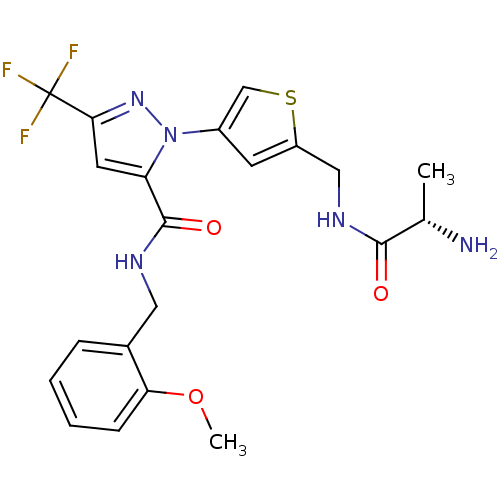
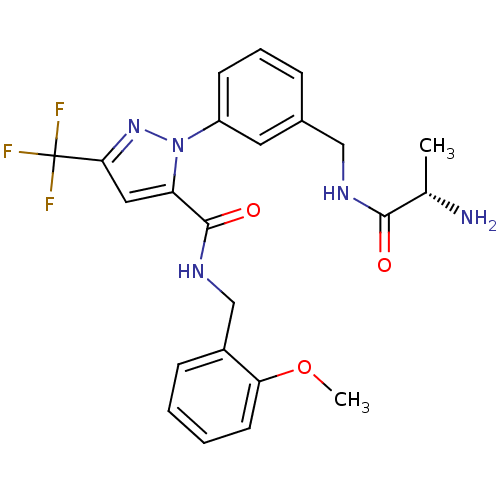
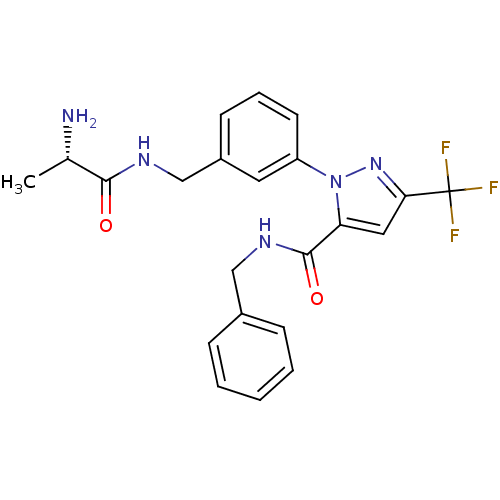
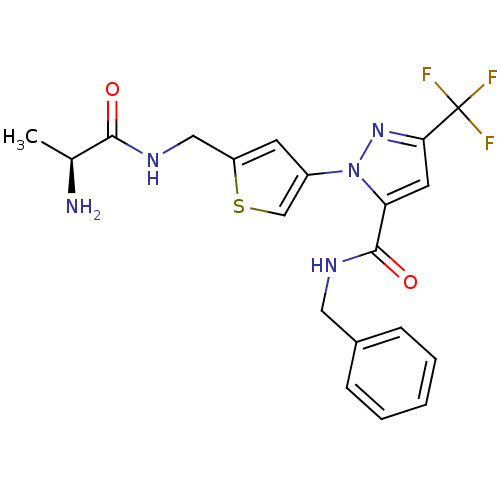
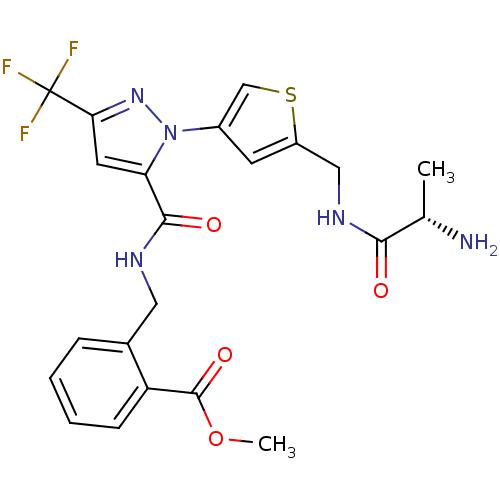
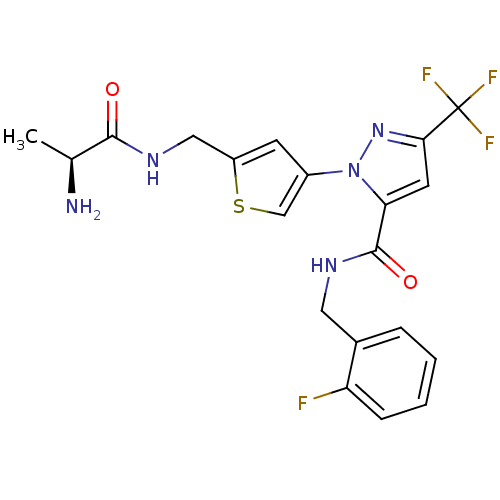
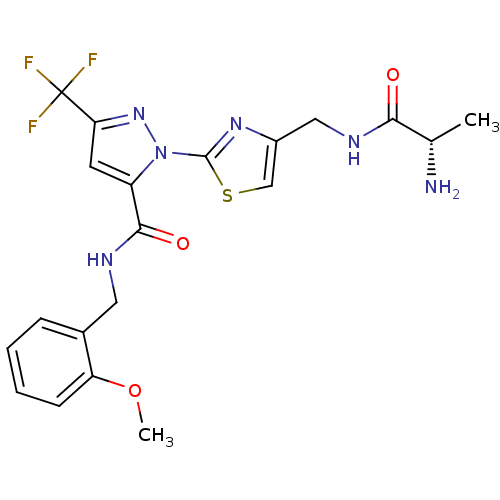
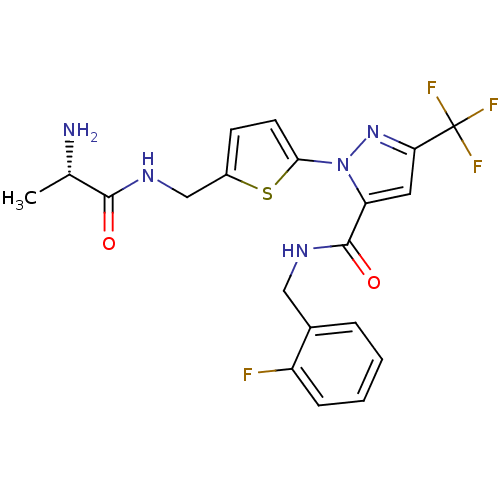
Report error Found 19 Enz. Inhib. hit(s) with all data for entry = 3074
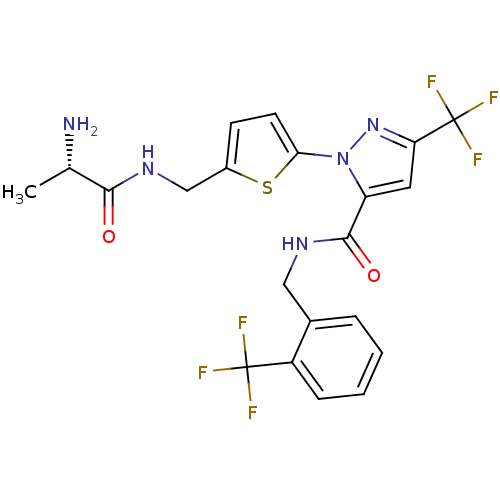
Affinity DataIC50: 60nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair
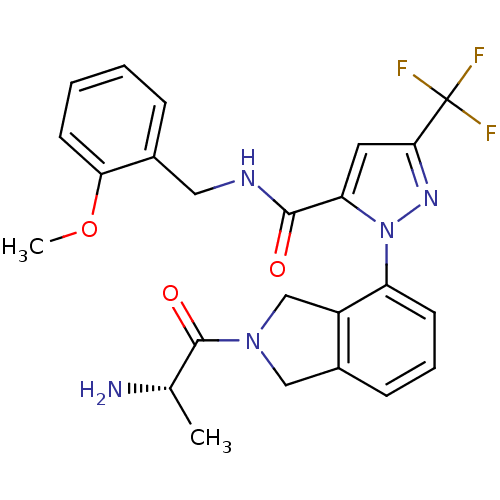
Affinity DataIC50: 60nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair
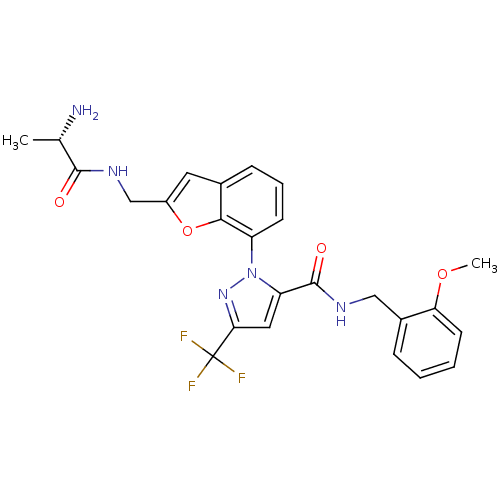
Affinity DataIC50: 80nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair
Affinity DataIC50: 220nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair
Affinity DataIC50: 230nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair
Affinity DataIC50: 260nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair
Affinity DataIC50: 300nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair
Affinity DataIC50: 370nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair
Affinity DataIC50: 770nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair
Affinity DataIC50: 790nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair
Affinity DataIC50: 2.31E+3nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair
Affinity DataIC50: 2.92E+3nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair
Affinity DataIC50: 3.05E+3nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair
Affinity DataIC50: 3.35E+3nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair
Affinity DataIC50: 4.20E+3nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair
Affinity DataIC50: 4.48E+3nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 9.0 T: 2°CAssay Description:Histone H3 is used as the substrate for CARM-1 enzyme, and the methylation is monitored using tritiated S-Adenosyl-Methionine (SAM) as a methyl donor...More data for this Ligand-Target Pair