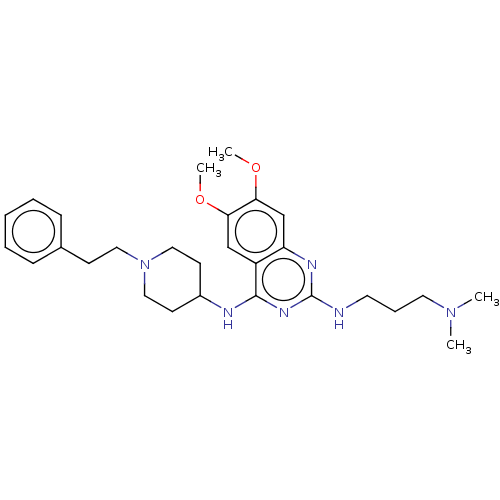
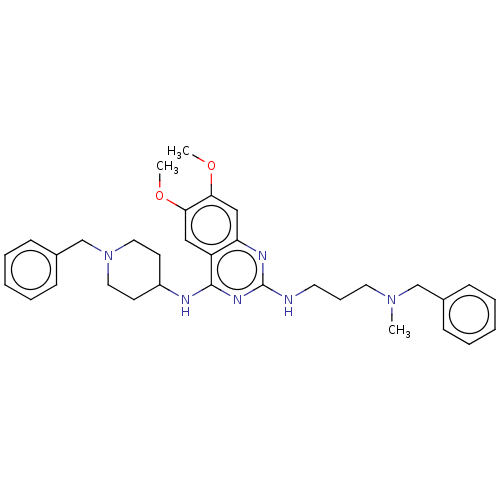
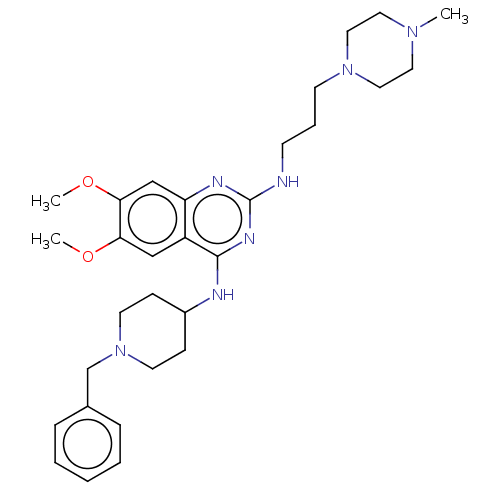
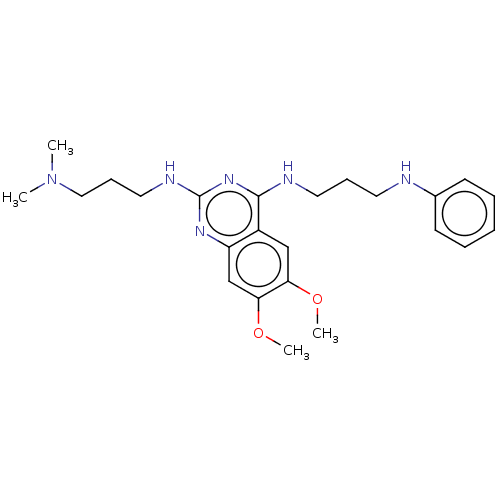
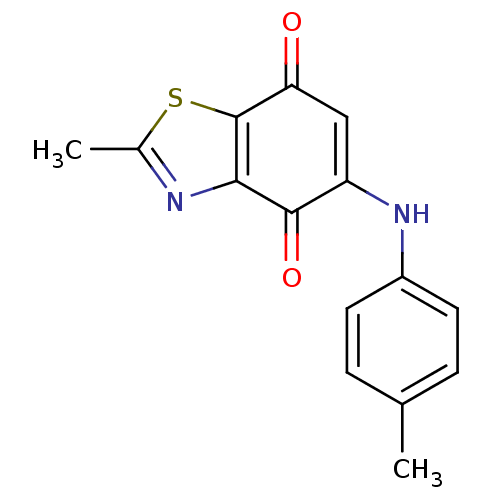
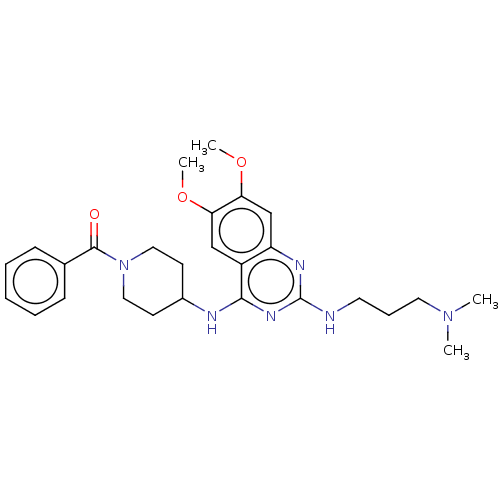
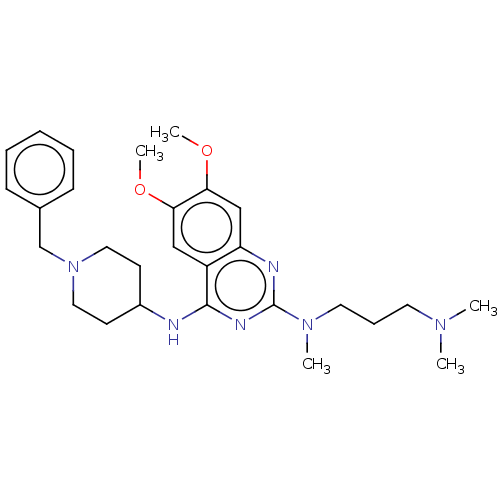
Report error Found 93 Enz. Inhib. hit(s) with all data for entry = 50016305
Affinity DataKd: 14nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
Affinity DataKi: 35nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 43nMAssay Description:Inhibition of human MAO-A using p-tyramine as substrate by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 48nMAssay Description:Inhibition of human MAO-B using p-tyramine as substrate by fluorimetric analysisMore data for this Ligand-Target Pair
Affinity DataKd: 64nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 67nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 68nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 75nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 81nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 82nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 82nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 82nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 83nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 84nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 86nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 108nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 112nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 115nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 127nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 127nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase, H3 lysine-79 specific(Human)
Sapienza University of Rome
Curated by ChEMBL
Sapienza University of Rome
Curated by ChEMBL
Affinity DataIC50: 140nMAssay Description:Inhibition of DOTL1 (unknown origin) using [3H]SAM and HeLa oligo nucleosomes as substrates incubated for 1 hrMore data for this Ligand-Target Pair
Affinity DataKi: 146nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 147nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 149nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 153nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 156nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 162nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 170nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 175nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 183nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 191nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 205nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 212nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 228nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 230nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 237nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 243nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 250nMAssay Description:Inhibition of PRMT1 (unknown origin) using [3H]SAM and chicken histone 4 as substrates incubated for 1 hrMore data for this Ligand-Target Pair
Affinity DataKd: 251nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 356nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 369nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 370nMAssay Description:Inhibition of SETD8 (unknown origin) using [3H]SAM and HeLa nucleosomes as substrates incubated for 1 hrMore data for this Ligand-Target Pair
Affinity DataKi: 440nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 569nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 680nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
Affinity DataKi: 690nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
Affinity DataKd: 975nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.18E+3nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
Affinity DataKd: 1.30E+3nMAssay Description:Binding affinity to human LSD1/CoREST assessed as dissociation constant by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
 3D Structure (crystal)
3D Structure (crystal)