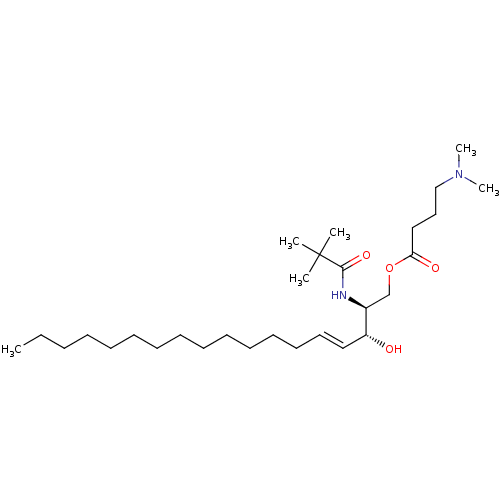
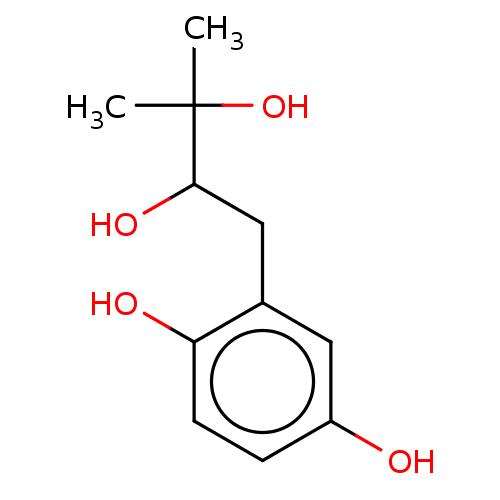
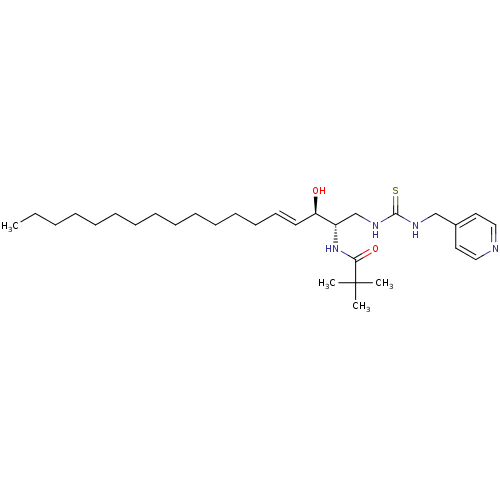
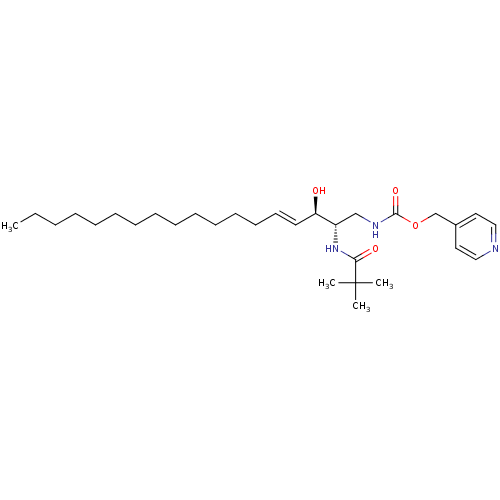
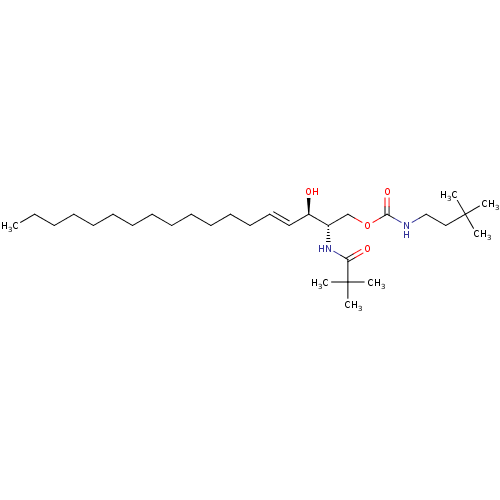
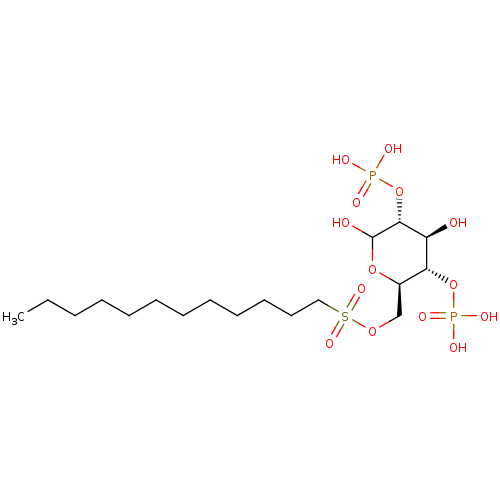
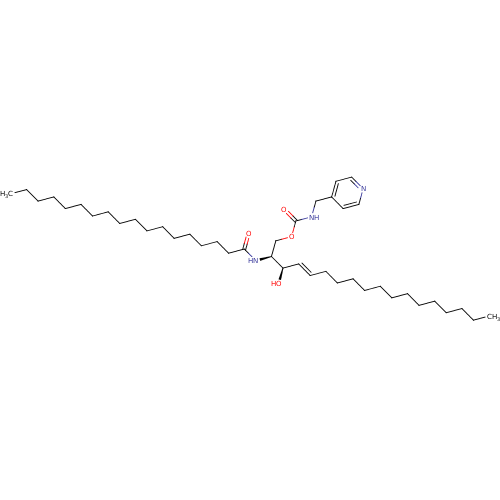
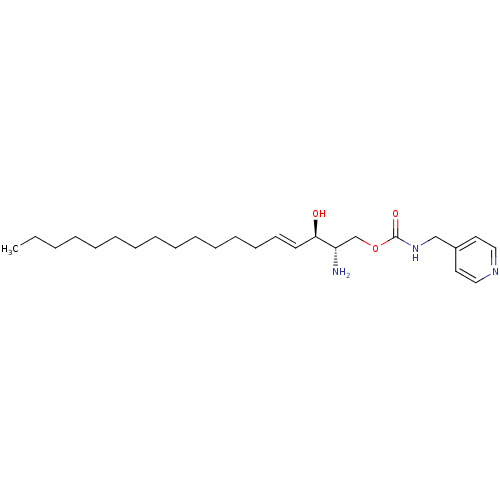
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of rat brain microsomes nSMaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibitory concentration against the membrane neutral magnesium-dependent Sphingomyelinase (N-SMase) using rat brain microsomes as the enzyme sourceMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of rat brain nSMaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibitory concentration against neutral sphingomyelinase (N-Smase) from rat brain microsomesMore data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibition of rat brain microsomes nSMaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibitory concentration against the membrane neutral magnesium-dependent Sphingomyelinase (N-SMase) using rat brain microsomes as the enzyme sourceMore data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+3nMAssay Description:Inhibitory concentration against the membrane neutral magnesium-dependent Sphingomyelinase (N-SMase) using rat brain microsomes as the enzyme sourceMore data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+3nMAssay Description:Inhibitory activity against N-SMase (neutral magnesium- dependent-sphingomyelinase) using rat brain microsomesMore data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+3nMAssay Description:Inhibitory concentration against the membrane neutral magnesium-dependent Sphingomyelinase (N-SMase) using rat brain microsomes as the enzyme sourceMore data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+3nMAssay Description:Inhibition of rat brain microsomes nSMaseMore data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+3nMAssay Description:Inhibitory activity against N-SMase (neutral magnesium- dependent-sphingomyelinase) using rat brain microsomesMore data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+3nMAssay Description:Inhibitory concentration against the membrane neutral magnesium-dependent Sphingomyelinase (N-SMase) using rat brain microsomes as the enzyme sourceMore data for this Ligand-Target Pair
Affinity DataIC50: 4.20E+3nMAssay Description:Inhibition of rat brain microsomes nSMaseMore data for this Ligand-Target Pair
Affinity DataIC50: 4.90E+3nMAssay Description:Inhibitory concentration against the membrane neutral magnesium-dependent Sphingomyelinase (N-SMase) using rat brain microsomes as the enzyme sourceMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibitory concentration against the membrane neutral magnesium-dependent Sphingomyelinase (N-SMase) using rat brain microsomes as the enzyme sourceMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of rat brain microsomes nSMaseMore data for this Ligand-Target Pair
Affinity DataIC50: 6.60E+3nMAssay Description:Inhibitory activity against N-SMase (neutral magnesium- dependent-sphingomyelinase) using rat brain microsomesMore data for this Ligand-Target Pair
Affinity DataIC50: 6.80E+3nMAssay Description:Inhibitory activity against N-SMase (neutral magnesium- dependent-sphingomyelinase) using rat brain microsomesMore data for this Ligand-Target Pair
Affinity DataIC50: 7.90E+3nMAssay Description:Inhibitory concentration against the membrane neutral magnesium-dependent Sphingomyelinase (N-SMase) using rat brain microsomes as the enzyme sourceMore data for this Ligand-Target Pair
Affinity DataIC50: 8.20E+3nMAssay Description:Inhibition of rat brain microsomes nSMaseMore data for this Ligand-Target Pair
Affinity DataIC50: 8.20E+3nMAssay Description:Inhibitory concentration against the membrane neutral magnesium-dependent Sphingomyelinase (N-SMase) using rat brain microsomes as the enzyme sourceMore data for this Ligand-Target Pair
Affinity DataIC50: 8.40E+3nMAssay Description:Inhibitory concentration against the membrane neutral magnesium-dependent Sphingomyelinase (N-SMase) using rat brain microsomes as the enzyme sourceMore data for this Ligand-Target Pair
Affinity DataIC50: 8.70E+3nMAssay Description:Inhibitory concentration against the membrane neutral magnesium-dependent Sphingomyelinase (N-SMase) using rat brain microsomes as the enzyme sourceMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+4nMAssay Description:Inhibition of rat brain microsomes nSMaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+4nMAssay Description:Inhibition of rat brain microsomes nSMaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+4nMAssay Description:Inhibitory concentration against the membrane neutral magnesium-dependent Sphingomyelinase (N-SMase) using rat brain microsomes as the enzyme sourceMore data for this Ligand-Target Pair
Affinity DataKi: 2.00E+4nMAssay Description:Inhibition of rat brain microsomes nSMase assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+4nMAssay Description:Inhibitory activity against N-SMase (neutral magnesium- dependent-sphingomyelinase) using rat brain microsomesMore data for this Ligand-Target Pair
Affinity DataIC50: 3.10E+4nMAssay Description:Inhibitory concentration against the membrane neutral magnesium-dependent Sphingomyelinase (N-SMase) using rat brain microsomes as the enzyme sourceMore data for this Ligand-Target Pair
Affinity DataIC50: 3.40E+4nMAssay Description:Inhibition of rat brain microsomes nSMaseMore data for this Ligand-Target Pair
Affinity DataIC50: 4.50E+4nMAssay Description:Inhibitory activity against N-SMase (neutral magnesium- dependent-sphingomyelinase) using rat brain microsomesMore data for this Ligand-Target Pair
Affinity DataIC50: 4.93E+4nMAssay Description:Inhibitory concentration against neutral sphingomyelinase (N-Smase) from rat brain microsomesMore data for this Ligand-Target Pair
Affinity DataIC50: 5.80E+4nMAssay Description:Inhibitory activity against N-SMase (neutral magnesium- dependent-sphingomyelinase) using rat brain microsomesMore data for this Ligand-Target Pair
Affinity DataIC50: 7.70E+4nMAssay Description:Inhibitory concentration against the membrane neutral magnesium-dependent Sphingomyelinase (N-SMase) using rat brain microsomes as the enzyme sourceMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of nSMase in rat brain microsomal homogenate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of rat brain microsomes nSMaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibitory concentration against the membrane neutral magnesium-dependent Sphingomyelinase (N-SMase) using rat brain microsomes as the enzyme sourceMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of nSMase in rat brain microsomal homogenate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibitory concentration against the membrane neutral magnesium-dependent Sphingomyelinase (N-SMase) using rat brain microsomes as the enzyme sourceMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of rat brain microsomes nSMaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of nSMase in rat brain microsomal homogenate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of nSMase in rat brain microsomal homogenate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of nSMase in rat brain microsomal homogenate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibitory concentration against the membrane neutral magnesium-dependent Sphingomyelinase (N-SMase) using rat brain microsomes as the enzyme sourceMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibitory concentration against the membrane neutral magnesium-dependent Sphingomyelinase (N-SMase) using rat brain microsomes as the enzyme sourceMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibitory activity against N-SMase (neutral magnesium- dependent-sphingomyelinase) using rat brain microsomesMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibitory activity against N-SMase (neutral magnesium- dependent-sphingomyelinase) using rat brain microsomesMore data for this Ligand-Target Pair