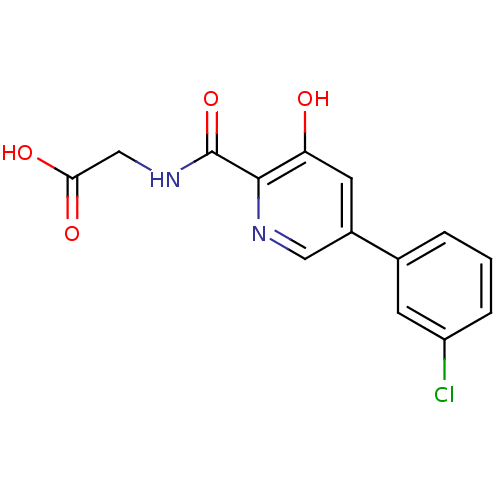
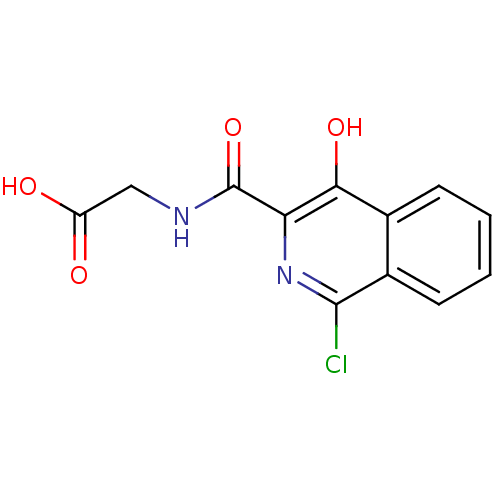
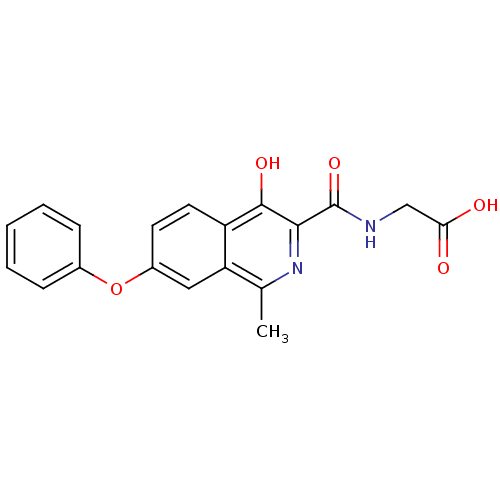
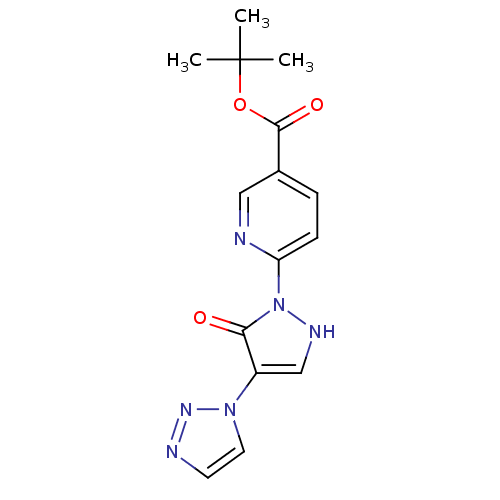
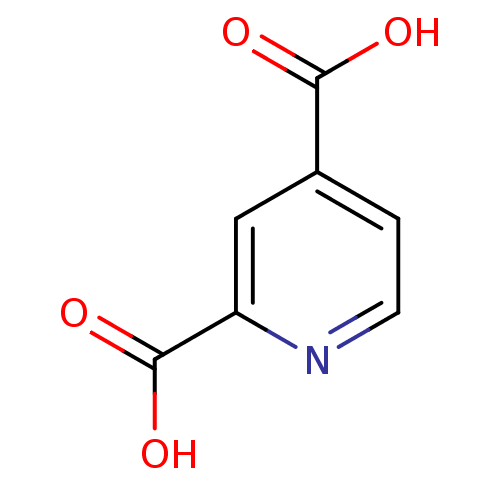
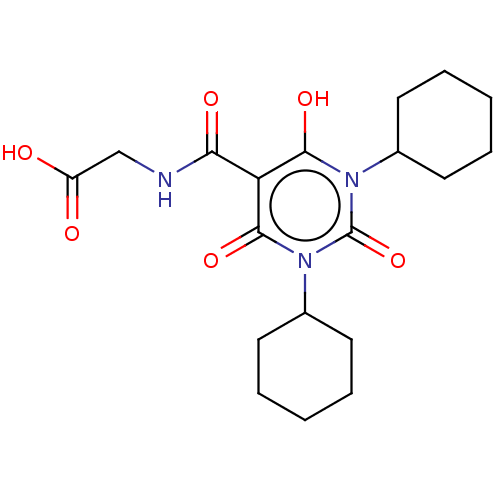
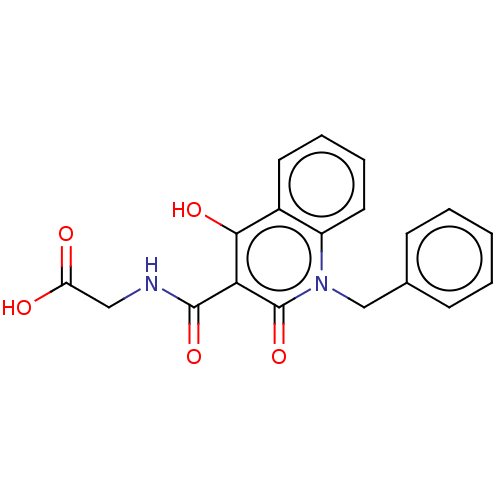
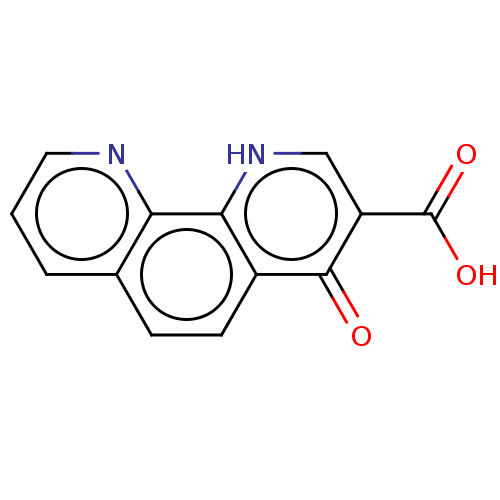
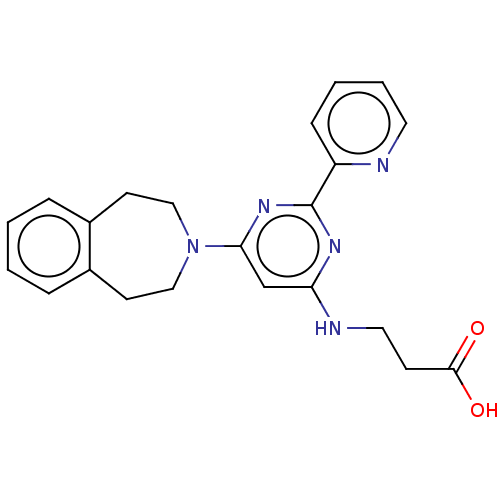
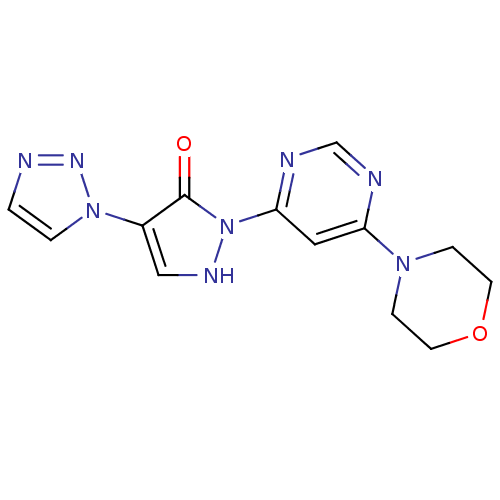
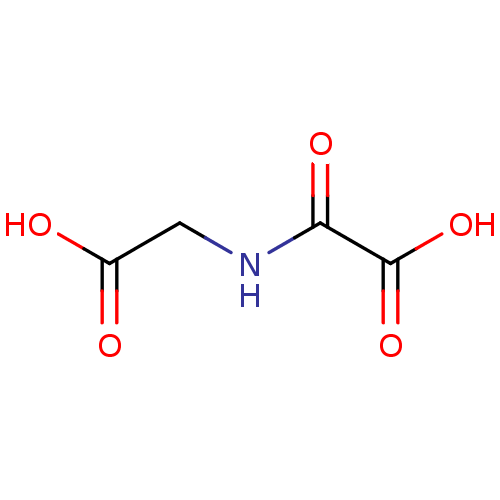
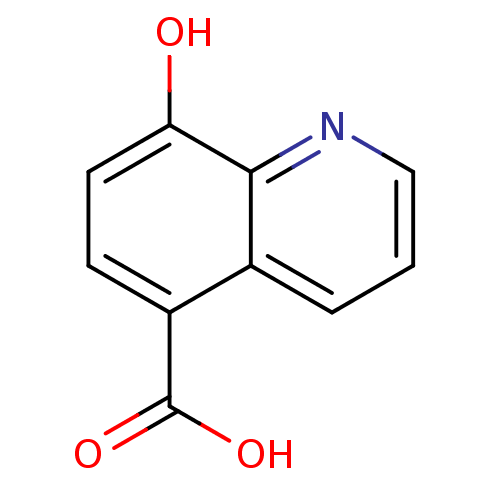
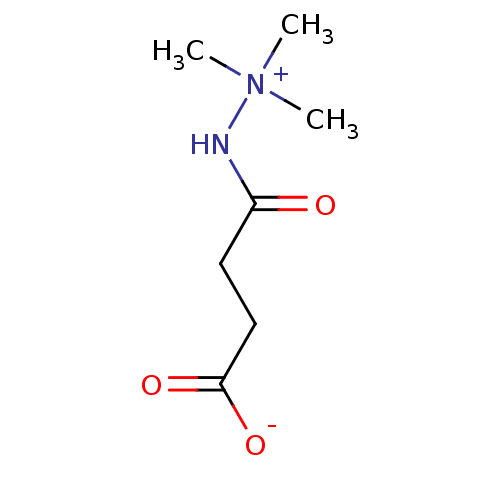
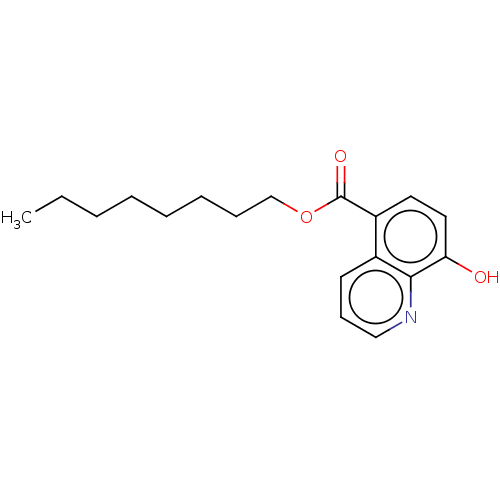
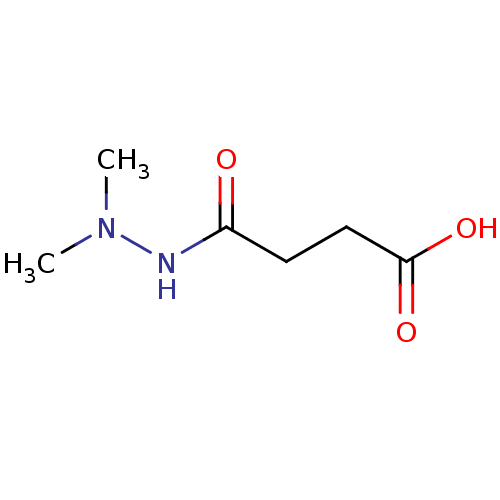
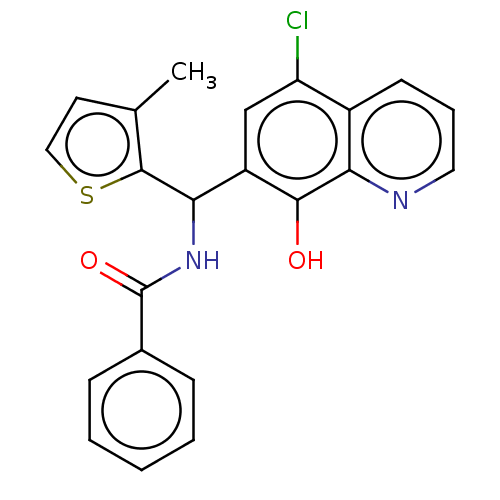
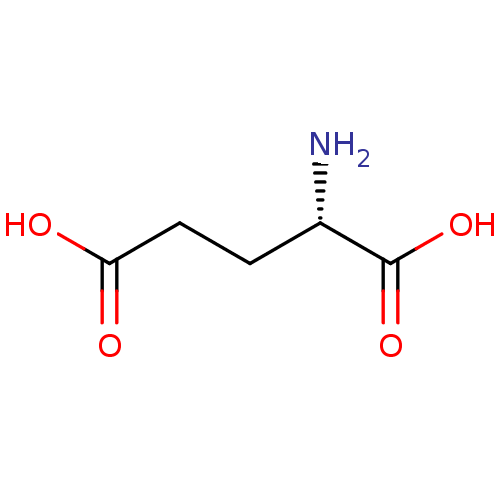
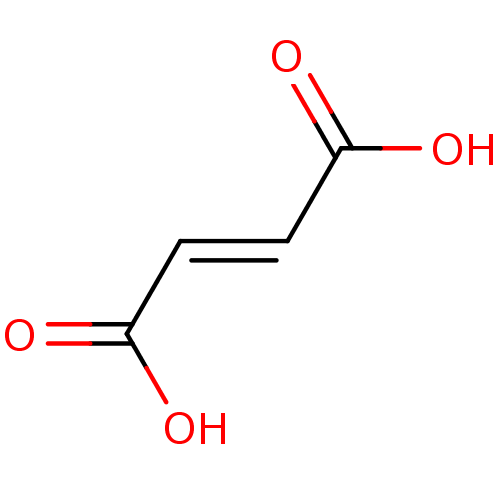
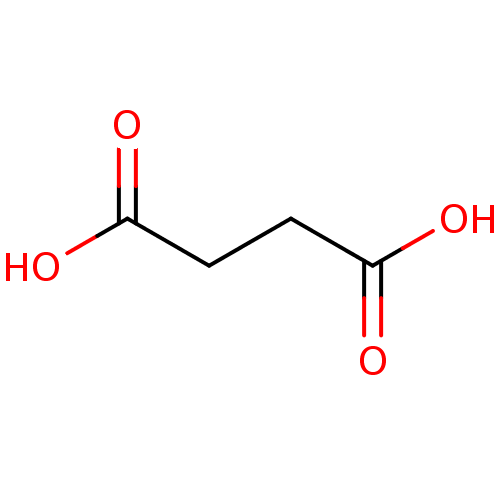
Affinity DataKi: 280nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataKi: 710nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataKi: 1.07E+3nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+3nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataKi: 3.28E+3nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 4.50E+3nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 5.30E+3nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataKi: 7.65E+3nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 8.50E+3nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.34E+4nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.57E+4nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.59E+4nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataKi: 1.61E+4nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 2.12E+4nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 2.68E+4nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 4.72E+4nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataKi: 4.87E+4nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 2.13E+5nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 2.84E+5nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of N-terminal His6-tagged recombinant Paramecium bursaria chlorella virus 1 CPH expressed in Escherichia coli Rosetta 2 (DE3) cells pre-in...More data for this Ligand-Target Pair