Report error Found 685 with Last Name = 'kwiatkowski' and Initial = 'n'
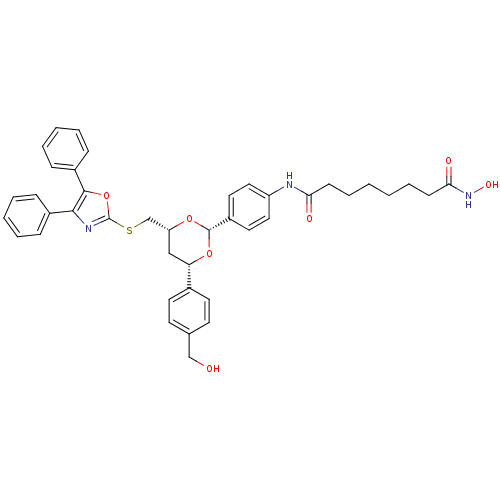
Affinity DataKi: 21nM ΔG°: -43.4kJ/molepH: 7.4 T: 22°CAssay Description:To assess the effect of test compounds on histone deacetylase enzyme function in Vitro, a fluorometric assay was performed using HDAC, which incubate...More data for this Ligand-Target Pair
Affinity DataKi: 28nM ΔG°: -42.7kJ/molepH: 7.4 T: 22°CAssay Description:To assess the effect of test compounds on histone deacetylase enzyme function in Vitro, a fluorometric assay was performed using HDAC, which incubate...More data for this Ligand-Target Pair
Affinity DataKi: 48nM ΔG°: -41.4kJ/molepH: 7.4 T: 22°CAssay Description:To assess the effect of test compounds on histone deacetylase enzyme function in Vitro, a fluorometric assay was performed using HDAC, which incubate...More data for this Ligand-Target Pair
Affinity DataKi: 88nM ΔG°: -39.9kJ/molepH: 7.4 T: 22°CAssay Description:To assess the effect of test compounds on histone deacetylase enzyme function in Vitro, a fluorometric assay was performed using HDAC, which incubate...More data for this Ligand-Target Pair
Affinity DataKi: 142nM ΔG°: -38.7kJ/molepH: 7.4 T: 22°CAssay Description:To assess the effect of test compounds on histone deacetylase enzyme function in Vitro, a fluorometric assay was performed using HDAC, which incubate...More data for this Ligand-Target Pair
Affinity DataKi: 995nM ΔG°: -33.9kJ/molepH: 7.4 T: 22°CAssay Description:To assess the effect of test compounds on histone deacetylase enzyme function in Vitro, a fluorometric assay was performed using HDAC, which incubate...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+3nM ΔG°: -32.2kJ/molepH: 7.4 T: 22°CAssay Description:To assess the effect of test compounds on histone deacetylase enzyme function in Vitro, a fluorometric assay was performed using HDAC, which incubate...More data for this Ligand-Target Pair
Affinity DataKi: 6.10E+3nM ΔG°: -29.5kJ/molepH: 7.4 T: 22°CAssay Description:To assess the effect of test compounds on histone deacetylase enzyme function in Vitro, a fluorometric assay was performed using HDAC, which incubate...More data for this Ligand-Target Pair
Affinity DataKi: 6.30E+3nM ΔG°: -29.4kJ/molepH: 7.4 T: 22°CAssay Description:To assess the effect of test compounds on histone deacetylase enzyme function in Vitro, a fluorometric assay was performed using HDAC, which incubate...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80nMAssay Description:Inhibition of recombinant human full-length His-tagged CDK9/Cyclin T1 expressed in baculovirus infection system using Cdk7/9tide as substrate measure...More data for this Ligand-Target Pair
Affinity DataIC50: 2.60nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 3.20nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 3.20nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 3.20nMAssay Description:Inhibition of recombinant human full-length His-tagged CDK9/Cyclin T1 expressed in baculovirus infection system using Cdk7/9tide as substrate measure...More data for this Ligand-Target Pair
Affinity DataIC50: 3.40nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 3.70nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 5.10nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 5.60nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 5.60nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 5.90nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Inhibition of recombinant human full-length His-tagged CDK9/Cyclin T1 expressed in baculovirus infection system using Cdk7/9tide as substrate measure...More data for this Ligand-Target Pair
Affinity DataIC50: 6.10nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 7.20nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:In vitro biochemical assays were performed in parallel to determine the most potent tool compound.More data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:Inhibition of human CDK19 (M1 to N360 residues) expressed in bacterial expression system by KINOMEscan assayMore data for this Ligand-Target Pair
Affinity DataIC50: 8.40nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 8.80nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 9.10nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 9.86nMAssay Description:Jurkat cells were treated with DMSO or concentration of compound indicated. 6 hours after treatment, cells were washed and harvested by resuspending ...More data for this Ligand-Target Pair
Affinity DataIC50: 10.7nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Inhibition of recombinant human full-length N-terminal GST-tagged CDK2/cyclinA1 expressed in baculovirus infected Sf9 insect cells using FRET-labeled...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Inhibition of recombinant human full-length N-terminal GST-tagged CDK2/cyclinA1 expressed in baculovirus infected Sf9 insect cells using FRET-labeled...More data for this Ligand-Target Pair
Affinity DataIC50: 11.3nMAssay Description:In vitro kinase assays with immunoprecipitated proteins (or recombinant CAK complexes) were performed as follows. Kinase reactions were performed in ...More data for this Ligand-Target Pair
Affinity DataIC50: 11.6nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 11.9nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 11.9nMAssay Description:Jurkat cells were treated with DMSO or concentration of compound indicated. 6 hours after treatment, cells were washed and harvested by resuspending ...More data for this Ligand-Target Pair
Affinity DataIC50: 13.4nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 13.9nMAssay Description:In vitro kinase assays with immunoprecipitated proteins (or recombinant CAK complexes) were performed as follows. Kinase reactions were performed in ...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMAssay Description:Inhibition of CDK9/Cyclin T1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 14.3nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 14.5nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 14.9nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Inhibition of CDK8 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Inhibition of recombinant human full-length His-tagged CDK9/Cyclin T1 expressed in baculovirus infection system using Cdk7/9tide as substrate measure...More data for this Ligand-Target Pair
Affinity DataIC50: 15.2nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 15.5nMAssay Description:In vitro kinase assay of Aurora A, B, and C using Z-LYTE technology (Invitrogen) and ATP at Km apparent for each kinase.More data for this Ligand-Target Pair
Affinity DataIC50: 15.7nMAssay Description:Jurkat cells were treated with DMSO or concentration of compound indicated. 6 hours after treatment, cells were washed and harvested by resuspending ...More data for this Ligand-Target Pair
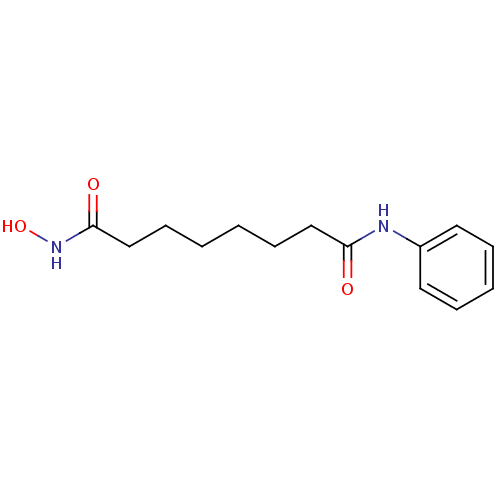
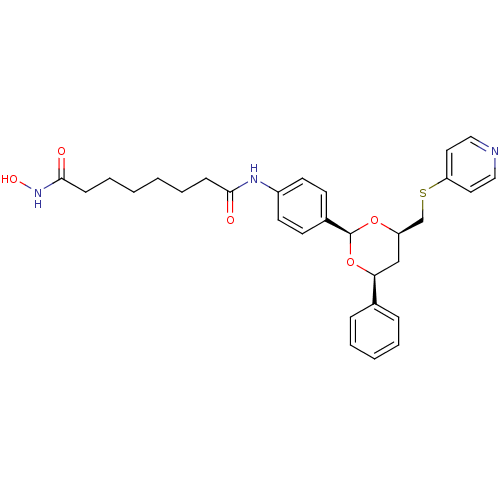
 3D Structure (crystal)
3D Structure (crystal)