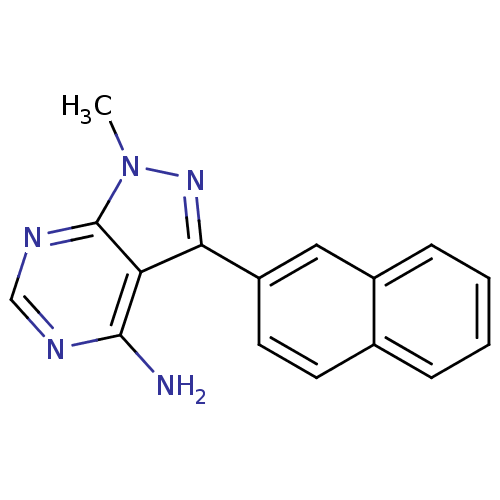
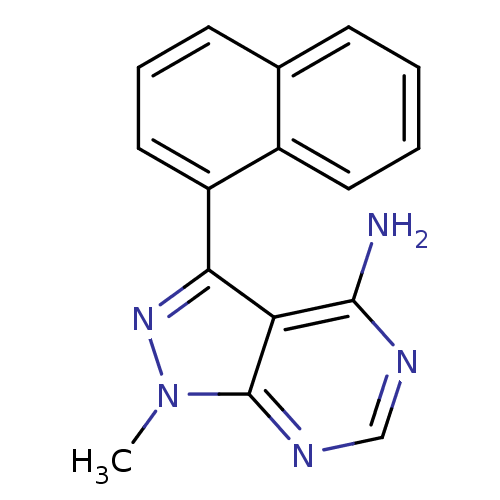
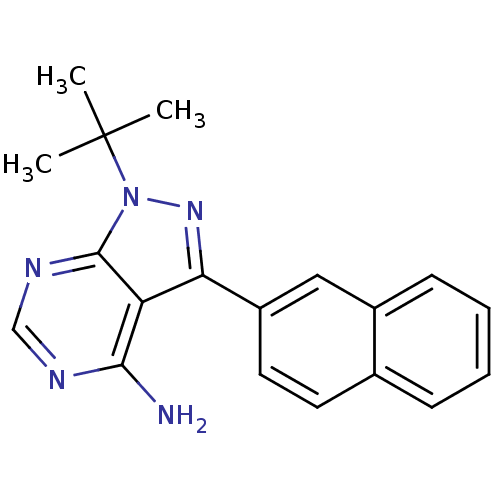
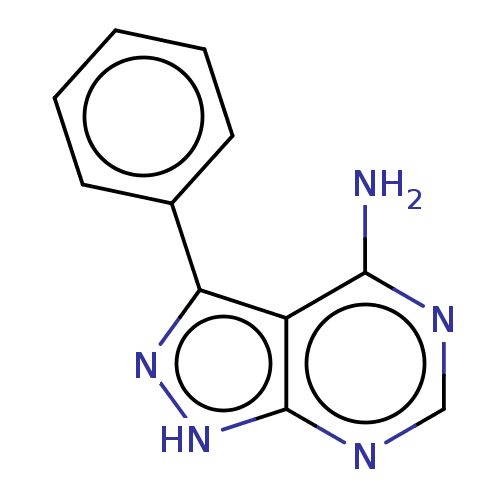
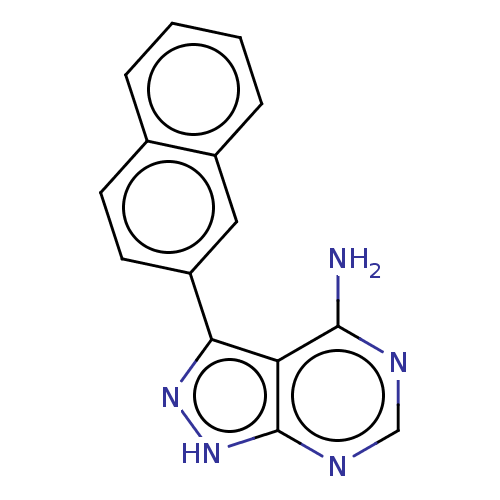
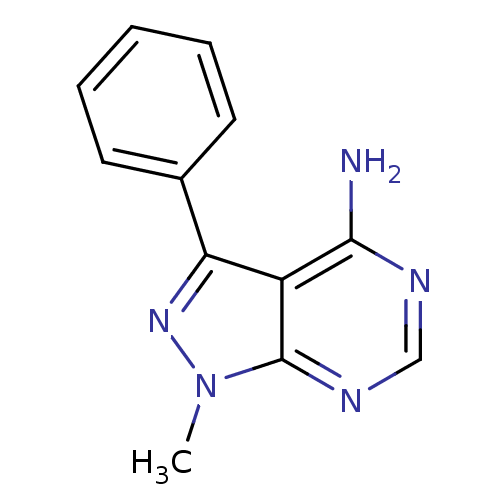
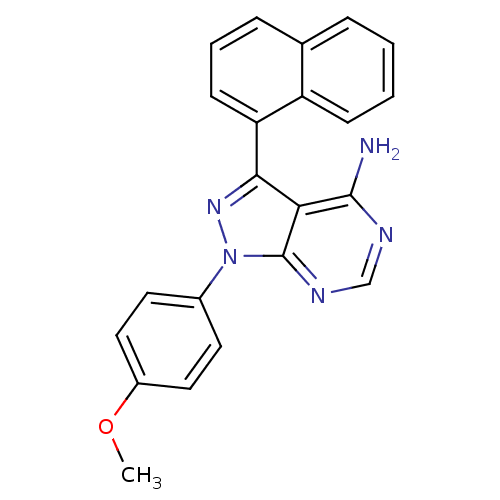
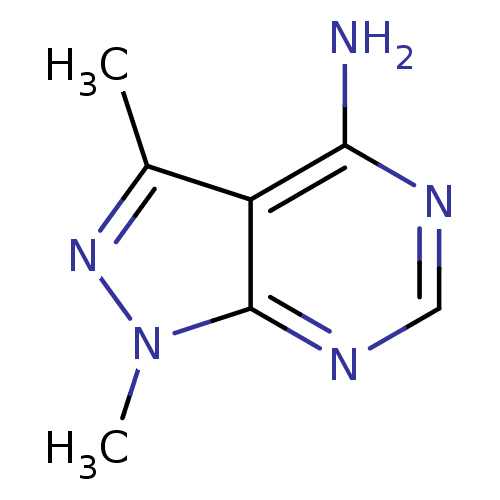
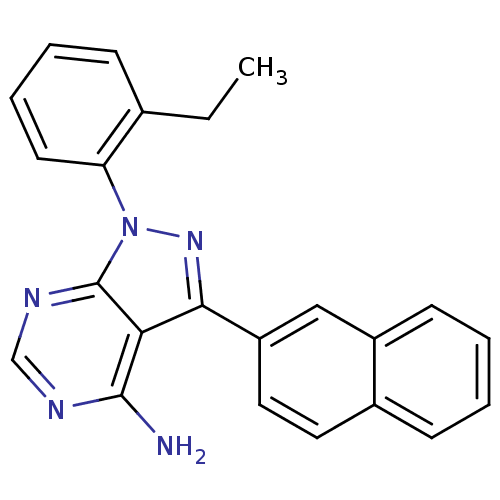
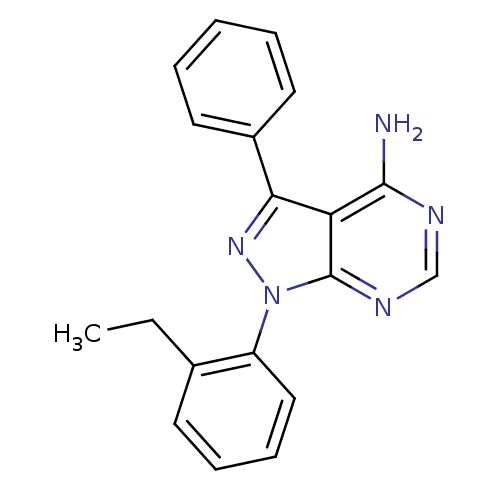
Affinity DataIC50: 12nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair
Affinity DataIC50: 25nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair
Affinity DataIC50: 65nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair
Affinity DataIC50: 105nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair
Affinity DataIC50: 125nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair
Affinity DataIC50: 250nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair
Affinity DataIC50: 350nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair
Affinity DataIC50: 500nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair
Affinity DataIC50: 875nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair
Affinity DataIC50: 900nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.25E+3nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.20E+3nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.55E+4nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+4nMpH: 8.0Assay Description:A NaOH stock solution (50 mM) was standardized with KHP then diluted in Millipore water (10-fold serial dilutions) then used to hold the pH constant ...More data for this Ligand-Target Pair