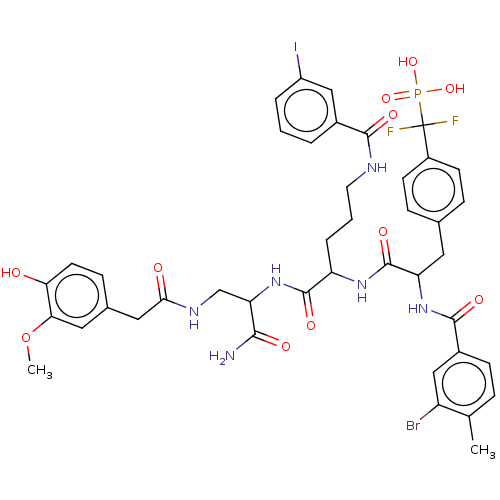
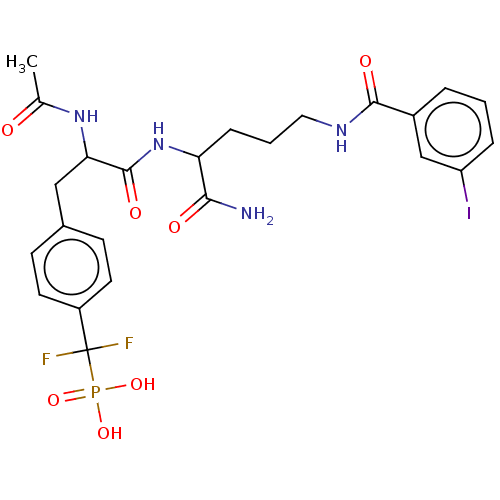
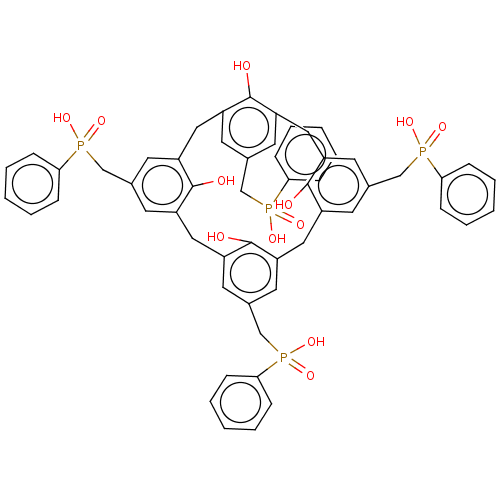
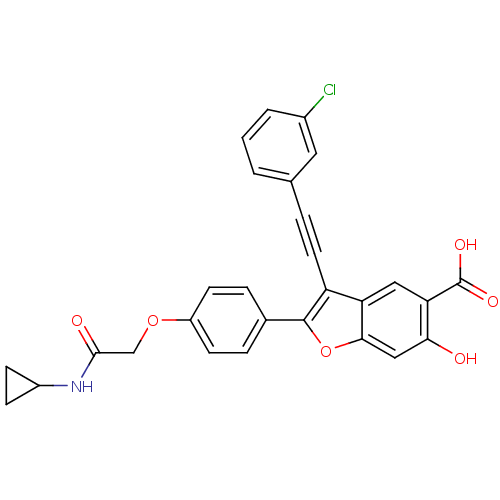
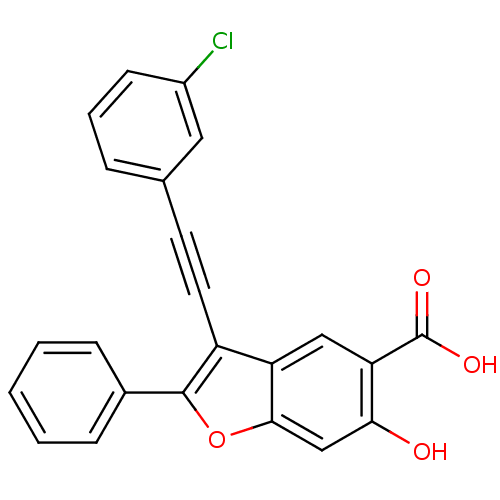
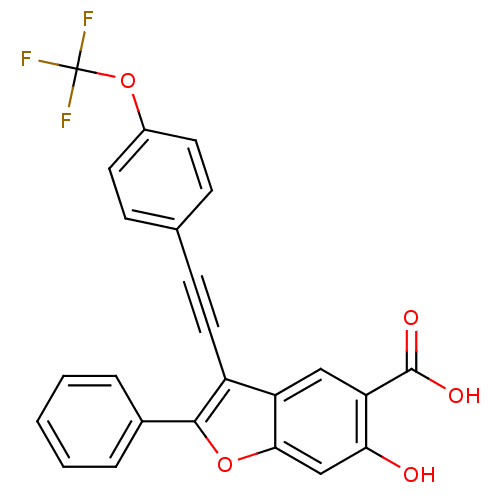
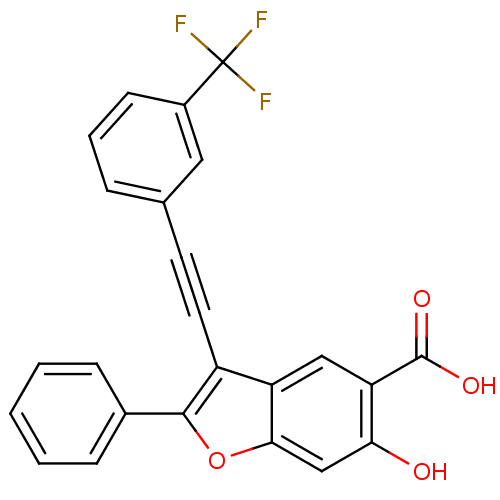
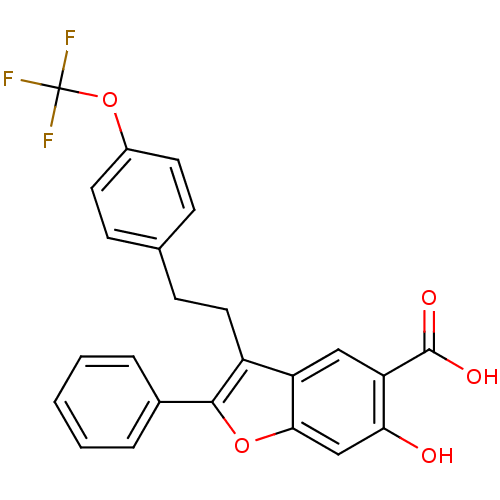
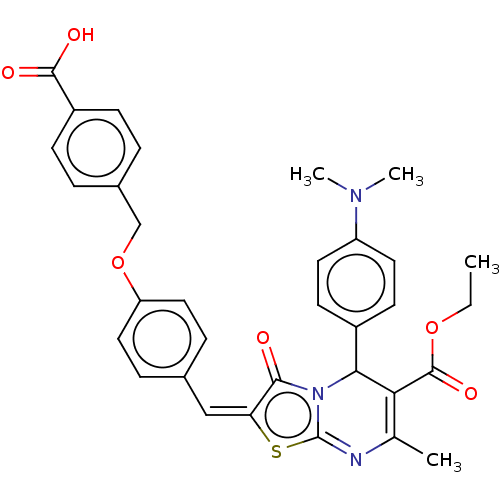
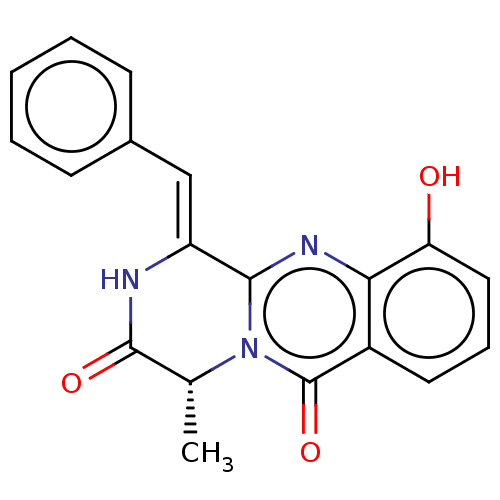
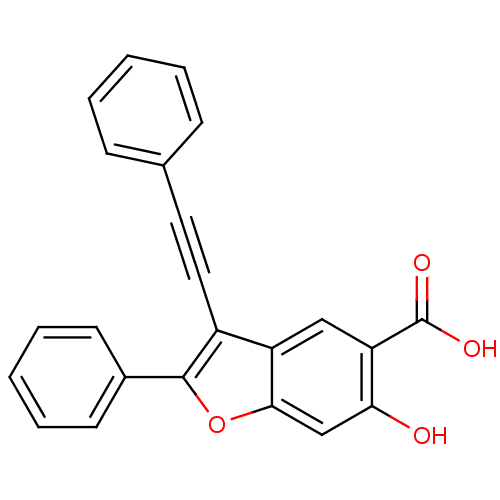
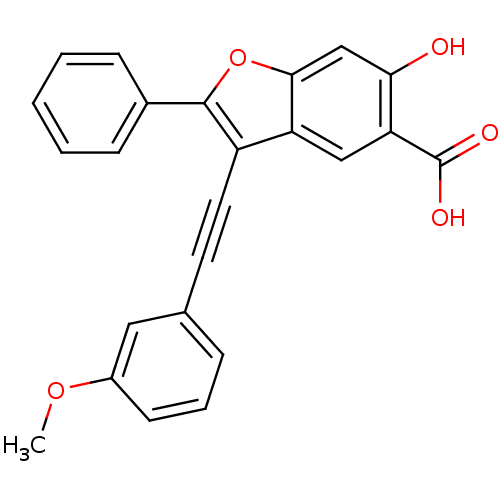
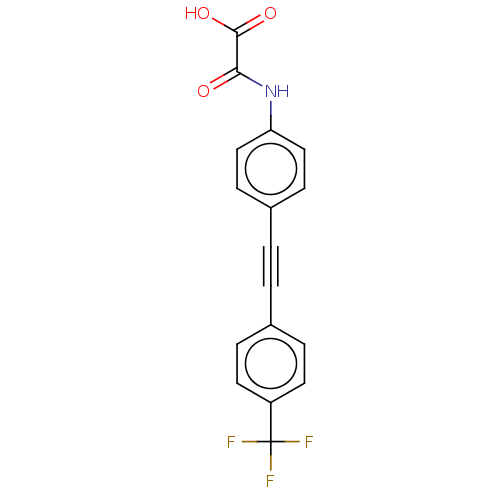
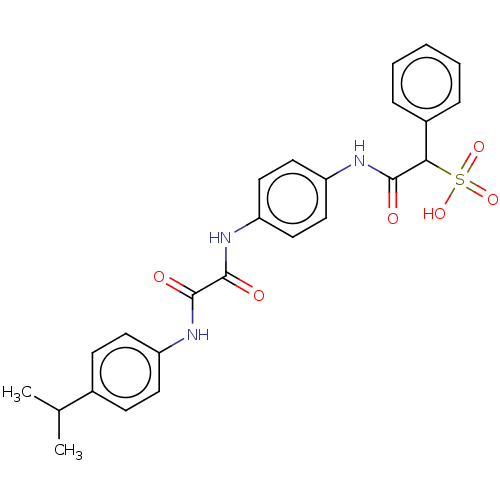
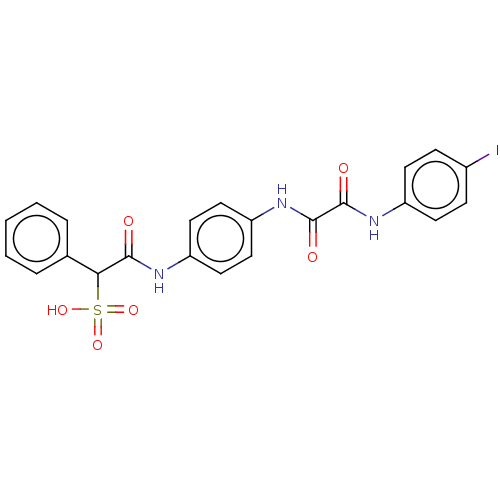
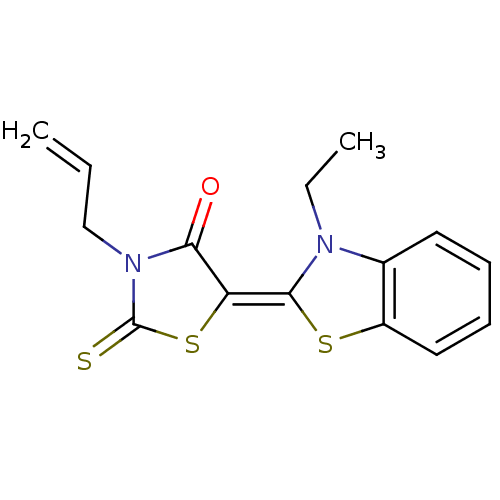
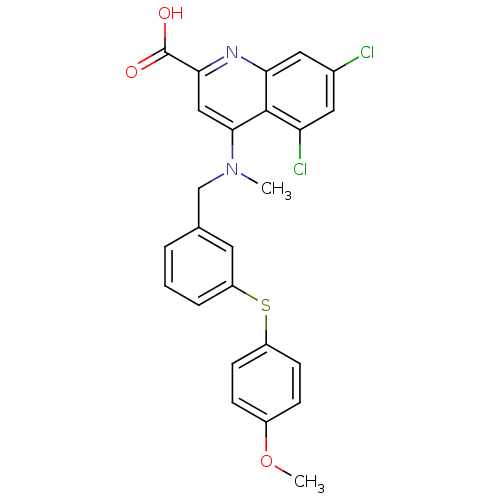
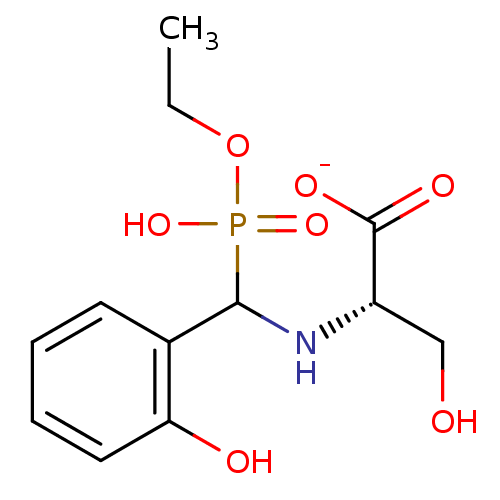
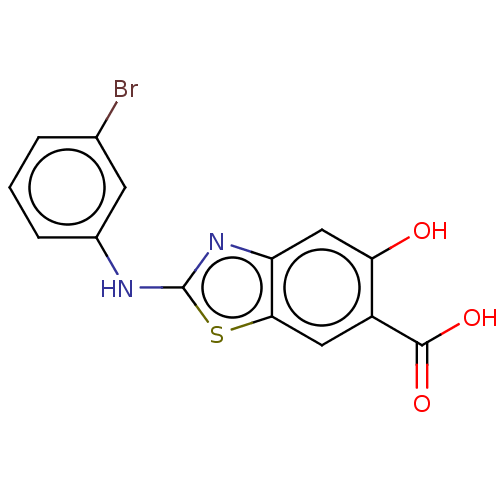
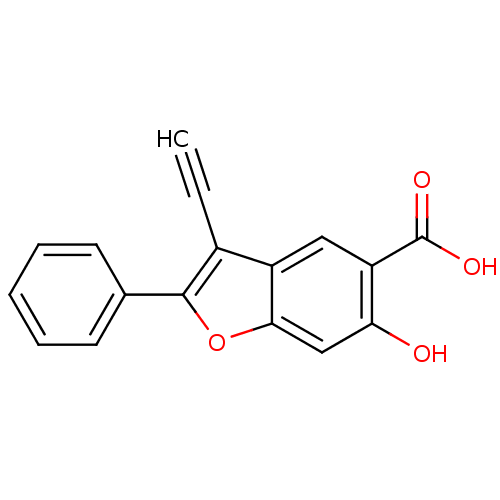
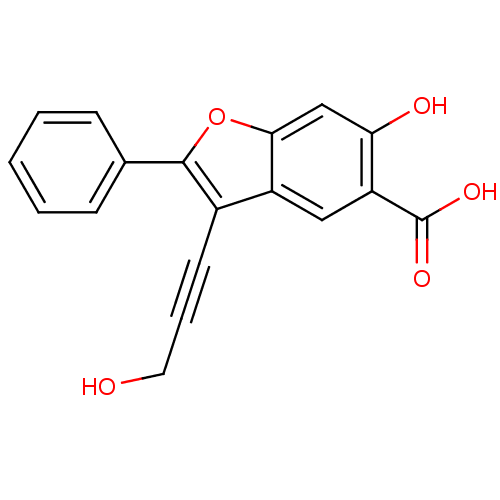
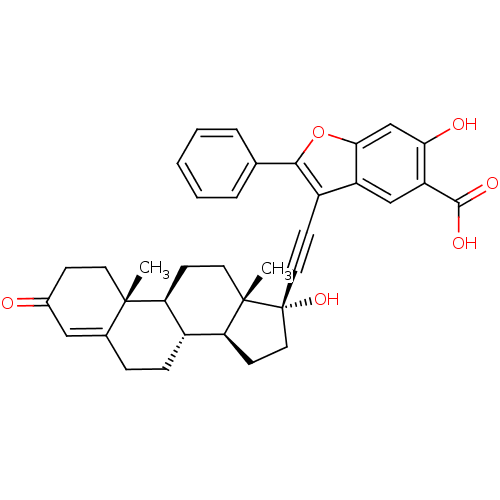
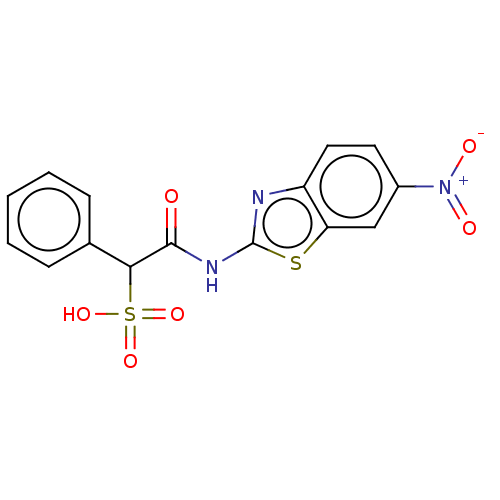
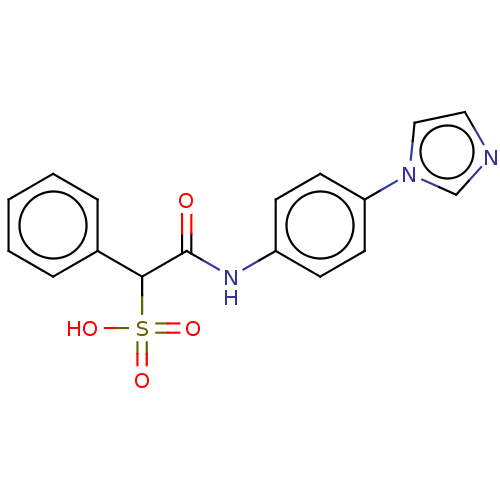
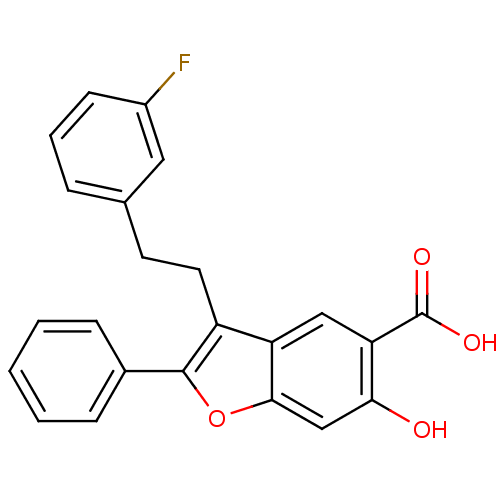
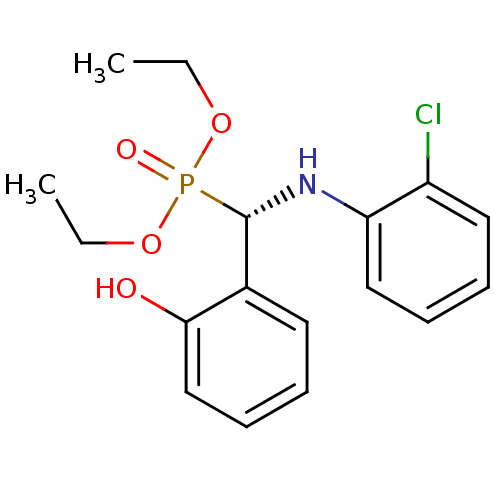
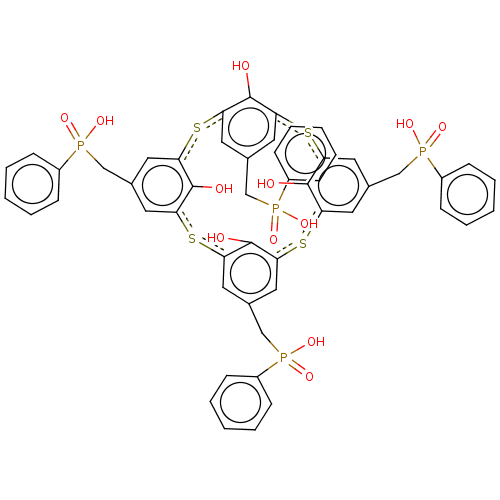TargetTyrosine-protein phosphatase non-receptor type 9 [277-582](Human)
Indiana University Research and Technology
US Patent
Indiana University Research and Technology
US Patent
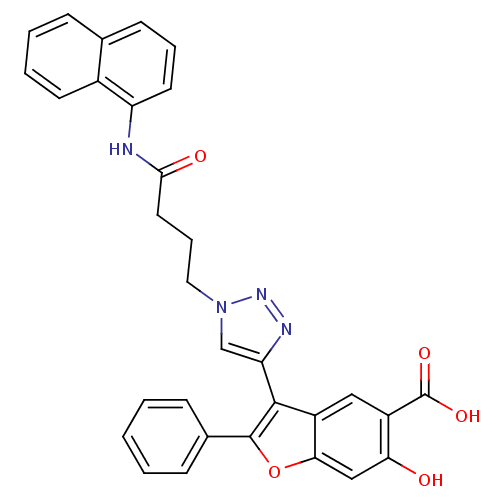
Affinity DataKi: 34nM IC50: 75nMpH: 7.0Assay Description:PTP activity was assayed using p-nitrophenyl phosphate (pNPP) as a substrate in 3,3-dimethylglutarate buffer (50 mM 3,3-dimethylglutarate, pH 7.0, 1 ...More data for this Ligand-Target Pair
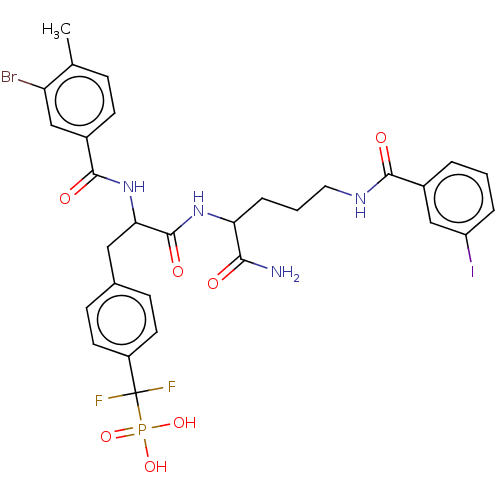
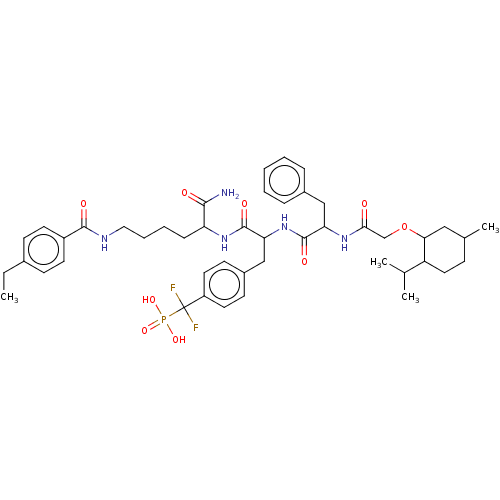
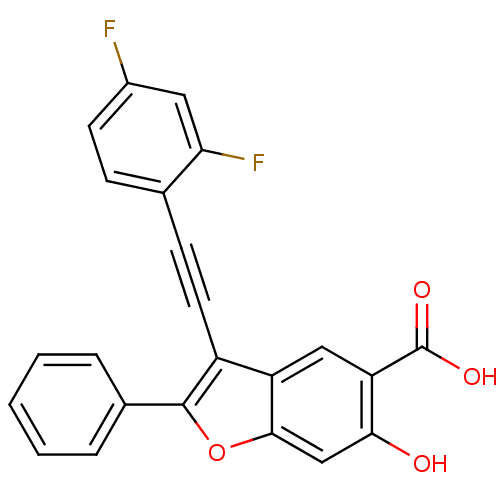
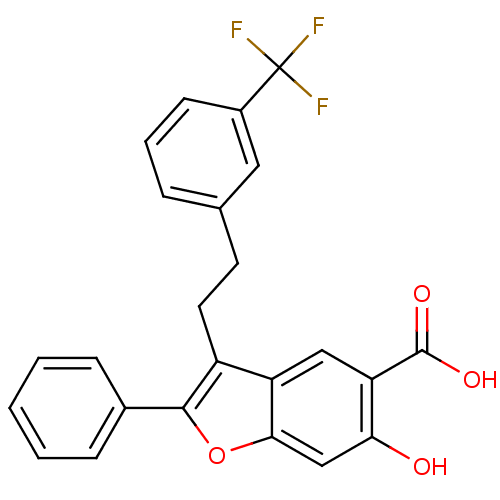
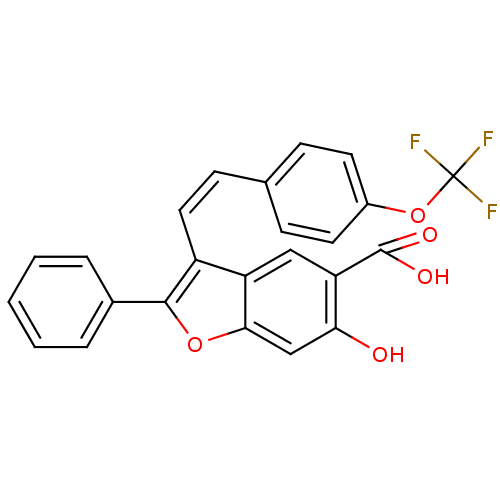
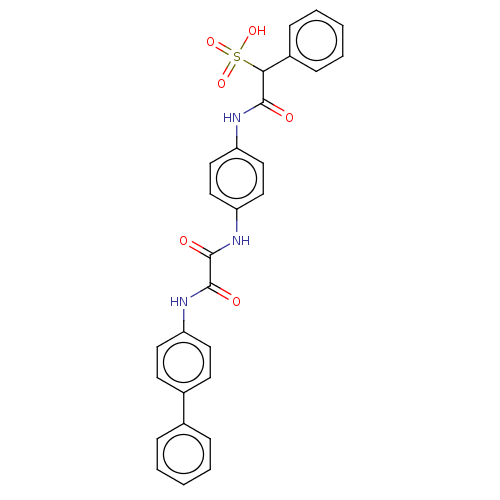
TargetTyrosine-protein phosphatase non-receptor type 9 [277-582](Human)
Indiana University Research and Technology
US Patent
Indiana University Research and Technology
US Patent
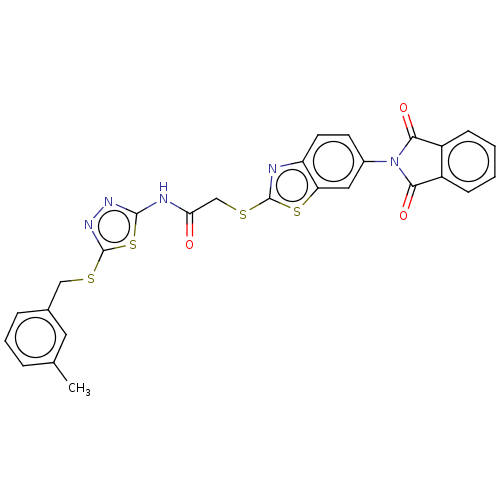
Affinity DataIC50: 270nMpH: 7.0Assay Description:PTP activity was assayed using p-nitrophenyl phosphate (pNPP) as a substrate in 3,3-dimethylglutarate buffer (50 mM 3,3-dimethylglutarate, pH 7.0, 1 ...More data for this Ligand-Target Pair
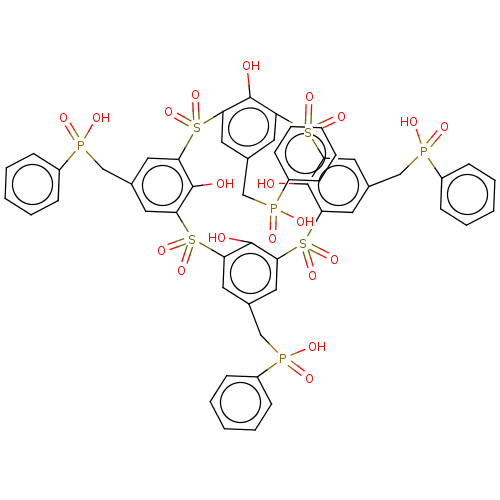
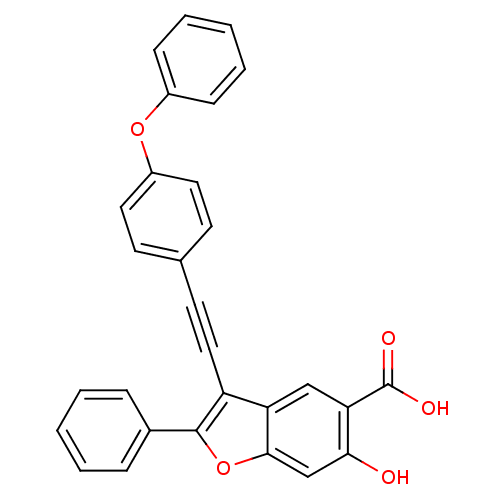
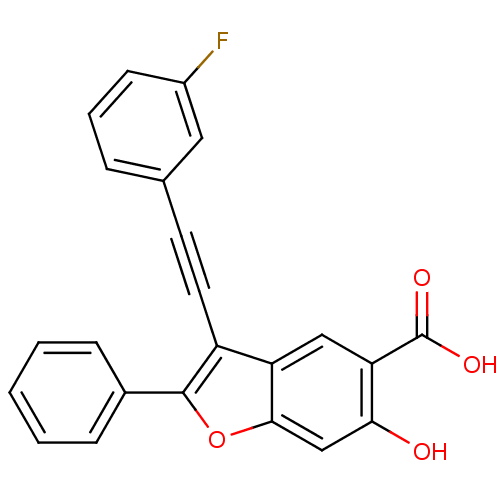
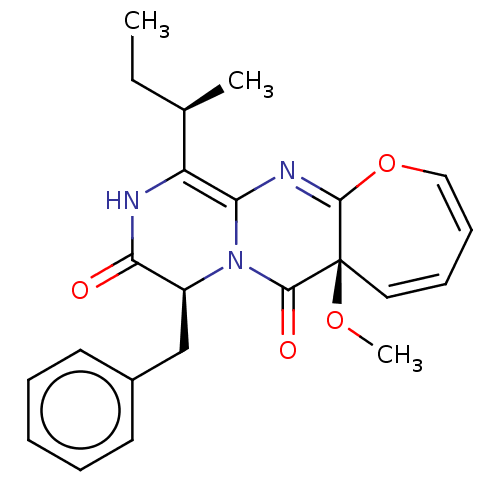
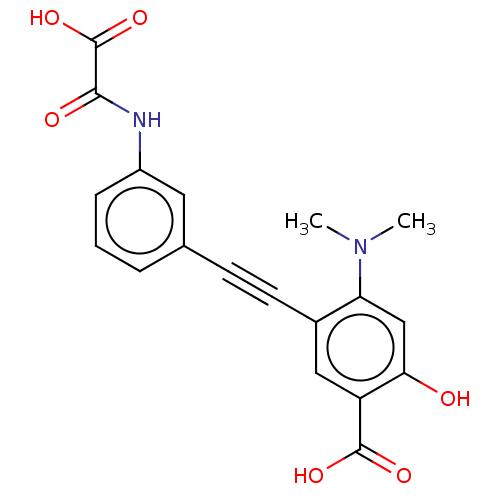
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
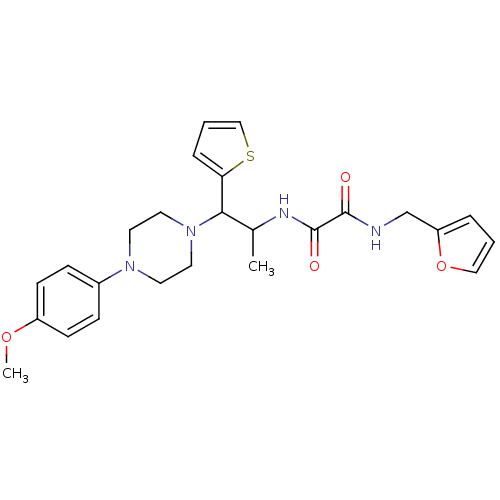
Affinity DataIC50: 410nMAssay Description:Inhibition of recombinant human N-terminal GST-tagged MEG2 catalytic domain (285 to 593 residues) expressed in Escherichia coli using p-nitrophenol a...More data for this Ligand-Target Pair
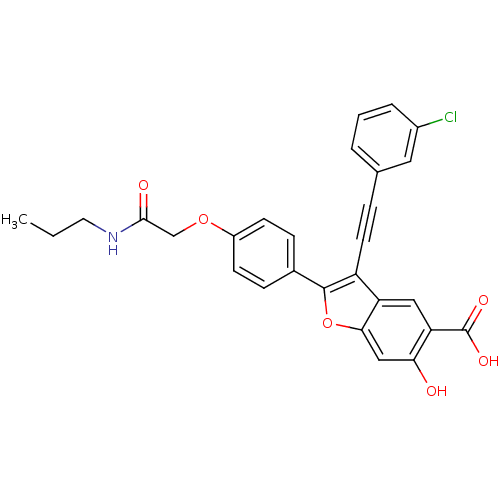
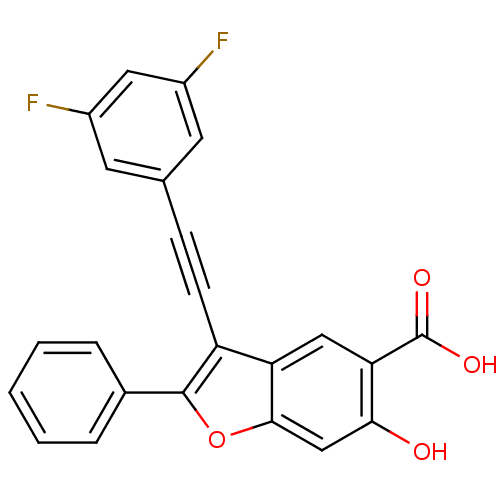
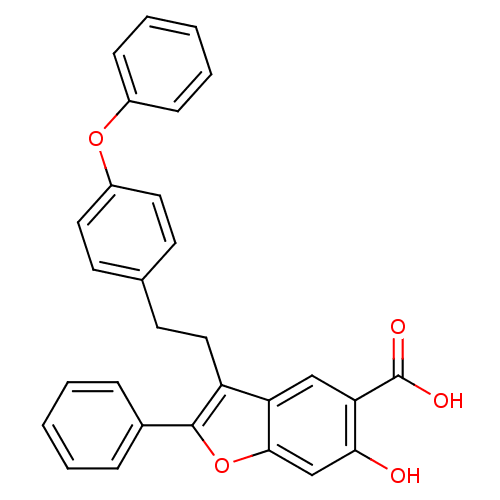
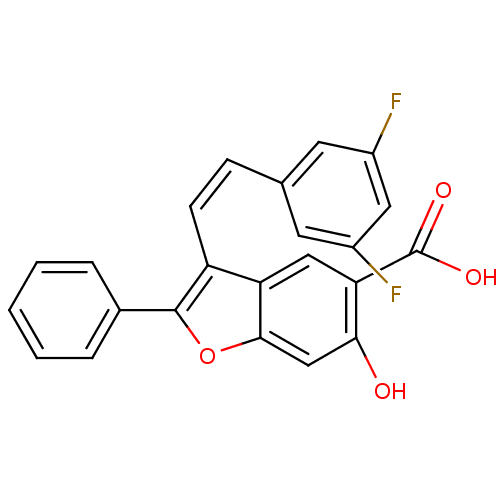
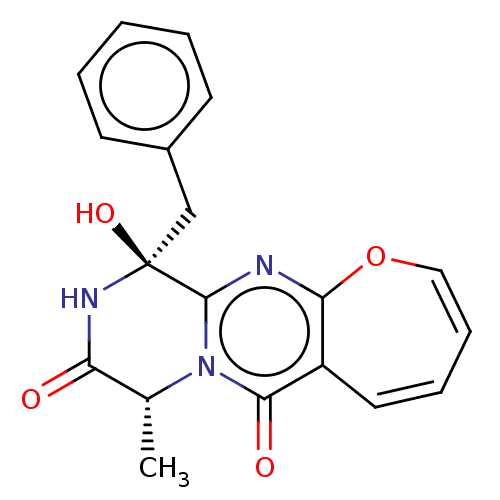
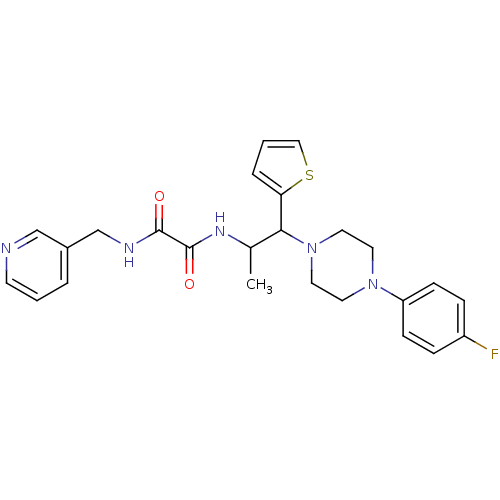
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
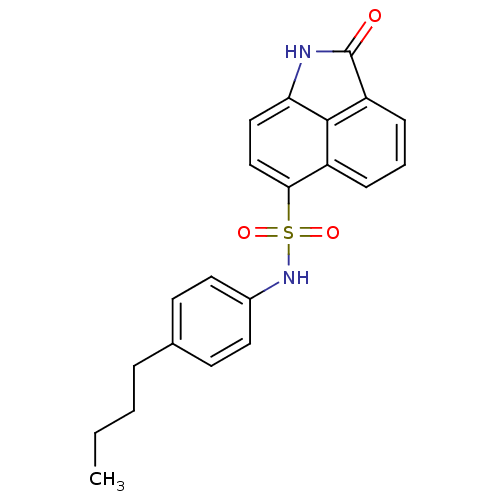
Affinity DataIC50: 590nMAssay Description:Inhibition of recombinant PTP-MEG2 (unknown origin) using pNPP as substrate by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9 [277-582](Human)
Indiana University Research and Technology
US Patent
Indiana University Research and Technology
US Patent
Affinity DataIC50: 900nMpH: 7.0Assay Description:PTP activity was assayed using p-nitrophenyl phosphate (pNPP) as a substrate in 3,3-dimethylglutarate buffer (50 mM 3,3-dimethylglutarate, pH 7.0, 1 ...More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataKi: >1.00E+3nM ΔG°: >-34.2kJ/molepH: 7.0 T: 2°CAssay Description:PTP activity was assayed using p-nitrophenyl phosphate (pNPP) as a substrate in DMG buffer (50 mM DMG, pH 7.0, 1 mM EDTA, 150 mM NaCl, 2 mM DTT, 0.1 ...More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 1.07E+3nMAssay Description:Inhibition of PTPMEG2 (unknown origin) expressed in Escherichia coli using pNPP substrate after 5 mins by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 2.30E+3nMAssay Description:Inhibition of PTPMEG2 (unknown origin) expressed in Escherichia coli using pNPP substrate after 5 mins by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 2.40E+3nMAssay Description:Inhibition of recombinant human N-terminal GST-tagged MEG2 catalytic domain (285 to 593 residues) expressed in Escherichia coli using p-nitrophenol a...More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of recombinant PTP-MEG2 (unknown origin) using pNPP as substrate by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 4.00E+3nMAssay Description:Inhibition of PTPMEG2 (unknown origin) expressed in Escherichia coli using pNPP substrate after 5 mins by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 4.00E+3nMAssay Description:Inhibition of PTPMEG2 (unknown origin) expressed in Escherichia coli using pNPP substrate after 5 mins by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 4.30E+3nMAssay Description:Inhibition of PTPMEG2 (unknown origin) expressed in Escherichia coli using pNPP substrate after 5 mins by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 4.80E+3nMAssay Description:Inhibition of PTPMEG2 (unknown origin) expressed in Escherichia coli using pNPP substrate after 5 mins by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 5.90E+3nMAssay Description:Inhibition of recombinant human N-terminal GST-tagged MEG2 catalytic domain (285 to 593 residues) expressed in Escherichia coli using p-nitrophenol a...More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 7.70E+3nMAssay Description:Inhibition of PTPMEG2 (unknown origin) expressed in Escherichia coli using pNPP substrate after 5 mins by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 8.40E+3nMAssay Description:Inhibition of PTPMEG2 (unknown origin) expressed in Escherichia coli using pNPP substrate after 5 mins by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 8.60E+3nMAssay Description:Inhibition of PTPMEG2 (unknown origin) expressed in Escherichia coli using pNPP substrate after 5 mins by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 9.00E+3nMAssay Description:Inhibition of PTPMEG2 (unknown origin) expressed in Escherichia coli using pNPP substrate after 5 mins by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 9.20E+3nMAssay Description:Inhibition of PTPMEG2 (unknown origin) expressed in Escherichia coli using pNPP substrate after 5 mins by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 1.07E+4nMAssay Description:Inhibition of MEG2 (unknown origin) assessed as reduction in pNPP hydrolysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 1.14E+4nMAssay Description:Inhibition of PTPMEG2 (unknown origin) expressed in Escherichia coli using pNPP substrate after 5 mins by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 1.40E+4nMAssay Description:Inhibition of recombinant human MEG2 expressed in Escherichia coli using pNPP as substrate preincubated for 15 mins followed by substrate addition an...More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 1.40E+4nMAssay Description:Inhibition of recombinant human MEG2 expressed in Escherichia coli using pNPP as substrate preincubated for 15 mins followed by substrate addition an...More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 1.40E+4nMAssay Description:Inhibition of recombinant human MEG2 expressed in Escherichia coli using pNPP as substrate preincubated for 15 mins followed by substrate addition an...More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 1.51E+4nMAssay Description:Inhibition of PTPMEG2 (unknown origin) expressed in Escherichia coli using pNPP substrate after 5 mins by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of PTPMEG2 (unknown origin) expressed in Escherichia coli using pNPP substrate after 5 mins by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of PTP-MEG2 (unknown origin) expressed in Escherichia coli BL21 using p-nitrophenyl phosphate as substrate measured after 30 mins by UV-vi...More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of PTP-MEG2 (unknown origin) expressed in Escherichia coli BL21 using p-nitrophenyl phosphate as substrate measured after 30 mins by UV-vi...More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 3.18E+4nMAssay Description:Inhibition of PTP-Meg2More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 3.75E+4nMAssay Description:Inhibition of MEG2 (unknown origin) assessed as reduction in pNPP hydrolysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of human MEG2 expressed in Escherichia coli after 5 mins by microplate spectrophotometer analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of human MEG2 expressed in Escherichia coli after 5 mins by microplate spectrophotometer analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of phosphatase activity of human Meg2 using pNPP as a substrate after 10 mins by spectrophotometer analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of phosphatase activity of human Meg2 using pNPP as a substrate after 10 mins by spectrophotometer analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of PTP-MEG2 (unknown origin) expressed in Escherichia coli BL21 using p-nitrophenyl phosphate as substrate measured after 30 mins by UV-vi...More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of human MEG2 expressed in Escherichia coli after 5 mins by microplate spectrophotometer analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of phosphatase activity of human Meg2 using pNPP as a substrate after 10 mins by spectrophotometer analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 5.94E+4nMAssay Description:Inhibition of GST-tagged MEG2 (unknown origin) using phospho-EGFR Asp-Ala-Asp-Glu-Tyr[PO3H2]-Leu-Ile-Pro-Gln-Gln-Gly as substrate preincubated for 30...More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 6.61E+4nMAssay Description:Inhibition of MEG2 (unknown origin) assessed as reduction in pNPP hydrolysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 6.86E+4nMAssay Description:Inhibition of human PTP-MEG2 using p-nitrophenol phosphate as substrate incubated for 35 mins prior to substrate addition measured after 30 mins by s...More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 6.90E+4nMAssay Description:Inhibition of human PTP-Meg2 using pNPP substrate by spectrophotometryMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 7.85E+4nMAssay Description:Inhibition of MEG2 (unknown origin) assessed as reduction in pNPP hydrolysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 8.50E+4nMAssay Description:Inhibition of PTPMEG2 (unknown origin) expressed in Escherichia coli using pNPP substrate after 5 mins by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of PTPMEG2 (unknown origin) expressed in Escherichia coli using pNPP substrate after 5 mins by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of PTPMEG2 (unknown origin) expressed in Escherichia coli using pNPP substrate after 5 mins by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of His-tagged PTP-MEG2 (unknown origin) expressed in Escherichia coli BL21 cells using para-nitrophenyl phosphate as substrate incubated f...More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of His-tagged PTP-MEG2 (unknown origin) expressed in Escherichia coli BL21 cells using para-nitrophenyl phosphate as substrate incubated f...More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 1.10E+5nMAssay Description:Inhibition of PTPMEG2 (unknown origin) expressed in Escherichia coli using pNPP substrate after 5 mins by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 9(Human)
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
V.P. Kukhar Institute of Bioorganic Chemistry and Petrochemistry of The National Academy of Sciences of Ukraine
Curated by ChEMBL
Affinity DataIC50: 1.46E+5nMAssay Description:Inhibition of human PTP-MEG2 using p-nitrophenol phosphate as substrate incubated for 35 mins prior to substrate addition measured after 30 mins by s...More data for this Ligand-Target Pair