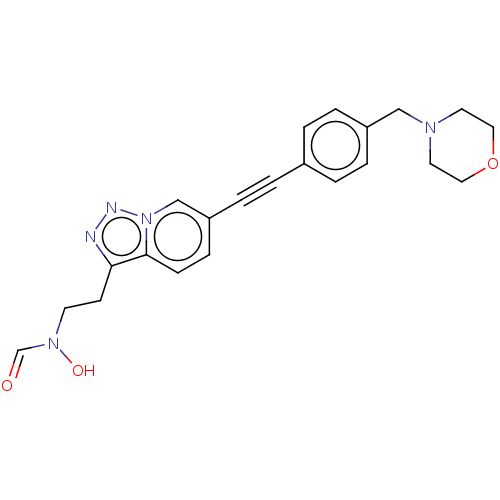
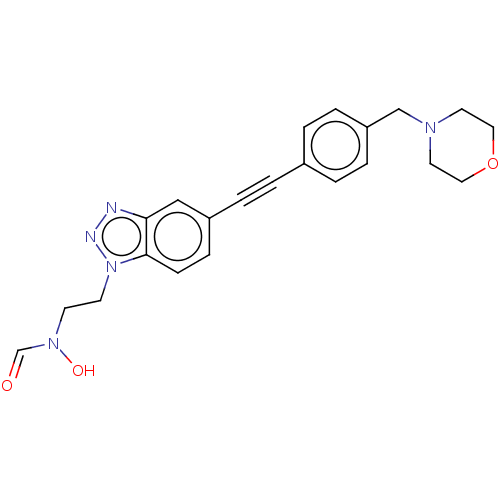
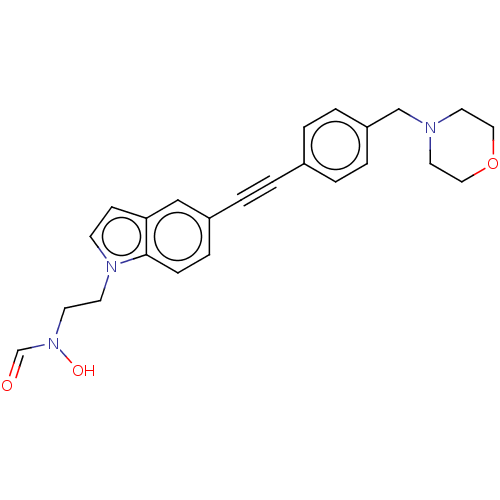
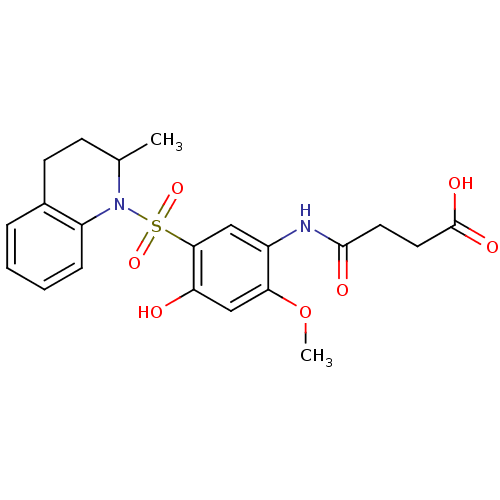
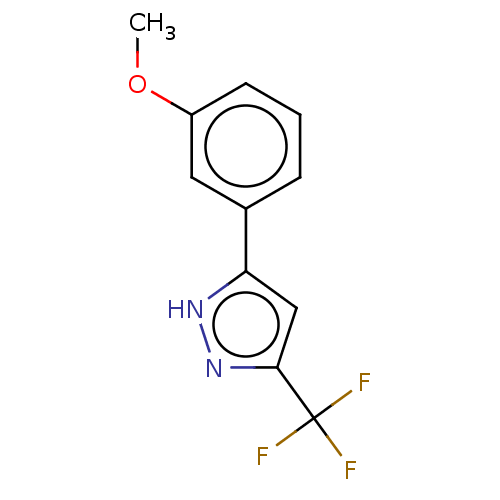
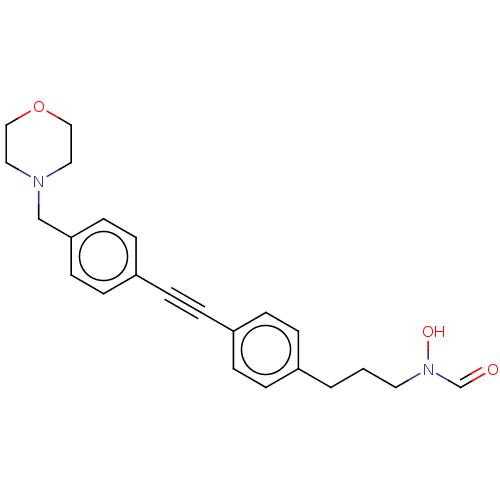
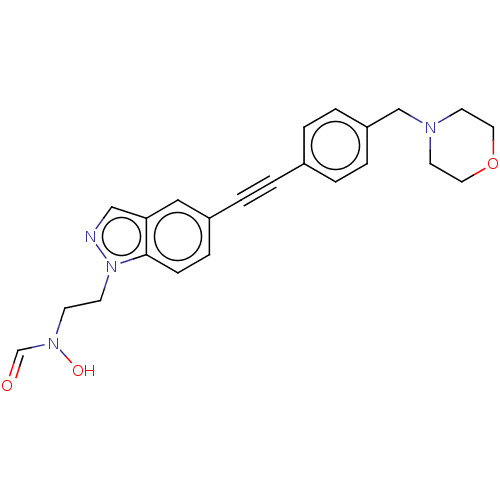
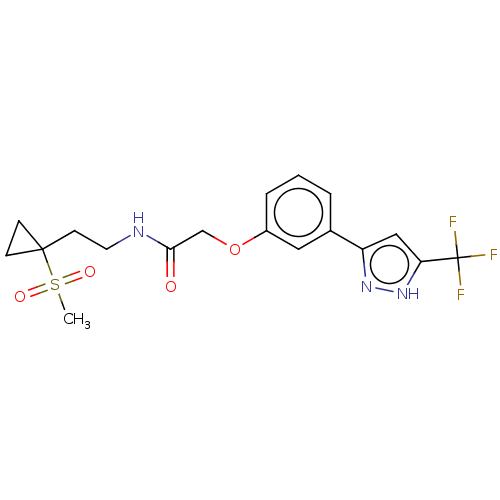
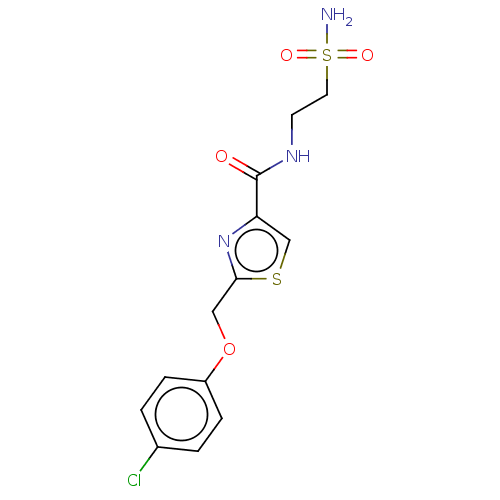
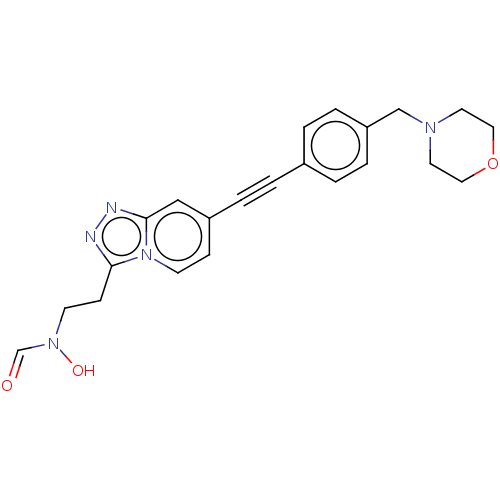
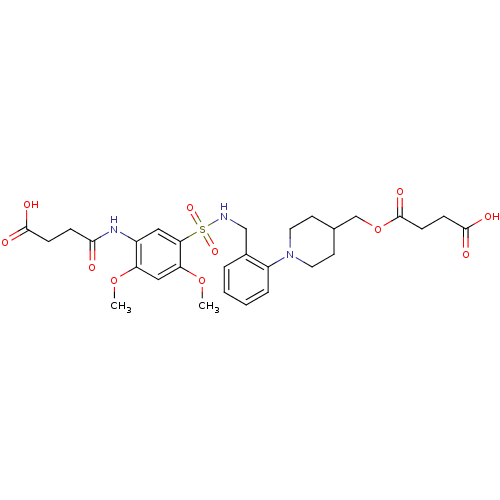
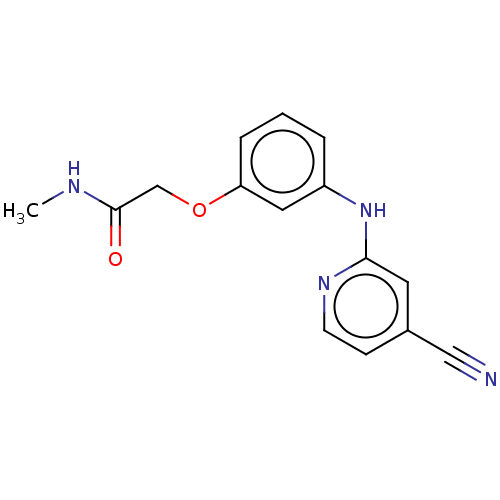
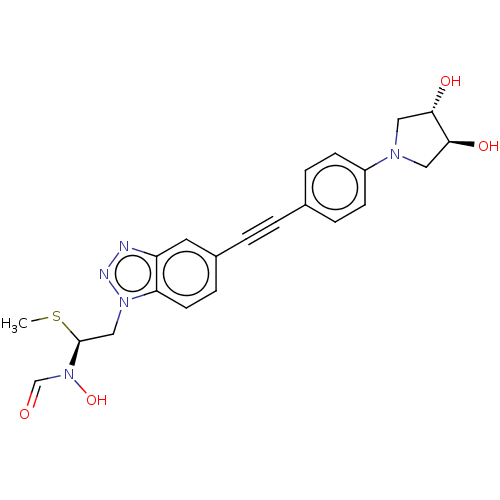
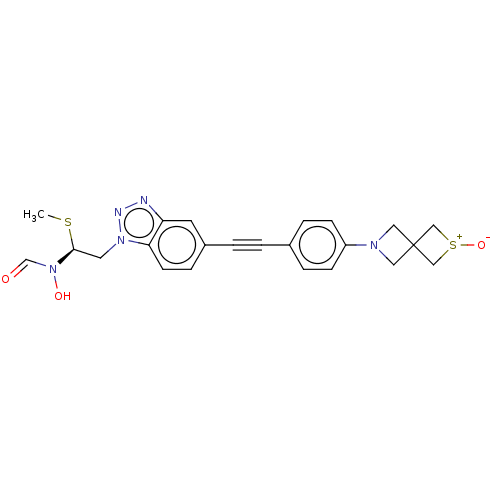
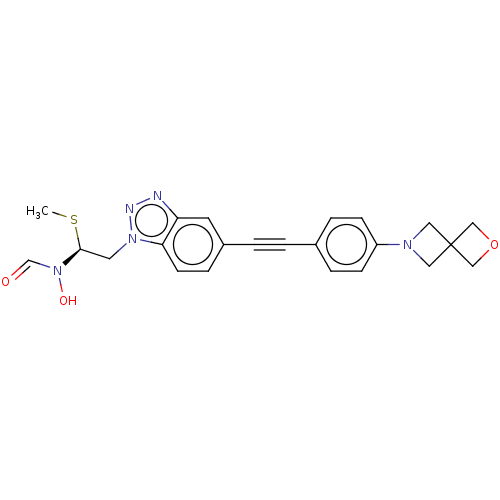
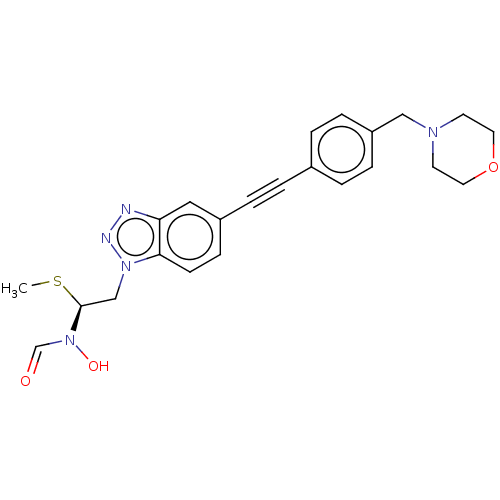
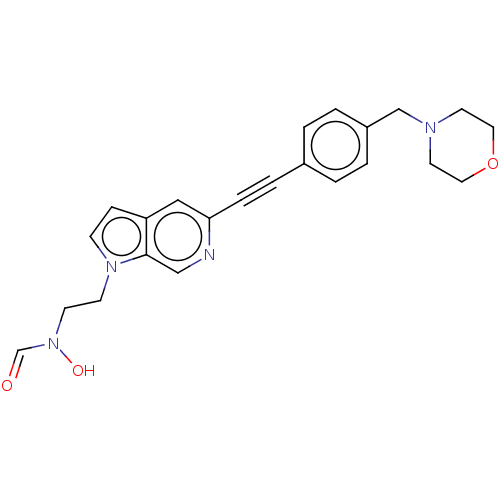
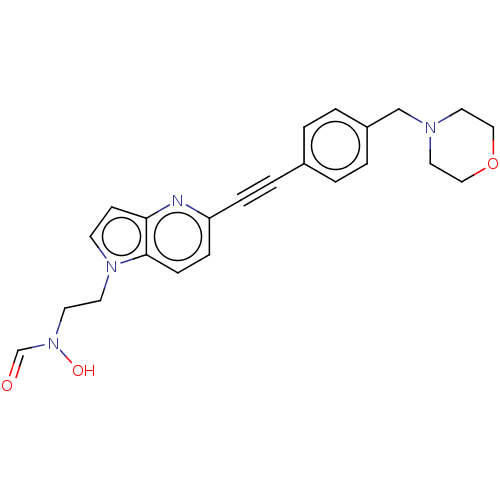
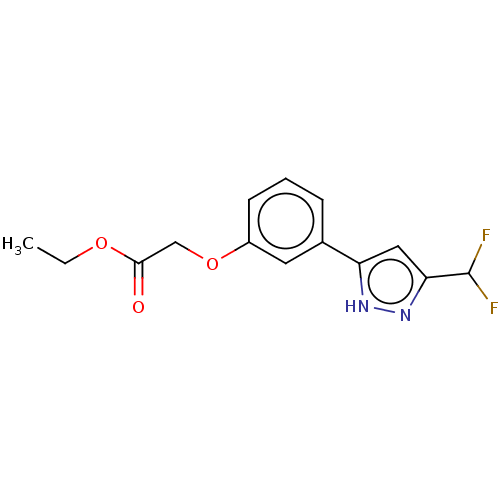
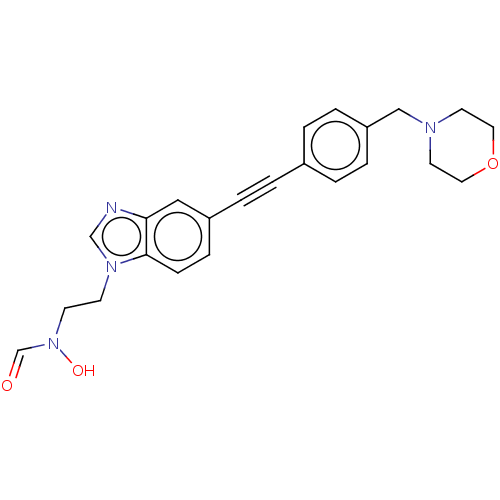
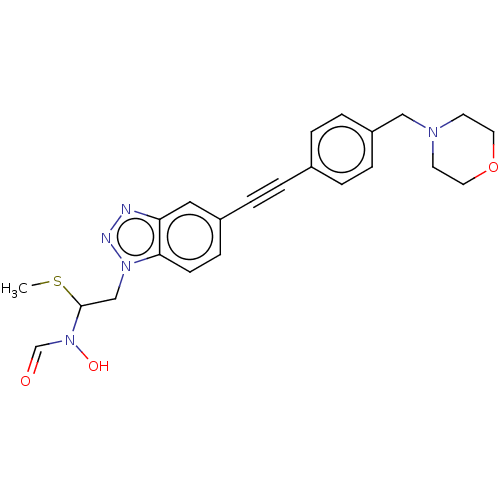
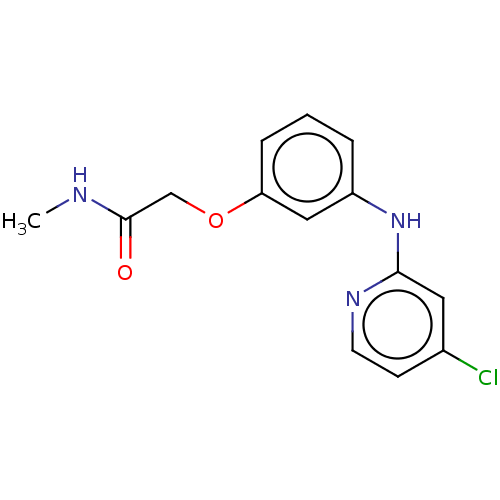
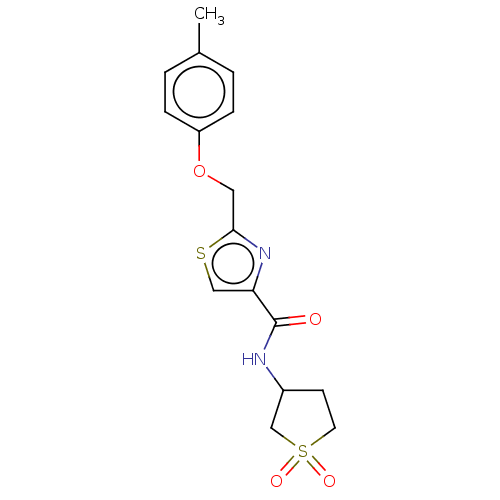
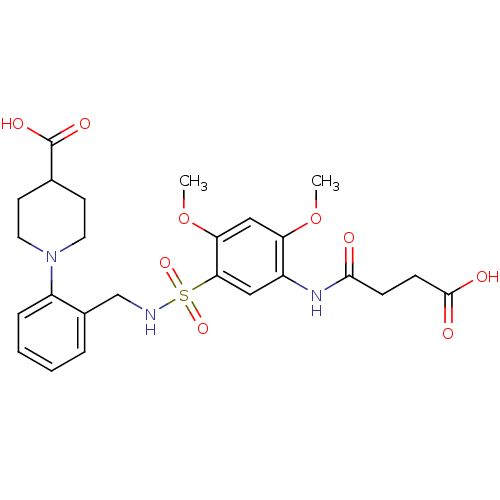
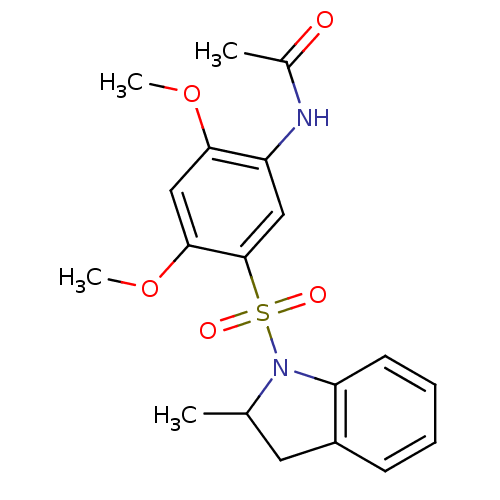
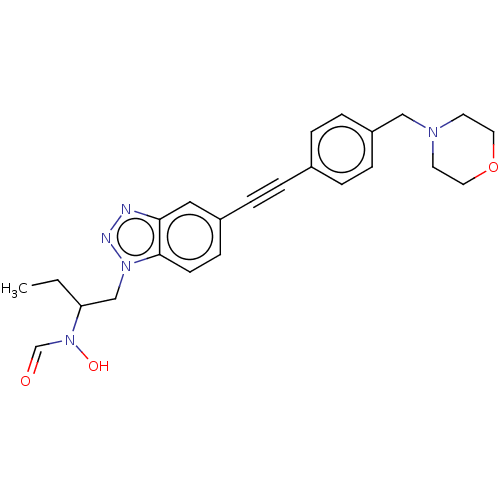
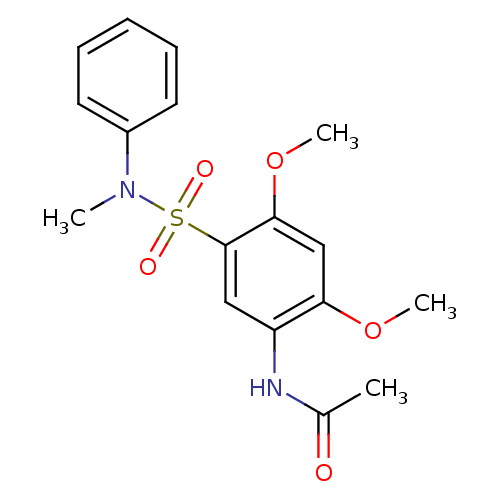
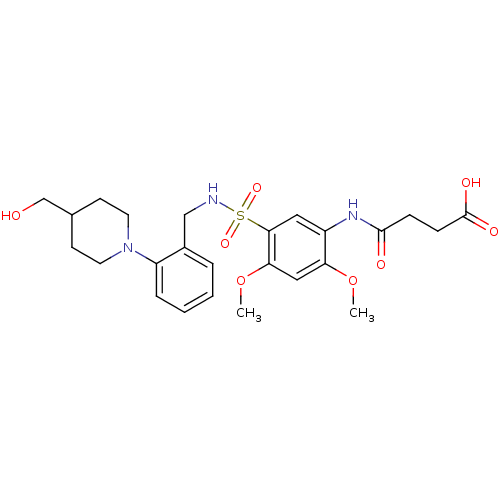
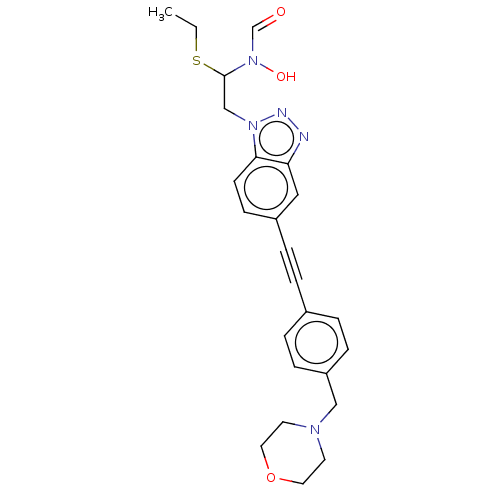
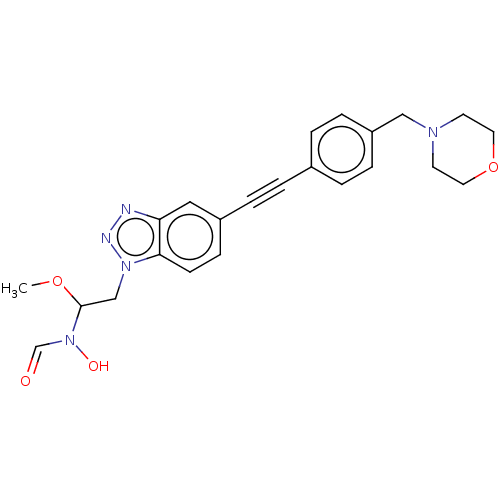
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: <1nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetPhenylalanine--tRNA ligase alpha/beta subunit(Escherichia coli (Enterobacteria))
Astrazeneca R&D Boston
Astrazeneca R&D Boston
Affinity DataIC50: 2.20nMpH: 8.0Assay Description:Compounds were solubilized in DMSO. Serial 2-fold dilutions covering two concentration ranges, 10 mM to 19.5 μM and 100 μM to 195 nM, were ...More data for this Ligand-Target Pair
TargetPhenylalanine--tRNA ligase alpha/beta subunit(Escherichia coli (Enterobacteria))
Astrazeneca R&D Boston
Astrazeneca R&D Boston
Affinity DataIC50: 2.70nMpH: 8.0Assay Description:Compounds were solubilized in DMSO. Serial 2-fold dilutions covering two concentration ranges, 10 mM to 19.5 μM and 100 μM to 195 nM, were ...More data for this Ligand-Target Pair
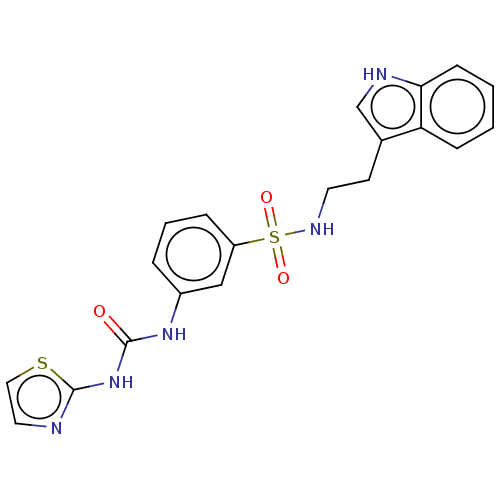
Affinity DataIC50: 7nMpH: 7.35Assay Description:Inhibition assay using GlmU bifuncation enzyme with various bacterial strains.More data for this Ligand-Target Pair
Affinity DataIC50: 13nMpH: 8.0Assay Description:Compounds were solubilized in DMSO. Serial 2-fold dilutions covering two concentration ranges, 10 mM to 19.5 μM and 100 μM to 195 nM, were ...More data for this Ligand-Target Pair
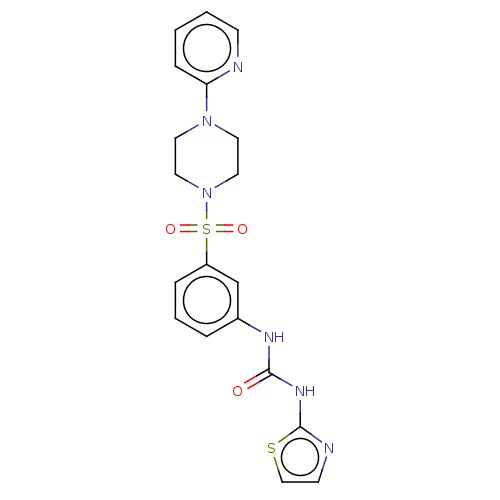
Affinity DataIC50: 18nMpH: 7.35Assay Description:Inhibition assay using GlmU bifuncation enzyme with various bacterial strains.More data for this Ligand-Target Pair
Affinity DataIC50: 22nMpH: 8.0Assay Description:Compounds were solubilized in DMSO. Serial 2-fold dilutions covering two concentration ranges, 10 mM to 19.5 μM and 100 μM to 195 nM, were ...More data for this Ligand-Target Pair
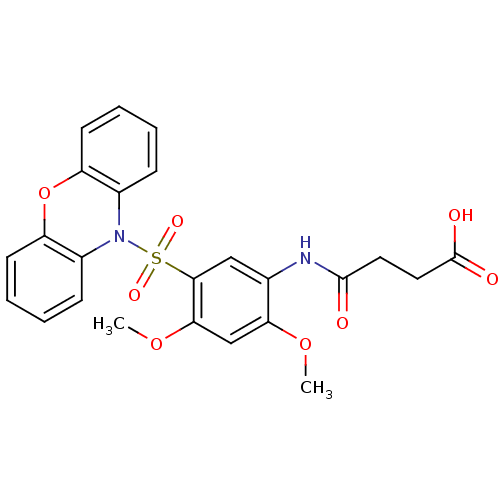
Affinity DataIC50: 23nMpH: 7.35Assay Description:Inhibition assay using GlmU bifuncation enzyme with various bacterial strains.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 37nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 44nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 45nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
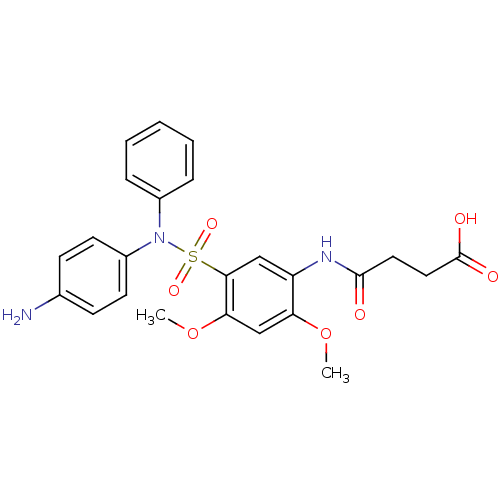
Affinity DataIC50: 58nMpH: 7.35Assay Description:Inhibition assay using GlmU bifuncation enzyme with various bacterial strains.More data for this Ligand-Target Pair
TargetPhenylalanine--tRNA ligase alpha/beta subunit(Escherichia coli (Enterobacteria))
Astrazeneca R&D Boston
Astrazeneca R&D Boston
Affinity DataIC50: 60nMpH: 8.0Assay Description:Compounds were solubilized in DMSO. Serial 2-fold dilutions covering two concentration ranges, 10 mM to 19.5 μM and 100 μM to 195 nM, were ...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 63nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 64nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetPhenylalanine--tRNA ligase alpha/beta subunit(Escherichia coli (Enterobacteria))
Astrazeneca R&D Boston
Astrazeneca R&D Boston
Affinity DataIC50: 70nMpH: 8.0Assay Description:Compounds were solubilized in DMSO. Serial 2-fold dilutions covering two concentration ranges, 10 mM to 19.5 μM and 100 μM to 195 nM, were ...More data for this Ligand-Target Pair
TargetPhenylalanine--tRNA ligase alpha/beta subunit(Escherichia coli (Enterobacteria))
Astrazeneca R&D Boston
Astrazeneca R&D Boston
Affinity DataIC50: 80nMpH: 8.0Assay Description:Compounds were solubilized in DMSO. Serial 2-fold dilutions covering two concentration ranges, 10 mM to 19.5 μM and 100 μM to 195 nM, were ...More data for this Ligand-Target Pair
TargetPhenylalanine--tRNA ligase alpha/beta subunit(Escherichia coli (Enterobacteria))
Astrazeneca R&D Boston
Astrazeneca R&D Boston
Affinity DataIC50: 100nMpH: 8.0Assay Description:Compounds were solubilized in DMSO. Serial 2-fold dilutions covering two concentration ranges, 10 mM to 19.5 μM and 100 μM to 195 nM, were ...More data for this Ligand-Target Pair
TargetPhenylalanine--tRNA ligase alpha/beta subunit(Escherichia coli (Enterobacteria))
Astrazeneca R&D Boston
Astrazeneca R&D Boston
Affinity DataIC50: 100nMpH: 8.0Assay Description:Compounds were solubilized in DMSO. Serial 2-fold dilutions covering two concentration ranges, 10 mM to 19.5 μM and 100 μM to 195 nM, were ...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 170nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetPhenylalanine--tRNA ligase alpha/beta subunit(Escherichia coli (Enterobacteria))
Astrazeneca R&D Boston
Astrazeneca R&D Boston
Affinity DataIC50: 190nMpH: 8.0Assay Description:Compounds were solubilized in DMSO. Serial 2-fold dilutions covering two concentration ranges, 10 mM to 19.5 μM and 100 μM to 195 nM, were ...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 280nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 280nMpH: 7.35Assay Description:Inhibition assay using GlmU bifuncation enzyme with various bacterial strains.More data for this Ligand-Target Pair
Affinity DataIC50: 400nMpH: 7.35Assay Description:Inhibition assay using GlmU bifuncation enzyme with various bacterial strains.More data for this Ligand-Target Pair
Affinity DataIC50: 500nMAssay Description:Inhibition of Streptococcus pneumoniae acetyltransferase activity of GlmU using acetyl-CoA and glucosamine-1-phosphate after 30 mins by Ellman's meth...More data for this Ligand-Target Pair
TargetPhenylalanine--tRNA ligase alpha/beta subunit(Escherichia coli (Enterobacteria))
Astrazeneca R&D Boston
Astrazeneca R&D Boston
Affinity DataIC50: 560nMpH: 8.0Assay Description:Compounds were solubilized in DMSO. Serial 2-fold dilutions covering two concentration ranges, 10 mM to 19.5 μM and 100 μM to 195 nM, were ...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: <1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: <1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: <1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: <1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: <1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetPhenylalanine--tRNA ligase alpha/beta subunit(Escherichia coli (Enterobacteria))
Astrazeneca R&D Boston
Astrazeneca R&D Boston
Affinity DataIC50: 1.00E+3nMpH: 8.0Assay Description:Compounds were solubilized in DMSO. Serial 2-fold dilutions covering two concentration ranges, 10 mM to 19.5 μM and 100 μM to 195 nM, were ...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.10E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.30E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetPhenylalanine--tRNA ligase alpha/beta subunit(Escherichia coli (Enterobacteria))
Astrazeneca R&D Boston
Astrazeneca R&D Boston
Affinity DataIC50: 1.40E+3nMpH: 8.0Assay Description:Compounds were solubilized in DMSO. Serial 2-fold dilutions covering two concentration ranges, 10 mM to 19.5 μM and 100 μM to 195 nM, were ...More data for this Ligand-Target Pair
TargetPhenylalanine--tRNA ligase alpha/beta subunit(Escherichia coli (Enterobacteria))
Astrazeneca R&D Boston
Astrazeneca R&D Boston
Affinity DataIC50: 1.40E+3nMpH: 8.0Assay Description:Compounds were solubilized in DMSO. Serial 2-fold dilutions covering two concentration ranges, 10 mM to 19.5 μM and 100 μM to 195 nM, were ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of Streptococcus pneumoniae acetyltransferase activity of GlmU using acetyl-CoA and glucosamine-1-phosphate after 30 mins by Ellman's meth...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of Streptococcus pneumoniae acetyltransferase activity of GlmU using acetyl-CoA and glucosamine-1-phosphate after 30 mins by Ellman's meth...More data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+3nMpH: 7.35Assay Description:Inhibition assay using GlmU bifuncation enzyme with various bacterial strains.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 2.40E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+3nM Kd: 1.90E+3nMpH: 7.35Assay Description:Inhibition assay using GlmU bifuncation enzyme with various bacterial strains.More data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+3nMAssay Description:Inhibition of human AchE expressed in HEK293 cells using acetylthiocholine as substrate by Ellman's methodMore data for this Ligand-Target Pair
Affinity DataIC50: 4.20E+3nMpH: 8.0Assay Description:Compounds were solubilized in DMSO. Serial 2-fold dilutions covering two concentration ranges, 10 mM to 19.5 μM and 100 μM to 195 nM, were ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.70E+3nMpH: 7.35Assay Description:Inhibition assay using GlmU bifuncation enzyme with various bacterial strains.More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of Streptococcus pneumoniae acetyltransferase activity of GlmU using acetyl-CoA and glucosamine-1-phosphate after 30 mins by Ellman's meth...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 5.40E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.40E+3nMpH: 7.35Assay Description:Inhibition assay using GlmU bifuncation enzyme with various bacterial strains.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 7.30E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair

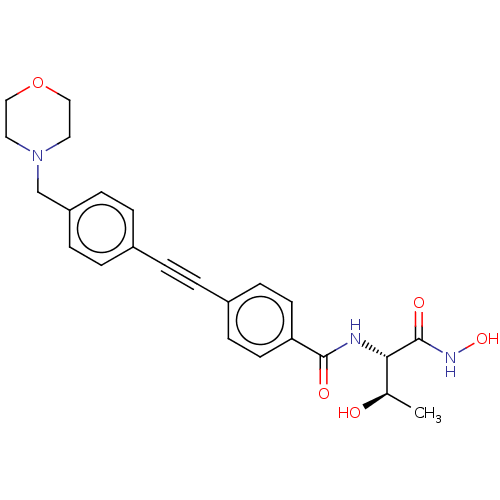
 3D Structure (crystal)
3D Structure (crystal)