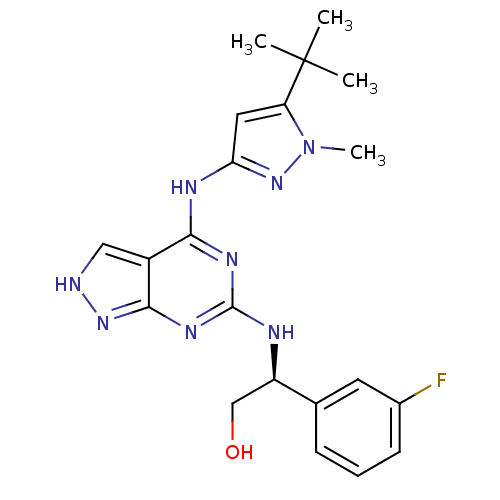
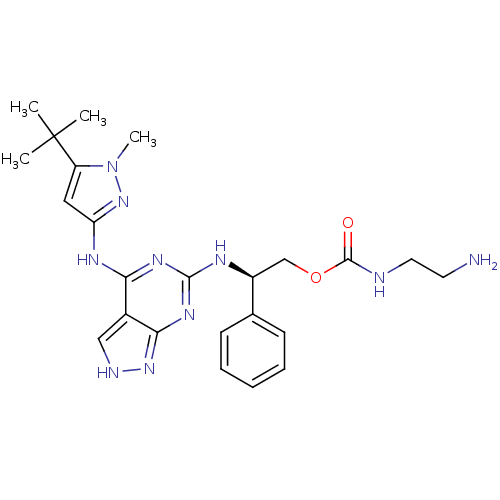
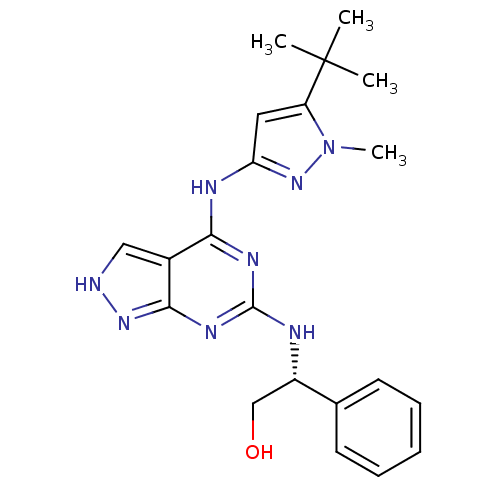
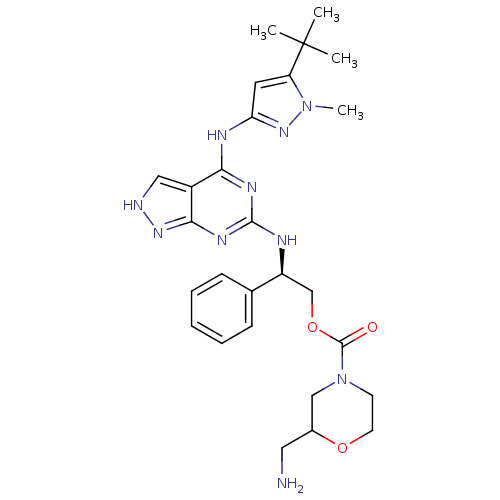
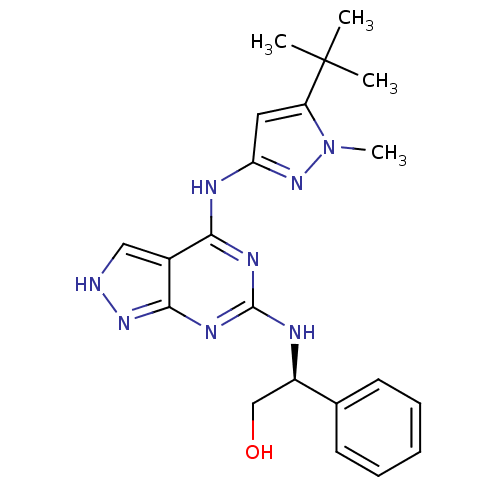
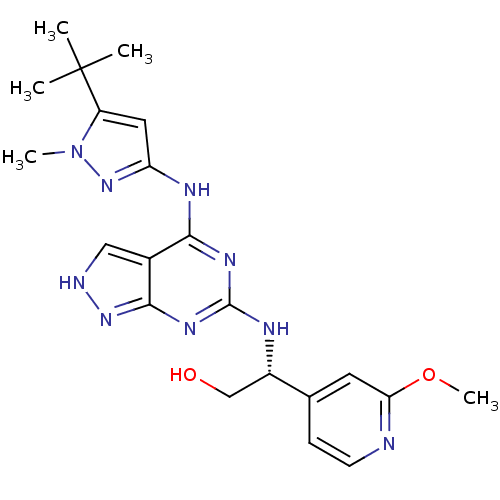
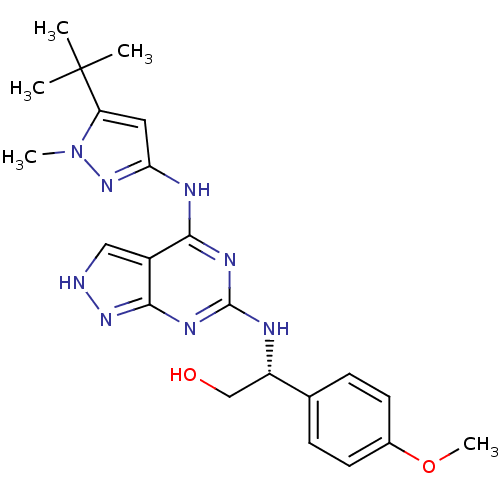
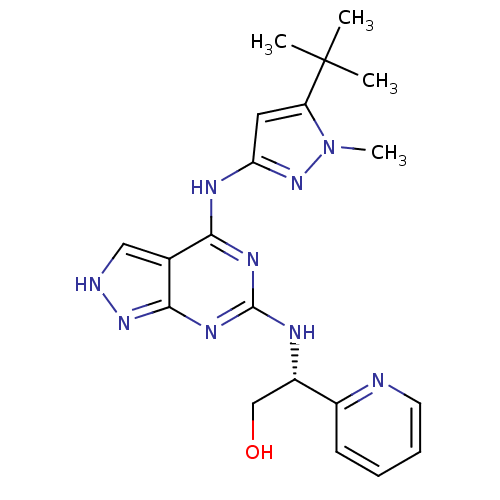
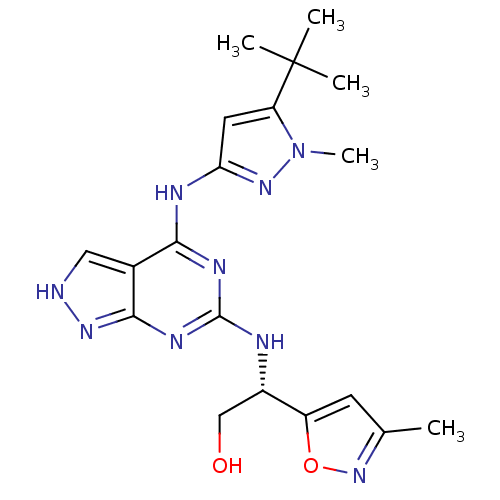
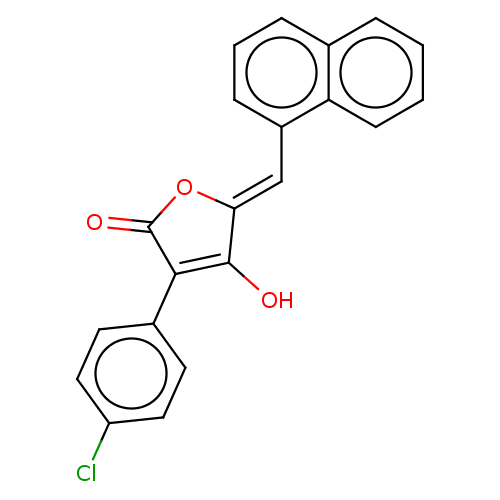
Affinity DataIC50: 1nMT: 2°CAssay Description:The reactions (25 μL) were carried out in 25 mM Tris-HCl pH 8.0, 10 mM ammonium sulfate, 1.25 mM DTT, 0.002% Brij-35, 10 mM MgCl2, 40 μM UN...More data for this Ligand-Target Pair
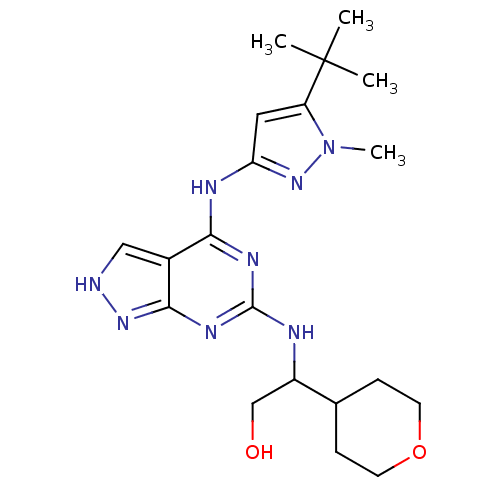
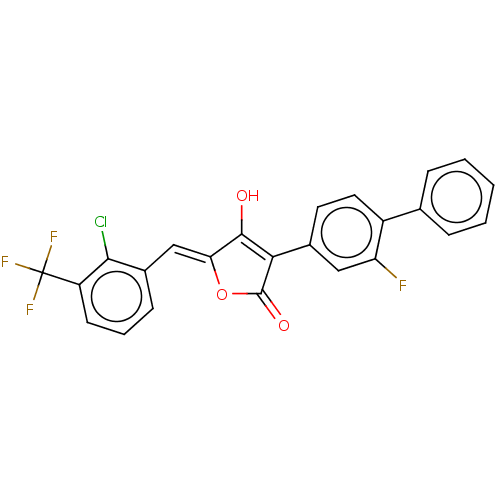
Affinity DataIC50: 2nMT: 2°CAssay Description:The reactions (25 μL) were carried out in 25 mM Tris-HCl pH 8.0, 10 mM ammonium sulfate, 1.25 mM DTT, 0.002% Brij-35, 10 mM MgCl2, 40 μM UN...More data for this Ligand-Target Pair
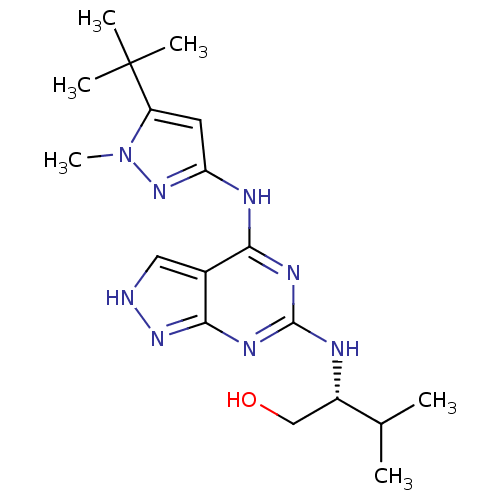
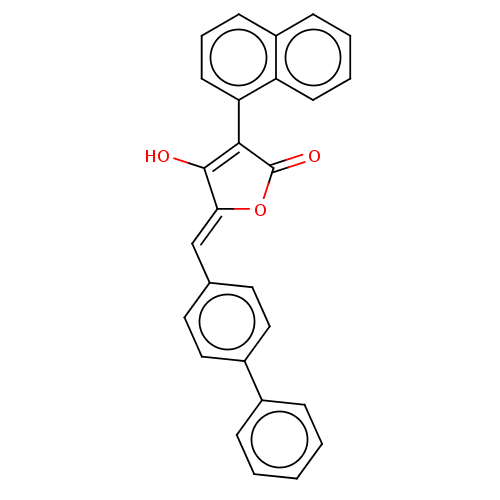
Affinity DataIC50: 3nMT: 2°CAssay Description:The reactions (25 μL) were carried out in 25 mM Tris-HCl pH 8.0, 10 mM ammonium sulfate, 1.25 mM DTT, 0.002% Brij-35, 10 mM MgCl2, 40 μM UN...More data for this Ligand-Target Pair
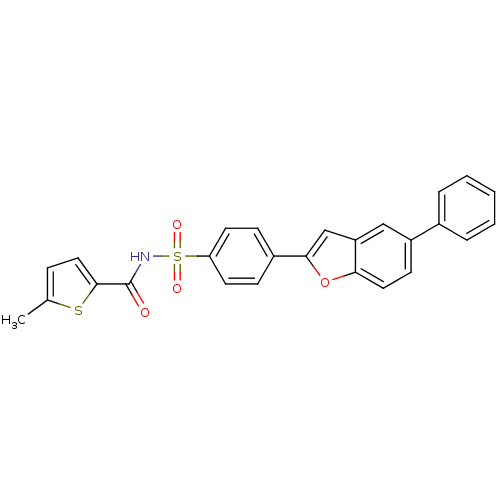
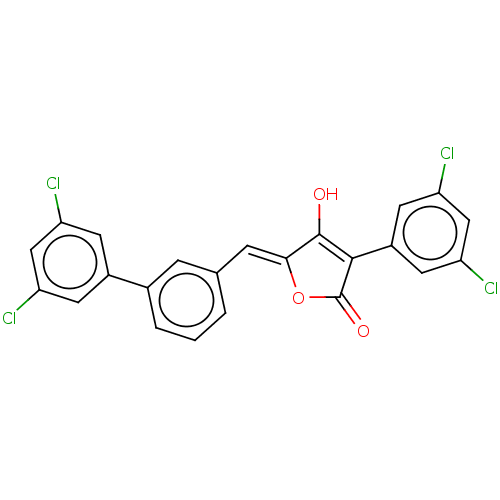
Affinity DataIC50: 3nMT: 2°CAssay Description:The reactions (25 μL) were carried out in 25 mM Tris-HCl pH 8.0, 10 mM ammonium sulfate, 1.25 mM DTT, 0.002% Brij-35, 10 mM MgCl2, 40 μM UN...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMT: 2°CAssay Description:The reactions (25 μL) were carried out in 25 mM Tris-HCl pH 8.0, 10 mM ammonium sulfate, 1.25 mM DTT, 0.002% Brij-35, 10 mM MgCl2, 40 μM UN...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMT: 2°CAssay Description:The reactions (25 μL) were carried out in 25 mM Tris-HCl pH 8.0, 10 mM ammonium sulfate, 1.25 mM DTT, 0.002% Brij-35, 10 mM MgCl2, 40 μM UN...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMT: 2°CAssay Description:The reactions (25 μL) were carried out in 25 mM Tris-HCl pH 8.0, 10 mM ammonium sulfate, 1.25 mM DTT, 0.002% Brij-35, 10 mM MgCl2, 40 μM UN...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMT: 2°CAssay Description:The reactions (25 μL) were carried out in 25 mM Tris-HCl pH 8.0, 10 mM ammonium sulfate, 1.25 mM DTT, 0.002% Brij-35, 10 mM MgCl2, 40 μM UN...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMT: 2°CAssay Description:The reactions (25 μL) were carried out in 25 mM Tris-HCl pH 8.0, 10 mM ammonium sulfate, 1.25 mM DTT, 0.002% Brij-35, 10 mM MgCl2, 40 μM UN...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMT: 2°CAssay Description:The reactions (25 μL) were carried out in 25 mM Tris-HCl pH 8.0, 10 mM ammonium sulfate, 1.25 mM DTT, 0.002% Brij-35, 10 mM MgCl2, 40 μM UN...More data for this Ligand-Target Pair
Affinity DataIC50: 21nMT: 2°CAssay Description:The reactions (25 μL) were carried out in 25 mM Tris-HCl pH 8.0, 10 mM ammonium sulfate, 1.25 mM DTT, 0.002% Brij-35, 10 mM MgCl2, 40 μM UN...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMT: 2°CAssay Description:The reactions (25 μL) were carried out in 25 mM Tris-HCl pH 8.0, 10 mM ammonium sulfate, 1.25 mM DTT, 0.002% Brij-35, 10 mM MgCl2, 40 μM UN...More data for this Ligand-Target Pair
Affinity DataIC50: 52nMT: 2°CAssay Description:The reactions (25 μL) were carried out in 25 mM Tris-HCl pH 8.0, 10 mM ammonium sulfate, 1.25 mM DTT, 0.002% Brij-35, 10 mM MgCl2, 40 μM UN...More data for this Ligand-Target Pair
Affinity DataIC50: 57nMT: 2°CAssay Description:The reactions (25 μL) were carried out in 25 mM Tris-HCl pH 8.0, 10 mM ammonium sulfate, 1.25 mM DTT, 0.002% Brij-35, 10 mM MgCl2, 40 μM UN...More data for this Ligand-Target Pair
Affinity DataIC50: 60nMT: 2°CAssay Description:The reactions (25 μL) were carried out in 25 mM Tris-HCl pH 8.0, 10 mM ammonium sulfate, 1.25 mM DTT, 0.002% Brij-35, 10 mM MgCl2, 40 μM UN...More data for this Ligand-Target Pair
Affinity DataIC50: 60nMT: 2°CAssay Description:The reactions (25 μL) were carried out in 25 mM Tris-HCl pH 8.0, 10 mM ammonium sulfate, 1.25 mM DTT, 0.002% Brij-35, 10 mM MgCl2, 40 μM UN...More data for this Ligand-Target Pair
Affinity DataIC50: 101nMT: 2°CAssay Description:The reactions (25 μL) were carried out in 25 mM Tris-HCl pH 8.0, 10 mM ammonium sulfate, 1.25 mM DTT, 0.002% Brij-35, 10 mM MgCl2, 40 μM UN...More data for this Ligand-Target Pair
Affinity DataIC50: 188nMT: 2°CAssay Description:The reactions (25 μL) were carried out in 25 mM Tris-HCl pH 8.0, 10 mM ammonium sulfate, 1.25 mM DTT, 0.002% Brij-35, 10 mM MgCl2, 40 μM UN...More data for this Ligand-Target Pair
Affinity DataIC50: 4.50E+3nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 1.08E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 1.22E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 1.27E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 1.56E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 1.56E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 1.56E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 1.63E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 1.77E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 1.87E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 1.95E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 2.17E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 2.56E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 2.95E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 3.40E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 3.55E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 3.86E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+4nMAssay Description:Inhibition of Escherichia coli MurC after 15 mins in presence of 0.005% Triton X-114 by Malachite green assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.26E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 4.40E+4nMAssay Description:Inhibition of Escherichia coli MurC after 15 mins in presence of 0.005% Triton X-114 by Malachite green assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.64E+4nMAssay Description:Inhibitory activity against MurC in Escherichia coliMore data for this Ligand-Target Pair