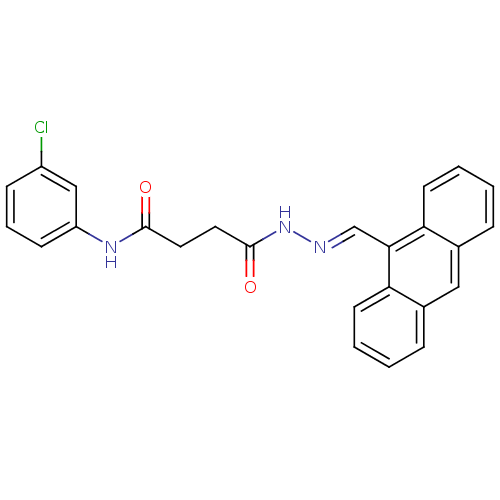
 3D Structure (crystal)
3D Structure (crystal)