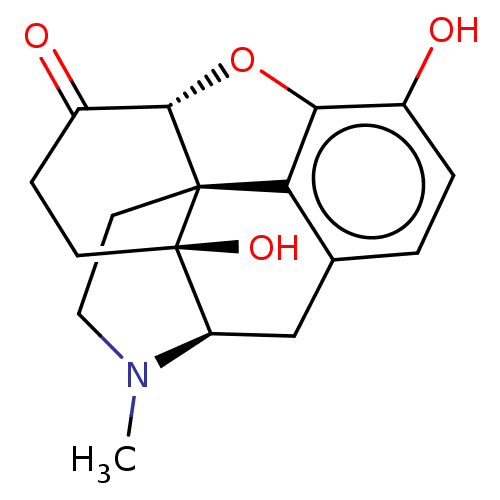
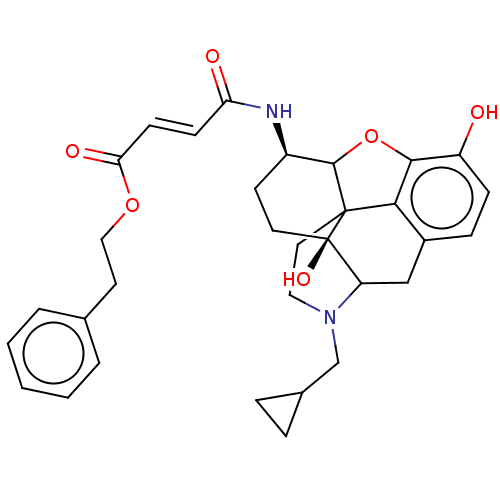
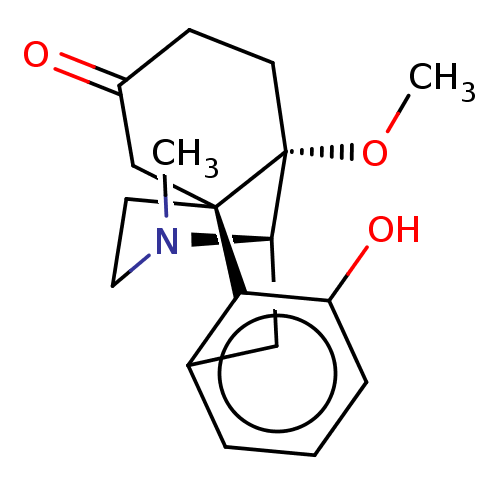
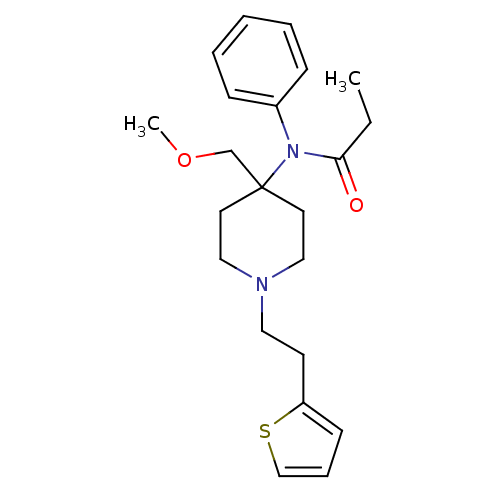
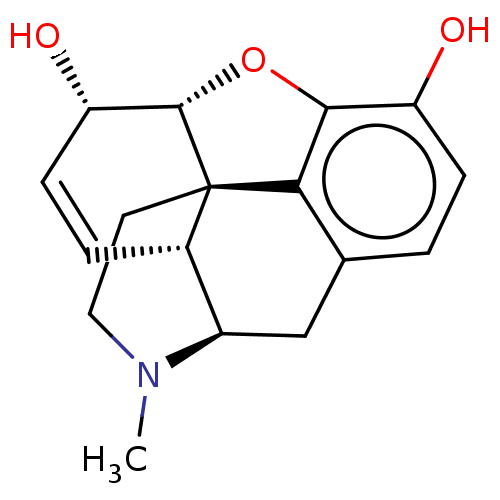
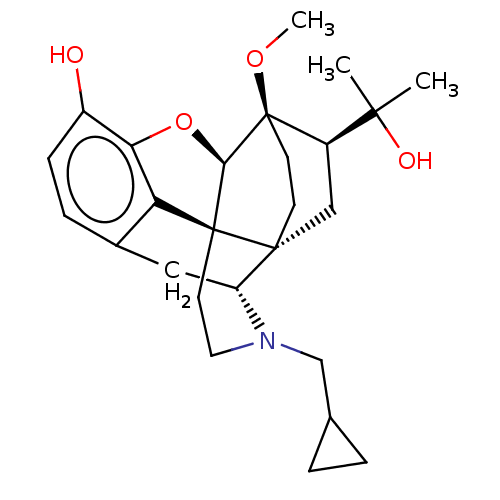
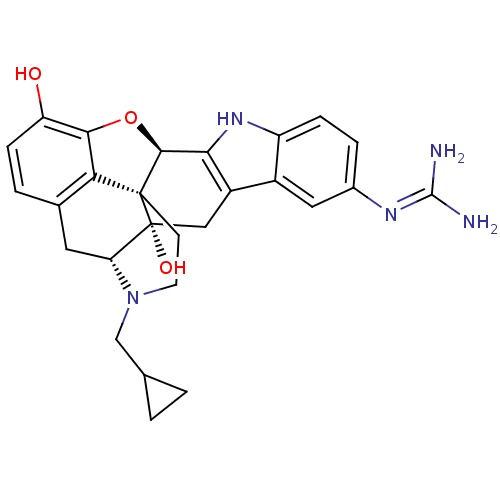
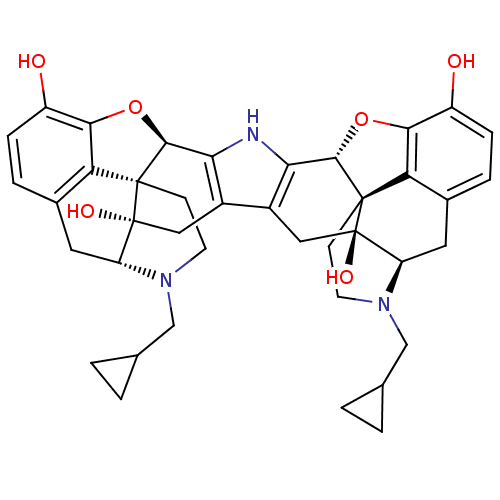
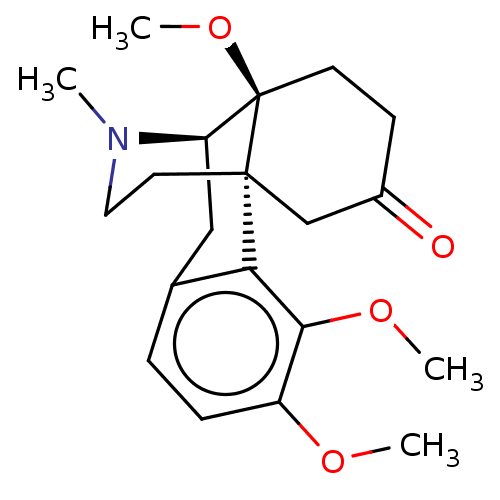
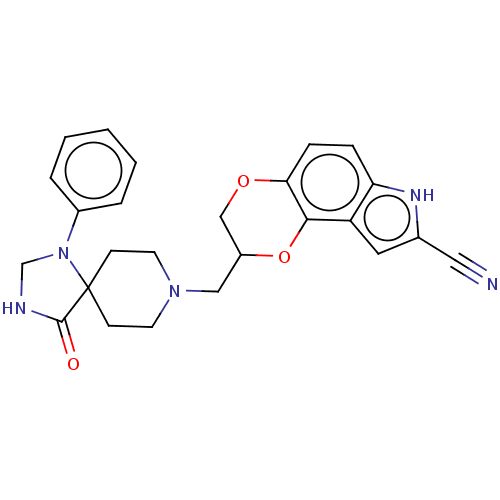
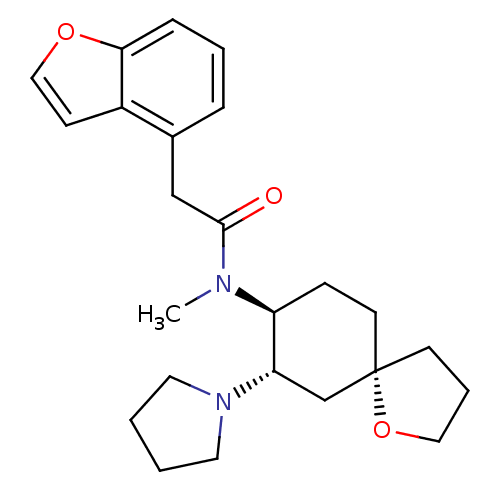
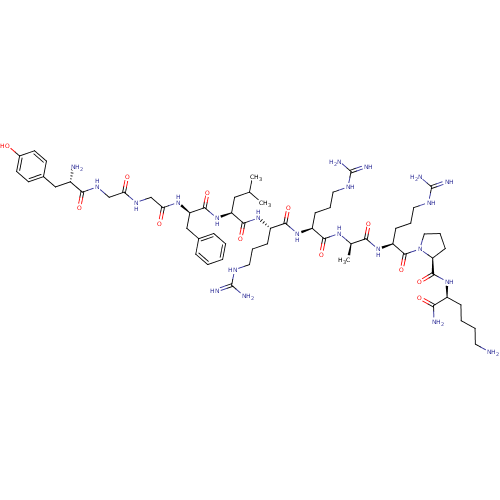
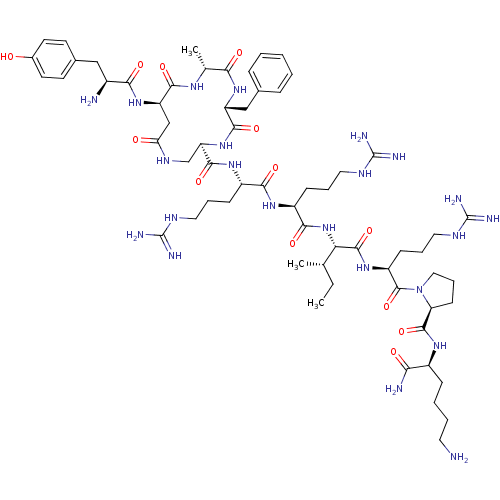
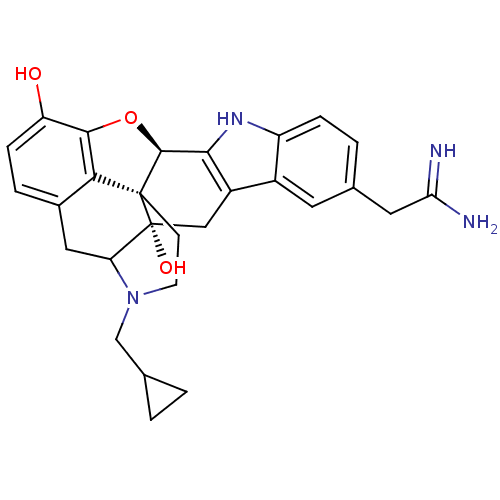
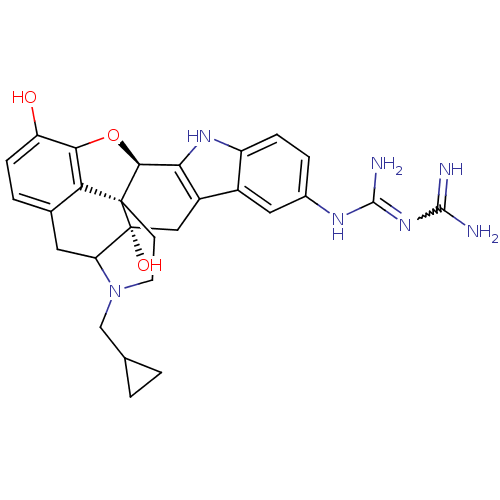
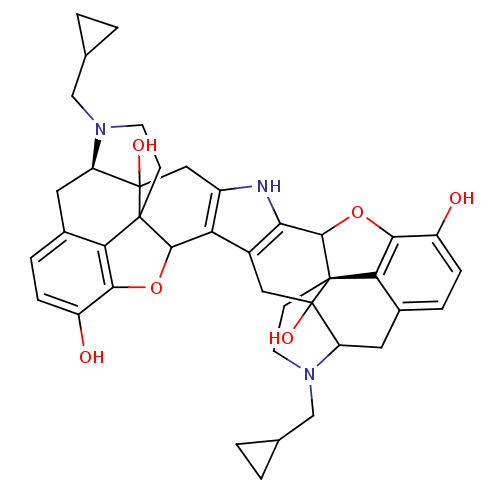
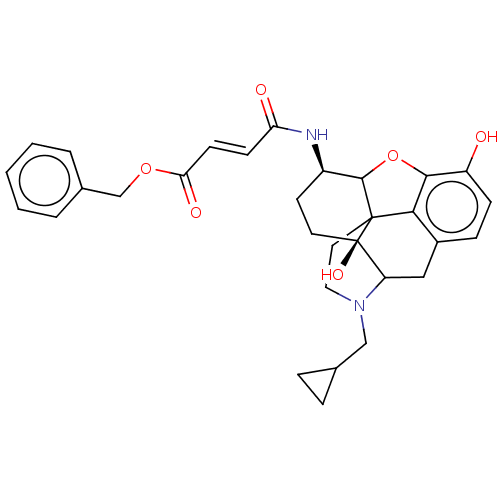
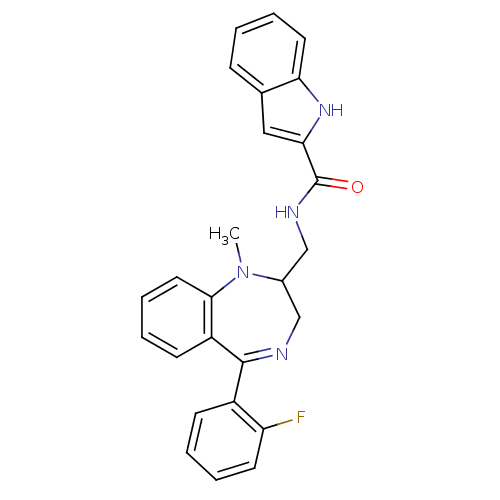
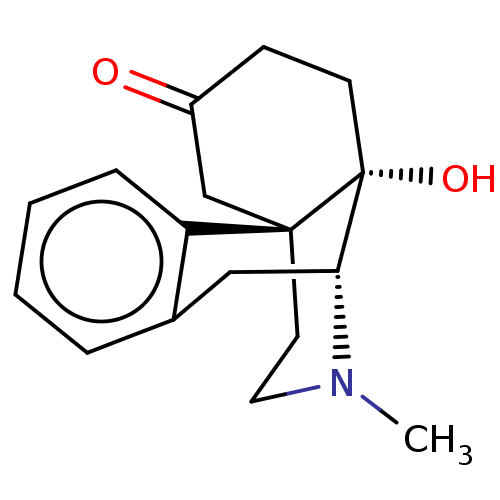
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.00300nMAssay Description:Concentration for 50% inhibition of [3H]naloxone (1 M) binding to opioid receptor in rat brain membrane was determined in the absence of NaClMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.00400nMAssay Description:50% Inhibition of stereospecific [3H]-naltrexone (10e-9 M) binding towards opiate receptor in rat brain homogenateMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.00400nMAssay Description:Concentration for 50% inhibition of [3H]naloxone (1 M) binding to opioid receptor in rat brain membrane was determined in the presence of NaClMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.00900nMAssay Description:Concentration for 50% inhibition of [3H]naloxone (1 M) binding to opioid receptor in rat brain membrane was determined in the absence of NaClMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.0130nMAssay Description:Concentration for 50% inhibition of [3H]naloxone (1 M) binding to opioid receptor in rat brain membrane was determined in the absence of NaClMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.0150nMAssay Description:The compound was tested for binding affinity against opioid receptor in rat brain membranes, using [3H]dihydromorphine as the radioligandMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.0200nMAssay Description:Concentration for 50% inhibition of [3H]naloxone (1 M) binding to opioid receptor in rat brain membrane was determined in the absence of NaClMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataKi: 0.0200nMAssay Description:In vitro affinity to displace [3H]naloxone from opiate receptor in freshly prepared rat brain homogenatesMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.0270nMAssay Description:50% Inhibition of stereospecific [3H]-naltrexone (10e-9 M) binding towards opiate receptor in rat brain homogenateMore data for this Ligand-Target Pair
Affinity DataKd: 0.0290nMAssay Description:Dissociation constant against wild type EL-2 Opioid receptor kappa 1 expressed in HEK cellsMore data for this Ligand-Target Pair
Affinity DataKd: 0.0380nMAssay Description:Dissociation constant against E203Q,D204N,D206N,E209Q EL-2 Opioid receptor kappa 1 expressed in HEK cellsMore data for this Ligand-Target Pair
Affinity DataKd: 0.0500nMAssay Description:Dissociation constant against E203Q,D204N,D206N EL-2 Opioid receptor kappa 1 expressed in HEK cellsMore data for this Ligand-Target Pair
Affinity DataKd: 0.0500nMAssay Description:Dissociation constant against D216N,D217N,E218Q EL-2 Opioid receptor kappa 1 expressed in HEK cellsMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.0500nMAssay Description:Concentration for 50% inhibition of [3H]naloxone (1 M) binding to opioid receptor in rat brain membrane was determined in the absence of NaClMore data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:Mutant factor of the compound against mu-[K303E] with Opioid receptor kappa 1 for binding affinityMore data for this Ligand-Target Pair
Affinity DataKi: 0.0700nMAssay Description:Displacement of [3H]diprenorphine from opioid receptor kappa 1 expressed in HEK 293 cellsMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.0800nMAssay Description:Concentration for 50% inhibition of [3H]naloxone (1 M) binding to opioid receptor in rat brain membrane was determined in the presence of NaClMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.0850nMAssay Description:Concentration for 50% inhibition of [3H]naloxone (1 M) binding to opioid receptor in rat brain membrane was determined in the absence of NaClMore data for this Ligand-Target Pair
Affinity DataKi: 0.0900nMAssay Description:Effect on binding affinity of mutational exchange of Glu297 in the Opioid receptor kappa 1 in transiently expressed rat cos-7 cells activity expresse...More data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataKi: 0.100nMAssay Description:Compound was evaluated for the binding affinity towards opioid receptor from rat whole brain using [3H]etorpine (33.2 Ci/mmol,0.2nanoM)as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 0.100nMAssay Description:Compound was evaluated for the inhibition of binding of [3H]U-69593 to cloned rat Opioid receptor kappa 1 expressed in CHO cell lineMore data for this Ligand-Target Pair
Affinity DataKi: 0.110nMAssay Description:Binding affinity towards rat Opioid receptor kappa 1 expressed in CHO cells of chinese hamster ovary was determined using [3H]diprenorphine as radiol...More data for this Ligand-Target Pair
Affinity DataKi: 0.120nMAssay Description:Effect on binding affinity of mutational exchange of Glu297 in the Opioid receptor kappa 1 in transiently expressed rat cos-7 cells activity expresse...More data for this Ligand-Target Pair
Affinity DataKi: 0.130nMAssay Description:Binding affinity against Opioid receptor kappa 1 isolated from rat brain membrane was determined using [3H]U-69593 as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 0.140nMAssay Description:Binding affinity towards Opioid receptor kappa 1 expressed in CHO cells was determined by using [3H]diprenorphine as radioligandMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.150nMAssay Description:Concentration for 50% inhibition of [3H]naloxone (1 M) binding to opioid receptor in rat brain membrane was determined in the presence of NaClMore data for this Ligand-Target Pair
Affinity DataKi: 0.160nMAssay Description:Binding affinity towards Opioid receptor kappa 1 expressed in CHO cells was determined by using [3H]diprenorphine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 0.180nMAssay Description:Binding affinity against Opioid receptor kappa 1 isolated from rat brain membrane was determined using [3H]U-69593 as radioligandMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.200nMAssay Description:Concentration for 50% inhibition of [3H]naloxone (1 M) binding to opioid receptor in rat brain membrane was determined in the absence of NaClMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.200nMAssay Description:Concentration for 50% inhibition of [3H]naloxone (1 M) binding to opioid receptor in rat brain membrane was determined in the absence of NaClMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Binding affinity towards Opioid receptor kappa 1 expressed in CHO cells was determined by using [3H]diprenorphine as radioligandMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.200nMAssay Description:Concentration for 50% inhibition of [3H]naloxone (1 M) binding to opioid receptor in rat brain membrane was determined in the absence of NaClMore data for this Ligand-Target Pair
Affinity DataKi: 0.210nMAssay Description:Binding affinity towards Opioid receptor kappa 1 expressed in CHO cells was determined by using [3H]diprenorphine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 0.214nMAssay Description:Binding affinity on HEK293 cells transiently transfected with plasmids encoding the rat Opioid receptor kappa 1More data for this Ligand-Target Pair
Affinity DataKi: 0.226nMAssay Description:Binding affinity on HEK293 cells transiently transfected with plasmids encoding the rat Opioid receptor kappa 1More data for this Ligand-Target Pair
Affinity DataKi: 0.240nMAssay Description:Displacement of [3H]diprenorphine from delta-W284E-receptor expressed in HEK 293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.244nMAssay Description:Binding affinity on HEK293 cells transiently transfected with plasmids encoding the rat Opioid receptor kappa 1More data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataKi: 0.25nMAssay Description:Inhibition of [3H]diprenorphine binding to rat brain membrane opioid receptors;(T= total opioid receptor family)More data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.25nMAssay Description:Displacement of [3H]nalotrexone from rat-brain Opioid receptorsMore data for this Ligand-Target Pair
Affinity DataKi: 0.260nMAssay Description:Binding affinity towards Opioid receptor kappa 1 expressed in CHO cells was determined by using [3H]diprenorphine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 0.270nMAssay Description:Effect on binding affinity of mutational exchange of Glu297 in the Opioid receptor kappa 1 in transiently expressed rat cos-7 cells activity expresse...More data for this Ligand-Target Pair
Affinity DataKi: 0.290nMAssay Description:Mutant factor against mu-[K303E] with Opioid receptor kappa 1 for binding affinityMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.300nMAssay Description:Concentration for 50% inhibition of [3H]naloxone (1 M) binding to opioid receptor in rat brain membrane was determined in the absence of NaClMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.300nMAssay Description:The compound was tested for binding affinity against opioid receptor in rat brain membranes, using [3H]dihydromorphine as the radioligandMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.300nMAssay Description:Displacement of [3H]dihydromorphine (DHM) from rat brain Opioid receptorsMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 0.300nMAssay Description:Concentration for 50% inhibition of [3H]naloxone (1 M) binding to opioid receptor in rat brain membrane was determined in the absence of NaClMore data for this Ligand-Target Pair
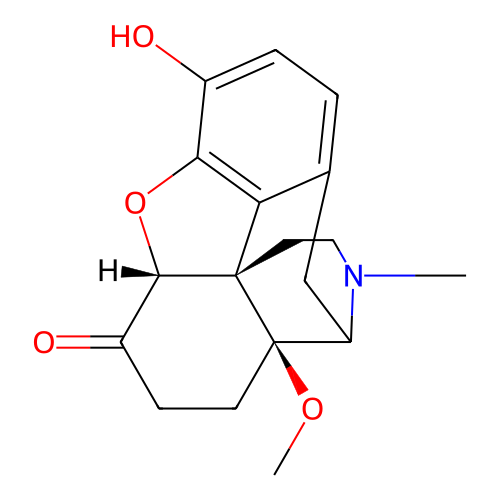
 3D Structure (crystal)
3D Structure (crystal)