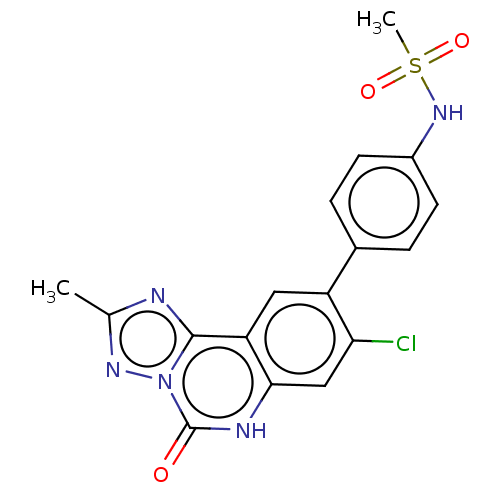
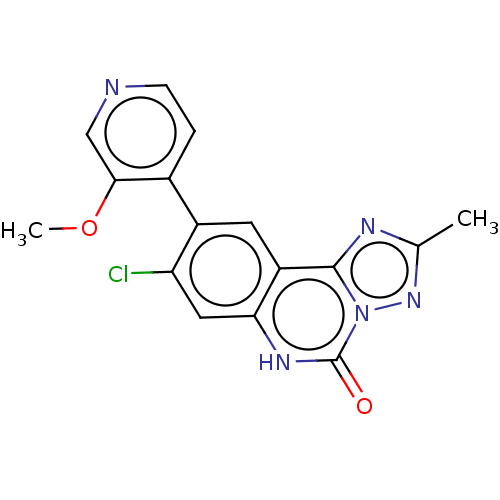
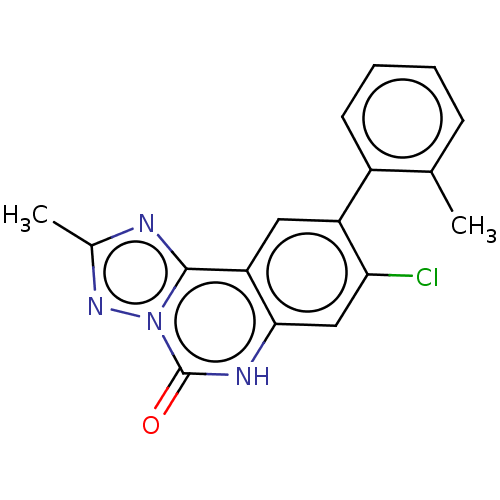
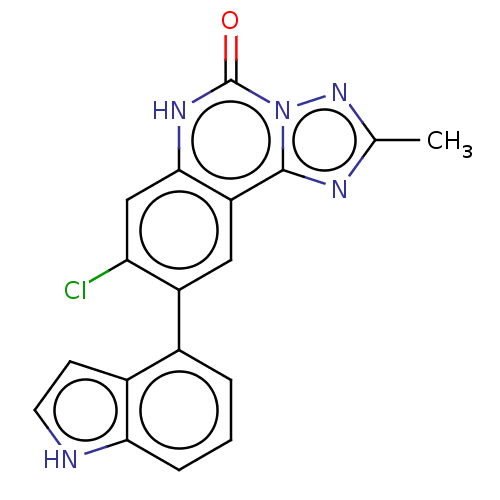
Affinity DataKi: 0.100nMAssay Description:Inhibition of bovine beta-trypsin type-3 using L-BAPNA as substrate preincubated for 5 mins followed by substrate addition measured over 60 minsMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Inhibition of bovine beta-trypsin using N-t-Boc Gln-Ala-Arg-AMC as substrate preincubated with enzyme followed by substrate addition by Dixon plot an...More data for this Ligand-Target Pair
Affinity DataKi: 2.40nMAssay Description:Inhibition of bovine beta-trypsin using BAPNA as substrate preincubated for 15 mins followed by substrate addition measured over 60 mins by Dixon plo...More data for this Ligand-Target Pair
Affinity DataKi: 4.5nMAssay Description:Displacement of [3H]-DAMGO from Wistar rat mu opioid receptor incubated for 1 hr by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 4.90nMAssay Description:Displacement of [3H]-DAMGO from Wistar rat mu opioid receptor incubated for 1 hr by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 5.10nMAssay Description:Inhibition of bovine beta-trypsin using BAPNA as substrate preincubated for 15 mins followed by substrate addition measured over 60 mins by Dixon plo...More data for this Ligand-Target Pair
Affinity DataKi: 5.5nMAssay Description:Displacement of [3H]-DAMGO from Wistar rat mu opioid receptor incubated for 1 hr by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 5.80nMAssay Description:Displacement of [3H]-DAMGO from Wistar rat mu opioid receptor incubated for 1 hr by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 8.70nMAssay Description:Displacement of [3H]-DAMGO from Wistar rat mu opioid receptor incubated for 1 hr by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 9.20nMAssay Description:Displacement of [3H]-DAMGO from Wistar rat mu opioid receptor incubated for 1 hr by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Inhibition of bovine beta-trypsinMore data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Displacement of [3H]-DAMGO from Wistar rat mu opioid receptor incubated for 1 hr by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:Displacement of [3H]-DAMGO from Wistar rat mu opioid receptor incubated for 1 hr by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:Displacement of [3H]-DAMGO from Wistar rat mu opioid receptor incubated for 1 hr by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 22nMAssay Description:Displacement of [3H]-DAMGO from Wistar rat mu opioid receptor incubated for 1 hr by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 32nMAssay Description:Displacement of [3H]-DAMGO from Wistar rat mu opioid receptor incubated for 1 hr by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 34nMAssay Description:Displacement of [3H]-DAMGO from Wistar rat mu opioid receptor incubated for 1 hr by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 53nMAssay Description:Displacement of [3H]-DAMGO from Wistar rat mu opioid receptor incubated for 1 hr by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 70nMAssay Description:Displacement of FITC-labelled SP2 probe from human TIM-3 IgV domain (residues 22 to 130) expressed in Escherichia coli BL21 (DE3) by FPA competition ...More data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 110nMAssay Description:Displacement of FITC-labelled SP2 probe from human TIM-3 IgV domain (residues 22 to 130) expressed in Escherichia coli BL21 (DE3) by FPA competition ...More data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 156nMAssay Description:Displacement of FITC-labelled SP2 probe from human TIM-3 IgV domain (residues 22 to 130) expressed in Escherichia coli BL21 (DE3) by FPA competition ...More data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 164nMAssay Description:Displacement of FITC-labelled SP2 probe from human TIM-3 IgV domain (residues 22 to 130) expressed in Escherichia coli BL21 (DE3) by FPA competition ...More data for this Ligand-Target Pair
Affinity DataKi: 172nMAssay Description:Displacement of [3H]-DAMGO from Wistar rat mu opioid receptor incubated for 1 hr by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 204nMAssay Description:Displacement of FITC-labelled SP2 probe from human TIM-3 IgV domain (residues 22 to 130) expressed in Escherichia coli BL21 (DE3) by FPA competition ...More data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 287nMAssay Description:Displacement of FITC-labelled SP2 probe from human TIM-3 IgV domain (residues 22 to 130) expressed in Escherichia coli BL21 (DE3) by FPA competition ...More data for this Ligand-Target Pair
Affinity DataKi: 320nMAssay Description:Displacement of [3H]-DPDPE from Wistar rat delta opioid receptor incubated for 3 hrs by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 405nMAssay Description:Displacement of [3H]-DAMGO from Wistar rat mu opioid receptor incubated for 1 hr by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 437nMAssay Description:Displacement of [3H]-DAMGO from Wistar rat mu opioid receptor incubated for 1 hr by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 440nMAssay Description:Displacement of FITC-labelled SP2 probe from human TIM-3 IgV domain (residues 22 to 130) expressed in Escherichia coli BL21 (DE3) by FPA competition ...More data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 500nMAssay Description:Displacement of FITC-labelled SP2 probe from human TIM-3 IgV domain (residues 22 to 130) expressed in Escherichia coli BL21 (DE3) by FPA competition ...More data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
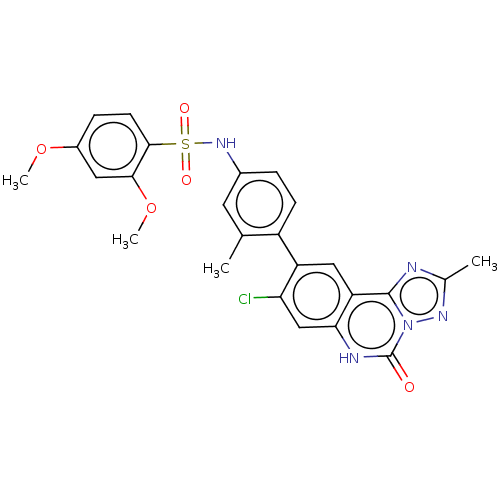
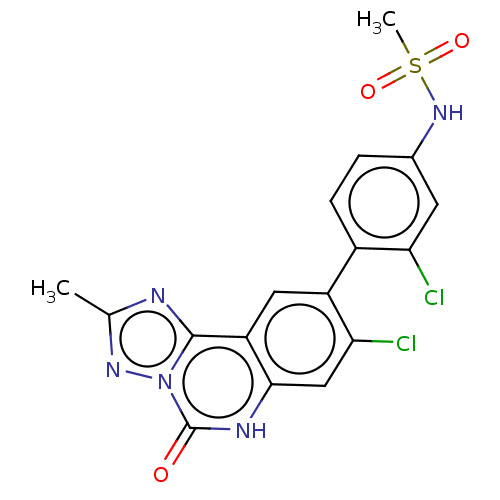
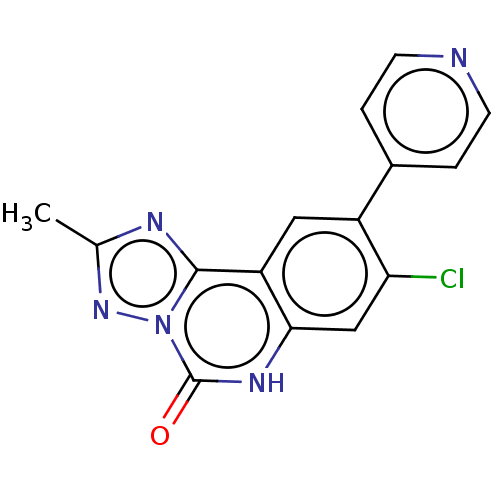
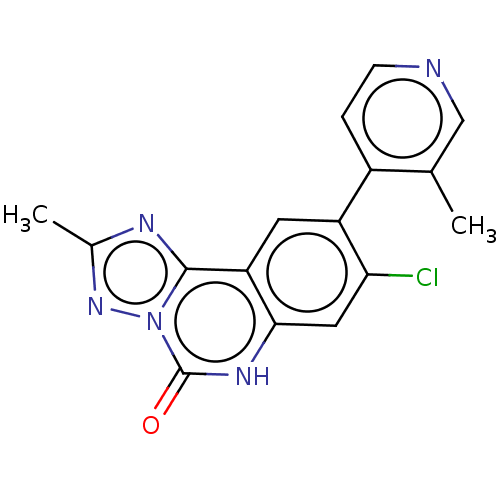
Affinity DataKi: 550nMAssay Description:Displacement of FITC-labelled 5-(((8-Chloro-9-(3-methylpyridin-4-yl)-5-oxo-5,6-dihydro-[1,2,4]triazolo[1,5-c]quinazolin-2-yl)methyl)carbamoyl)-2-(6-h...More data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 750nMAssay Description:Displacement of FITC-labelled 5-(((8-Chloro-9-(3-methylpyridin-4-yl)-5-oxo-5,6-dihydro-[1,2,4]triazolo[1,5-c]quinazolin-2-yl)methyl)carbamoyl)-2-(6-h...More data for this Ligand-Target Pair
Affinity DataKi: 922nMAssay Description:Displacement of [3H]-DPDPE from Wistar rat delta opioid receptor incubated for 3 hrs by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 955nMAssay Description:Displacement of [3H]-DPDPE from Wistar rat delta opioid receptor incubated for 3 hrs by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 970nMAssay Description:Displacement of FITC-labelled 5-(((8-Chloro-9-(3-methylpyridin-4-yl)-5-oxo-5,6-dihydro-[1,2,4]triazolo[1,5-c]quinazolin-2-yl)methyl)carbamoyl)-2-(6-h...More data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 1.30E+3nMAssay Description:Displacement of FITC-labelled 5-(((8-Chloro-9-(3-methylpyridin-4-yl)-5-oxo-5,6-dihydro-[1,2,4]triazolo[1,5-c]quinazolin-2-yl)methyl)carbamoyl)-2-(6-h...More data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 1.50E+3nMAssay Description:Displacement of FITC-labelled 5-(((8-Chloro-9-(3-methylpyridin-4-yl)-5-oxo-5,6-dihydro-[1,2,4]triazolo[1,5-c]quinazolin-2-yl)methyl)carbamoyl)-2-(6-h...More data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 1.50E+3nMAssay Description:Displacement of FITC-labelled 5-(((8-Chloro-9-(3-methylpyridin-4-yl)-5-oxo-5,6-dihydro-[1,2,4]triazolo[1,5-c]quinazolin-2-yl)methyl)carbamoyl)-2-(6-h...More data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 1.60E+3nMAssay Description:Displacement of FITC-labelled 5-(((8-Chloro-9-(3-methylpyridin-4-yl)-5-oxo-5,6-dihydro-[1,2,4]triazolo[1,5-c]quinazolin-2-yl)methyl)carbamoyl)-2-(6-h...More data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 1.80E+3nMAssay Description:Displacement of FITC-labelled 5-(((8-Chloro-9-(3-methylpyridin-4-yl)-5-oxo-5,6-dihydro-[1,2,4]triazolo[1,5-c]quinazolin-2-yl)methyl)carbamoyl)-2-(6-h...More data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 2.10E+3nMAssay Description:Displacement of FITC-labelled 5-(((8-Chloro-9-(3-methylpyridin-4-yl)-5-oxo-5,6-dihydro-[1,2,4]triazolo[1,5-c]quinazolin-2-yl)methyl)carbamoyl)-2-(6-h...More data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 2.20E+3nMAssay Description:Displacement of FITC-labelled 5-(((8-Chloro-9-(3-methylpyridin-4-yl)-5-oxo-5,6-dihydro-[1,2,4]triazolo[1,5-c]quinazolin-2-yl)methyl)carbamoyl)-2-(6-h...More data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 2.30E+3nMAssay Description:Displacement of FITC-labelled 5-(((8-Chloro-9-(3-methylpyridin-4-yl)-5-oxo-5,6-dihydro-[1,2,4]triazolo[1,5-c]quinazolin-2-yl)methyl)carbamoyl)-2-(6-h...More data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 2.60E+3nMAssay Description:Displacement of FITC-labelled 5-(((8-Chloro-9-(3-methylpyridin-4-yl)-5-oxo-5,6-dihydro-[1,2,4]triazolo[1,5-c]quinazolin-2-yl)methyl)carbamoyl)-2-(6-h...More data for this Ligand-Target Pair
Affinity DataKi: 2.66E+3nMAssay Description:Displacement of [3H]-DPDPE from Wistar rat delta opioid receptor incubated for 3 hrs by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 3.58E+3nMAssay Description:Displacement of [3H]-DPDPE from Wistar rat delta opioid receptor incubated for 3 hrs by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetHepatitis A virus cellular receptor 2(Homo sapiens)
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 4.90E+3nMAssay Description:Displacement of FITC-labelled 5-(((8-Chloro-9-(3-methylpyridin-4-yl)-5-oxo-5,6-dihydro-[1,2,4]triazolo[1,5-c]quinazolin-2-yl)methyl)carbamoyl)-2-(6-h...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]-DPDPE from Wistar rat delta opioid receptor incubated for 3 hrs by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]-DPDPE from Wistar rat delta opioid receptor incubated for 3 hrs by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]-DPDPE from Wistar rat delta opioid receptor incubated for 3 hrs by liquid scintillation counting methodMore data for this Ligand-Target Pair

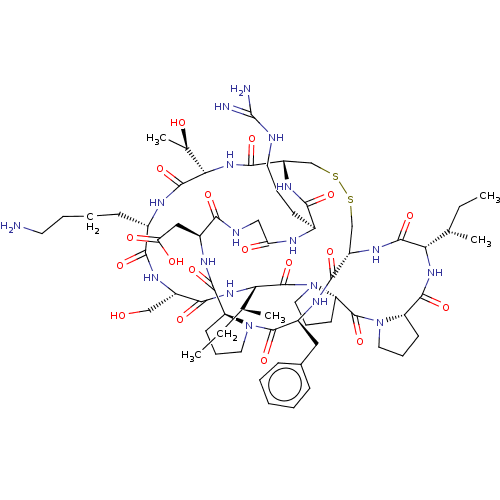
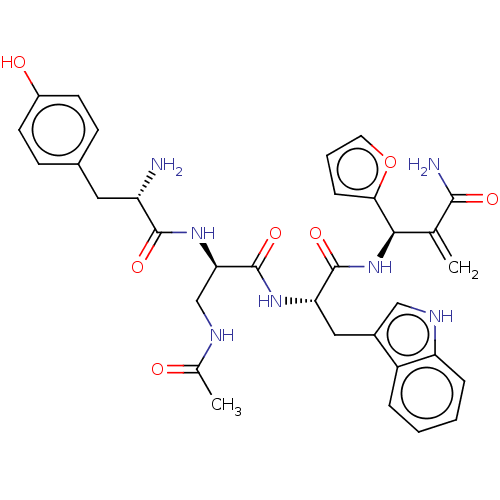
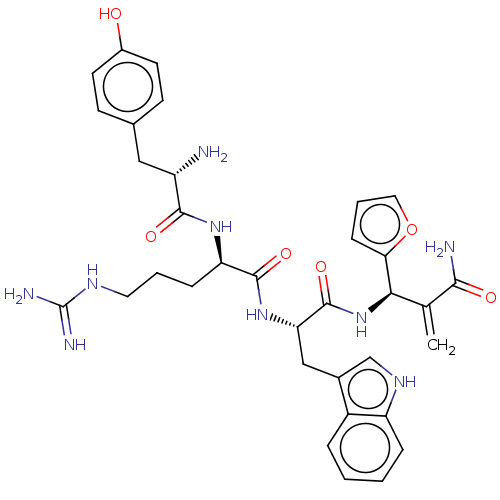
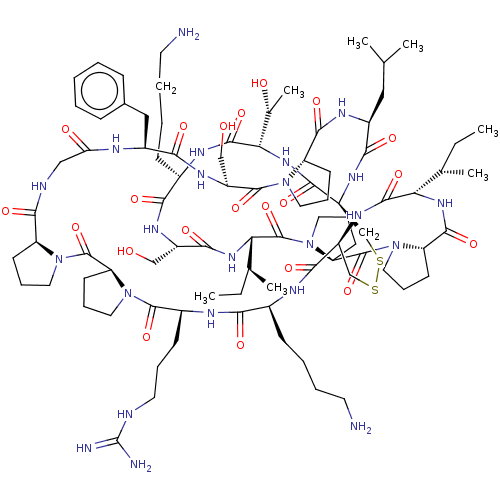
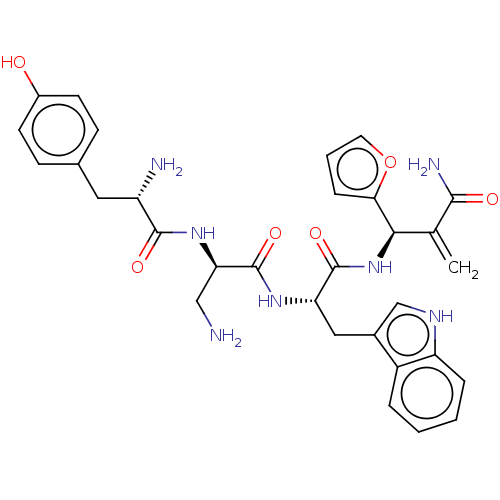
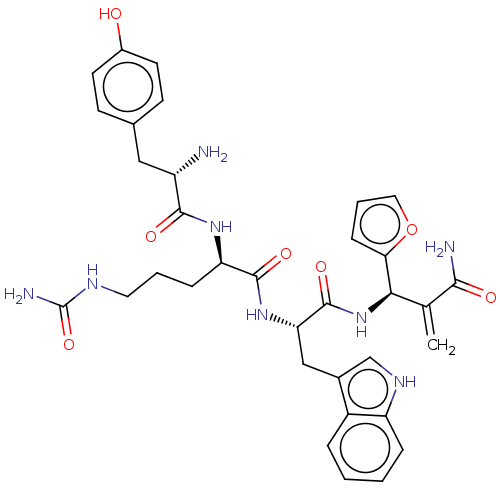
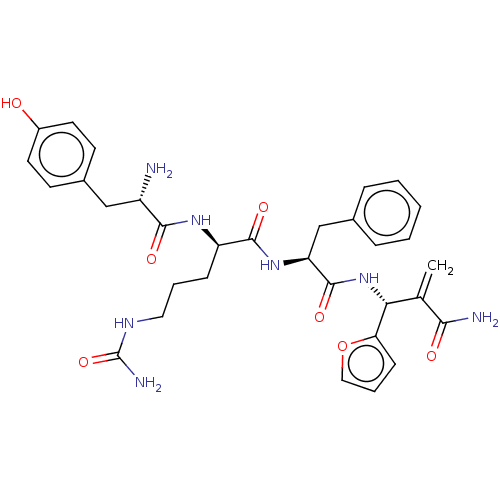
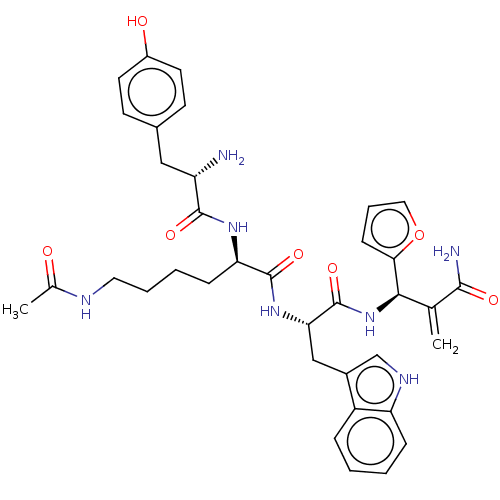
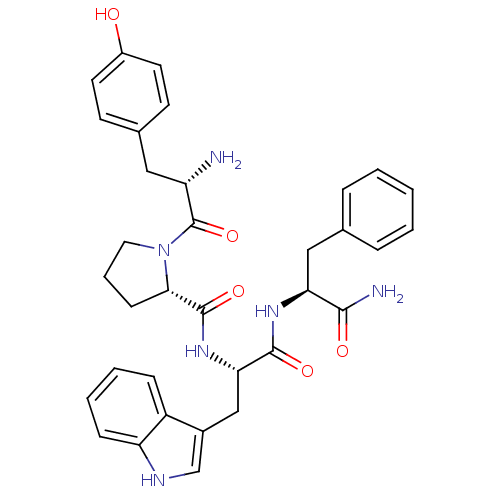
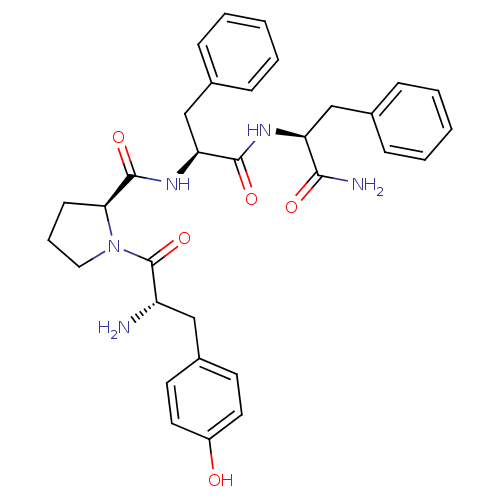
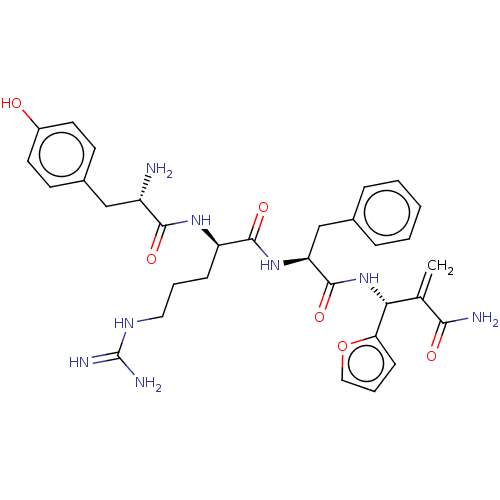
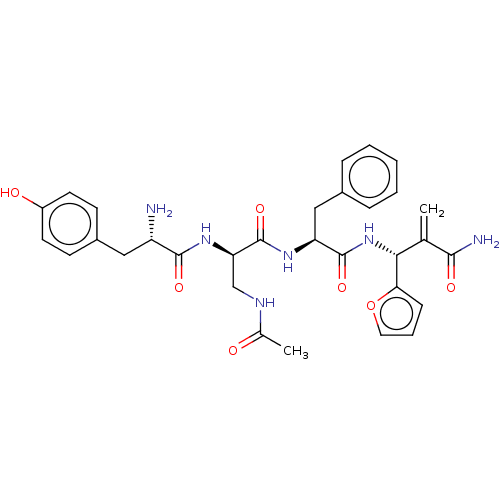
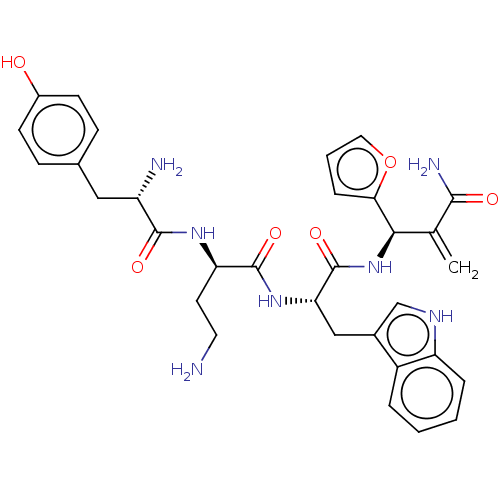
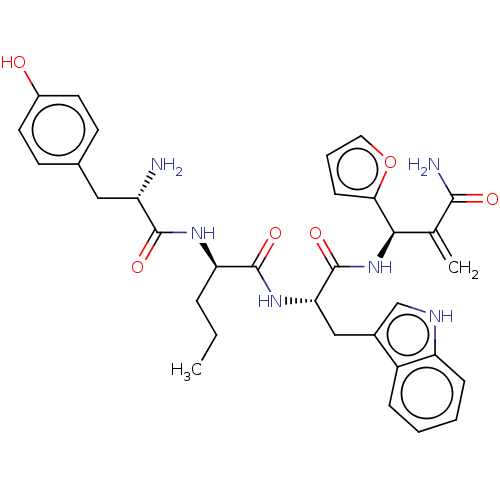
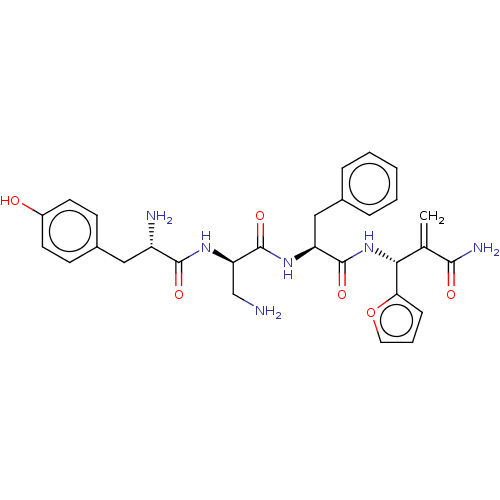

 3D Structure (crystal)
3D Structure (crystal)