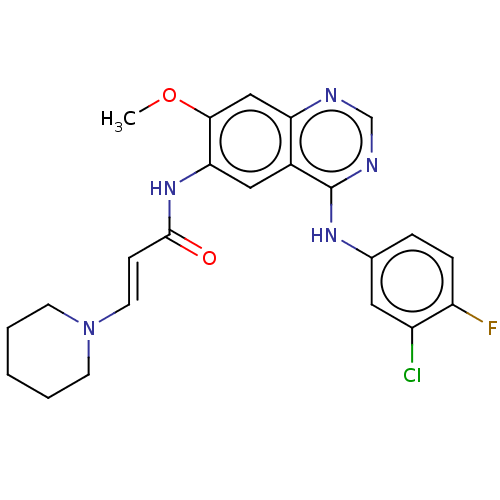
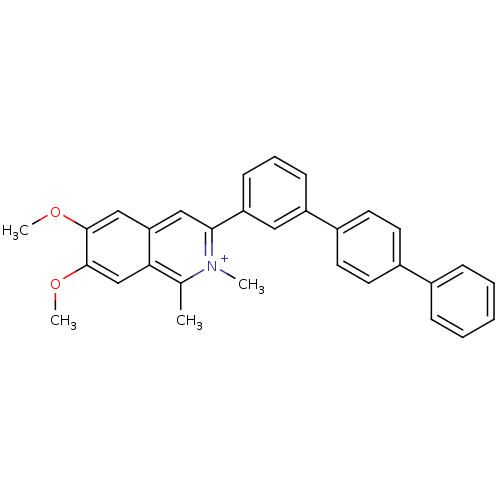
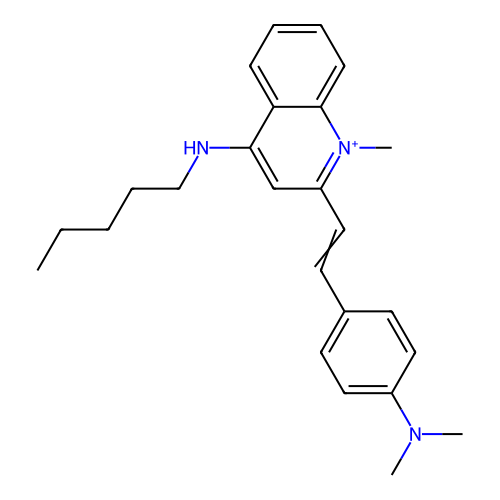
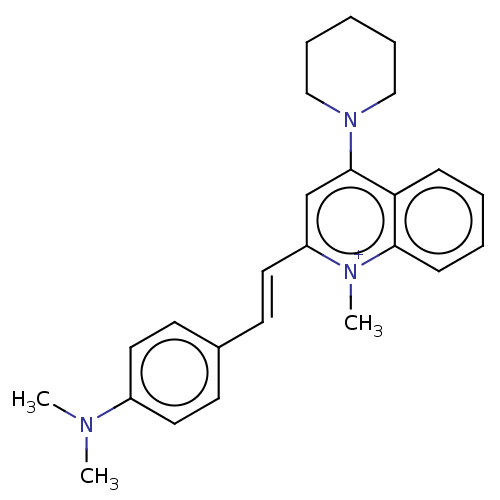
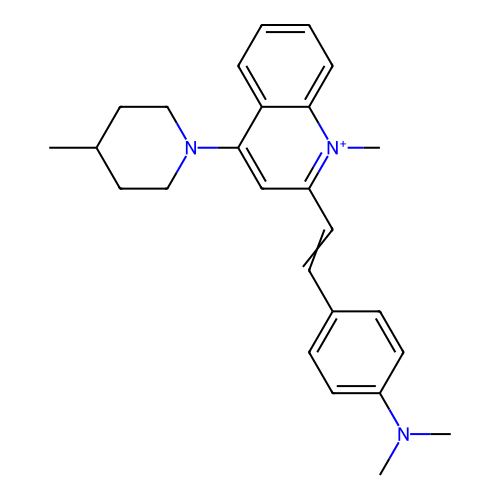
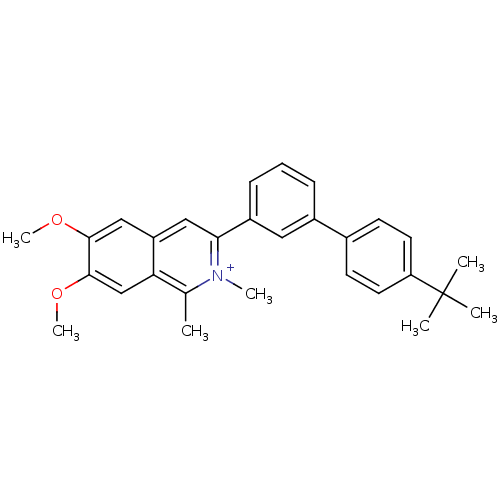
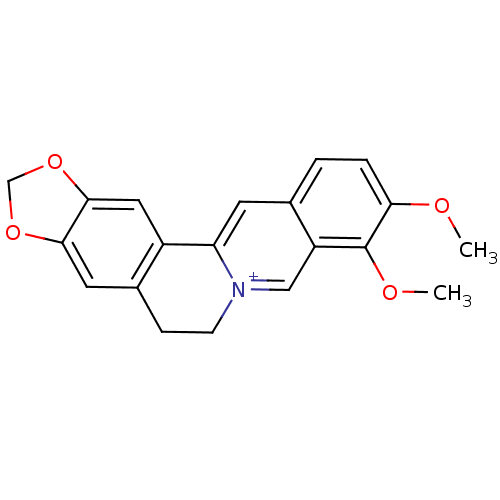
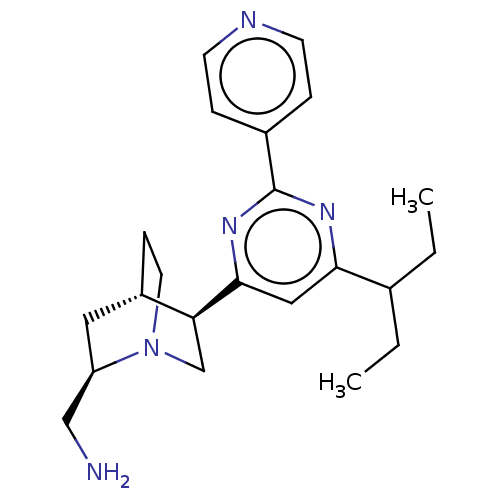
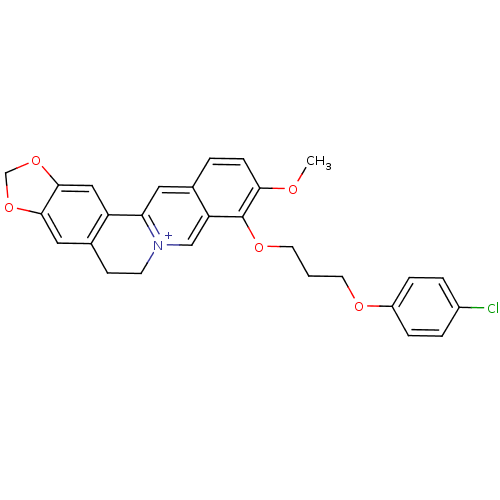
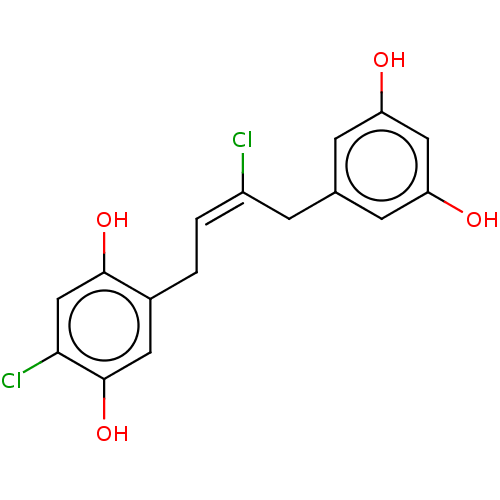
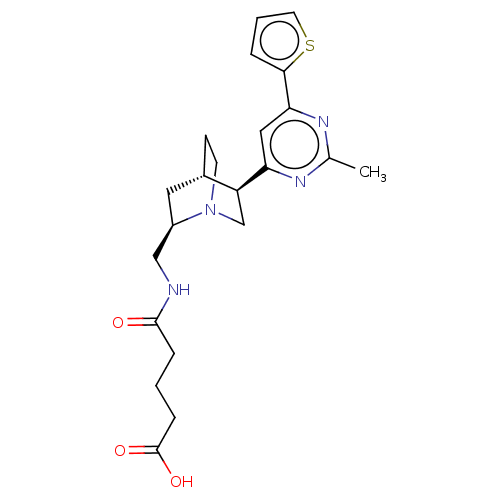
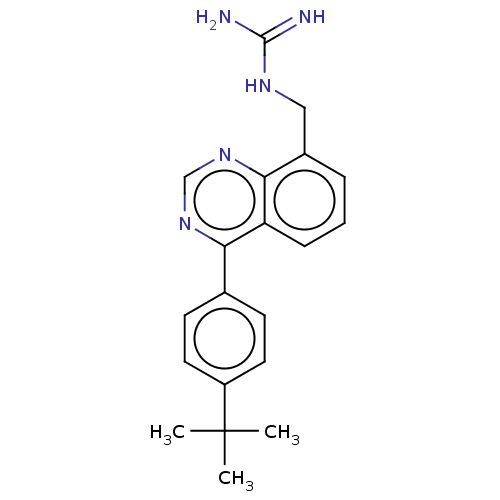
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataIC50: 150nMAssay Description:Inhibition of Staphylococcus aureus FTsZ GTPase activityMore data for this Ligand-Target Pair
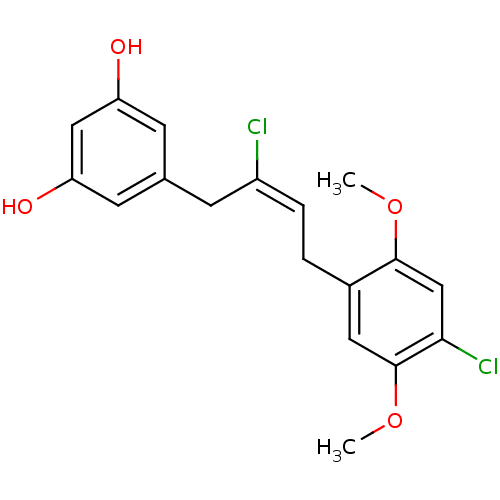
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataIC50: 154nMAssay Description:Inhibition of Staphylococcus aureus FtsZ GTPase activityMore data for this Ligand-Target Pair
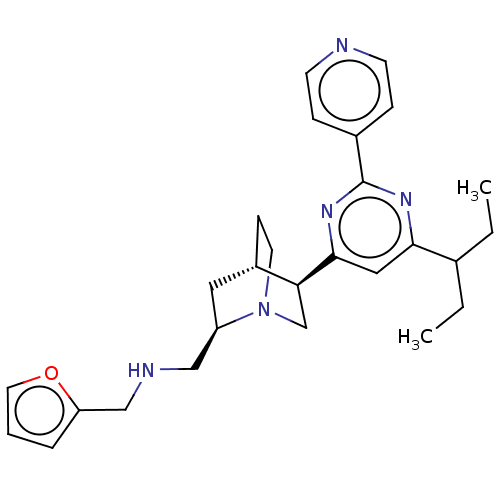
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataIC50: 154nMAssay Description:Inhibition of Staphylococcus aureus FtsZ GTPase activityMore data for this Ligand-Target Pair
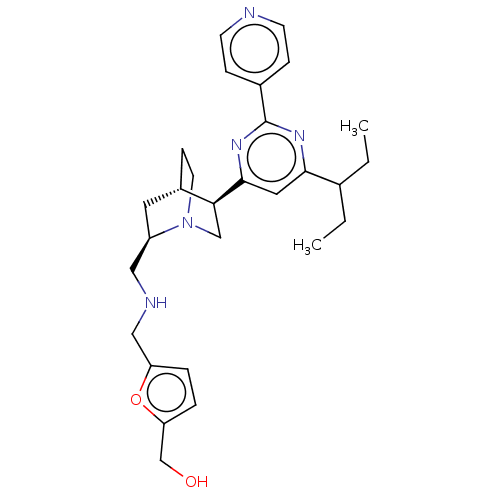
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataIC50: 154nMAssay Description:Inhibition of Staphylococcus aureus FtsZ GTPase activityMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataKd: 600nMAssay Description:Binding affinity to Staphylococcus aureus FtsZ assessed as dissociation constant by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataKd: 2.00E+3nMAssay Description:Binding affinity to Staphylococcus aureus FtsZ expressed in Escherichia coli cells assessed as dissociation constant by fluorescence spectroscopyMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataKd: 2.38E+3nMAssay Description:Binding affinity to Staphylococcus aureus FtsZ assessed as dissociation constant by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataKd: 2.49E+3nMAssay Description:Binding affinity to Staphylococcus aureus FtsZ assessed as dissociation constant by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataKd: 3.50E+3nMAssay Description:Binding affinity to Staphylococcus aureus FtsZ expressed in Escherichia coli BL21 (DE3) at 25 degC by fluorescence anisotropy assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataKd: 3.84E+3nMAssay Description:Binding affinity to Staphylococcus aureus FtsZ assessed as dissociation constant by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataKd: 3.86E+3nMAssay Description:Binding affinity to Staphylococcus aureus FtsZ assessed as dissociation constant by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataEC50: 4.00E+3nMAssay Description:Activation of Staphylococcus aureus FtsZ GTPase activityMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataKd: 5.40E+3nMAssay Description:Binding affinity to Staphylococcus aureus FtsZ expressed in Escherichia coli cells assessed as dissociation constant by fluorescence spectroscopyMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Staphylococcus aureus recombinant FTsZ preincubated for 10 mins followed by compound addition measured after 30 mins by Cytophos phosph...More data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataIC50: 3.75E+4nMAssay Description:Inhibition of Staphylococcus aureus ATCC 29213 N-terminal His6-tagged FtsZ GTPase activity expressed in Escherichia coli BL21(DE3) assessed as reduct...More data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataIC50: 3.75E+4nMAssay Description:Inhibition of Staphylococcus aureus recombinant FTsZ preincubated for 10 mins followed by compound addition measured after 30 mins by Cytophos phosph...More data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataIC50: 3.78E+4nMAssay Description:Inhibition of Staphylococcus aureus recombinant FTsZ preincubated for 10 mins followed by compound addition measured after 30 mins by Cytophos phosph...More data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataIC50: 3.80E+4nMAssay Description:Inhibition of recombinant Staphylococcus aureus FtsZ GTPase activity expressed in Escherichia coli BL21 (DE3) assessed as production of inorganic pho...More data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataIC50: 3.80E+4nMAssay Description:Inhibition of Staphylococcus aureus FtsZ GTPase activityMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataIC50: 4.40E+4nMpH: 6.5 T: 2°CAssay Description:GTPase assays were performed in 50 mM MES, pH 6.5, 50 mM KCl, 5 mM MgCl2 in the presence of 2 uM FtsZ in the presence of varying concentrations of co...More data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataIC50: 5.57E+4nMAssay Description:Inhibition of Staphylococcus aureus recombinant FTsZ preincubated for 10 mins followed by compound addition measured after 30 mins by Cytophos phosph...More data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataIC50: 6.70E+4nMAssay Description:Inhibition of recombinant Staphylococcus aureus FtsZ GTPase activity expressed in Escherichia coli BL21 (DE3) assessed as production of inorganic pho...More data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataIC50: 7.72E+4nMAssay Description:Inhibition of Staphylococcus aureus recombinant FTsZ preincubated for 10 mins followed by compound addition measured after 30 mins by Cytophos phosph...More data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataIC50: 7.74E+4nMAssay Description:Inhibition of Staphylococcus aureus N-terminal His6-tagged FtsZ GTPase activity expressed in Escherichia coli BL21(DE3) preincubated for 10 mins foll...More data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataIC50: 8.80E+4nMpH: 6.5 T: 2°CAssay Description:GTPase assays were performed in 50 mM MES, pH 6.5, 50 mM KCl, 5 mM MgCl2 in the presence of 2 uM FtsZ in the presence of varying concentrations of co...More data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataIC50: 3.17E+5nMAssay Description:Inhibition of Staphylococcus aureus recombinant FTsZ preincubated for 10 mins followed by compound addition measured after 30 mins by Cytophos phosph...More data for this Ligand-Target Pair
TargetCell division protein FtsZ(Staphylococcus aureus)
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Indian Institute of Science Education and Research Bhopal
Curated by ChEMBL
Affinity DataIC50: 3.17E+5nMAssay Description:Inhibition of Staphylococcus aureus ATCC 29213 N-terminal His6-tagged FtsZ GTPase activity expressed in Escherichia coli BL21(DE3) assessed as reduct...More data for this Ligand-Target Pair
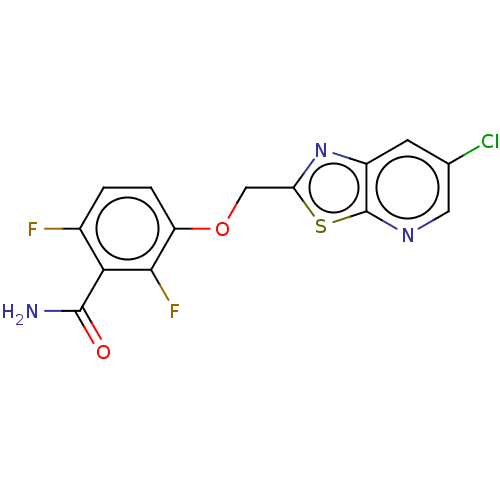
 3D Structure (crystal)
3D Structure (crystal)