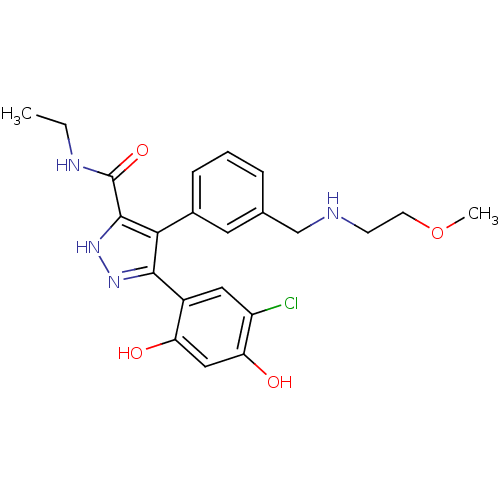
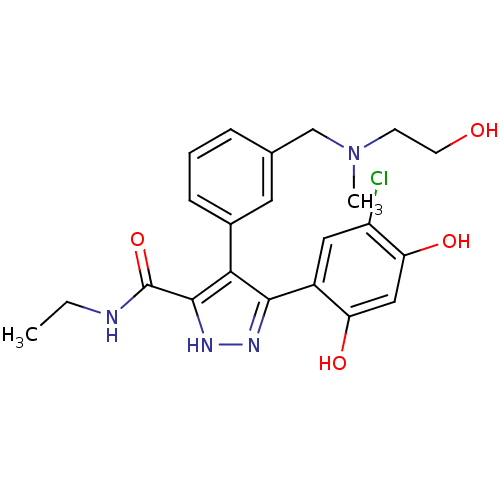
Report error Found 55 Enz. Inhib. hit(s) with all data for entry = 2464
Affinity DataIC50: 6nM EC50: 6nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 6nM EC50: 40nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 8nM EC50: 70nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 11nM EC50: 65nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 13nM EC50: 41nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 14nM EC50: 1.98E+3nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 14nM EC50: 86nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 14nM EC50: 114nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 15nM EC50: 80nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 21nM EC50: 16nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 18nM EC50: 69nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 19nM EC50: 61nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 20nM EC50: 39nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 21nM EC50: 83nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 21nM EC50: 470nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 22nM EC50: 152nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 25nM EC50: 260nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 29nM EC50: 25nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 26nM EC50: 58nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 26nM EC50: 29nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 27nM EC50: 826nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 28nM EC50: 1.29E+3nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 28nM EC50: 120nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 32nM EC50: 29nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 35nM EC50: 2.16E+3nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 36nM EC50: 35nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 35nM EC50: 133nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 37nM EC50: 290nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 39nM EC50: 125nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 39nM EC50: 87nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 39nM EC50: 118nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 64nM EC50: 56nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 57nM EC50: 275nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 74nM EC50: 91nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 115nM EC50: 3.47E+3nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 127nM EC50: 634nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 142nM EC50: 7.80E+3nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 146nM EC50: 315nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 1.27E+3nM EC50: 160nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 222nM EC50: 2.69E+3nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 231nM EC50: 3.11E+3nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 239nM EC50: 6.76E+3nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 258nM EC50: 1.16E+4nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 280nM EC50: 5.80E+3nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 343nM EC50: 2.78E+3nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 431nM EC50: 1.25E+4nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 540nM EC50: 2.15E+3nMAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 600nM EC50: 6.50E+3nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 728nM EC50: 1.29E+4nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair
Affinity DataIC50: 914nM EC50: 9.24E+3nMpH: 7.4 T: 2°CAssay Description:The assay is based upon displacement of a fluorescently labeled molecule, which binds specifically to the ATP-binding site of full-length human Hsp90...More data for this Ligand-Target Pair