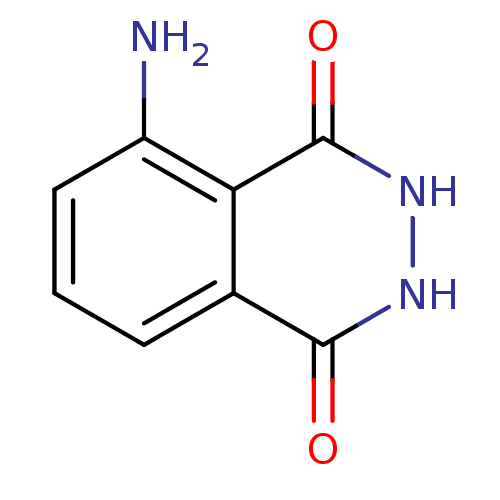
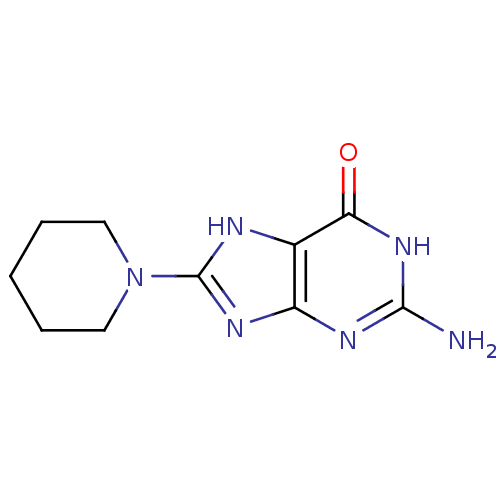
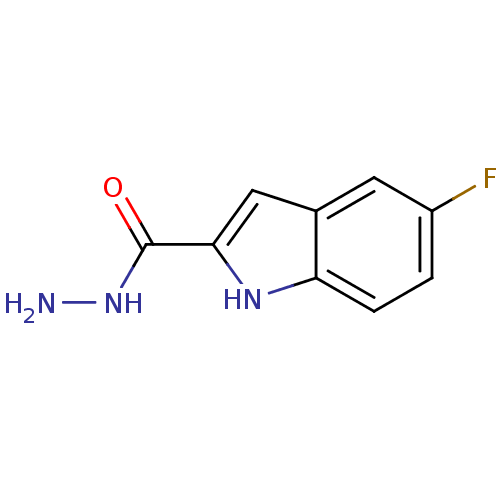
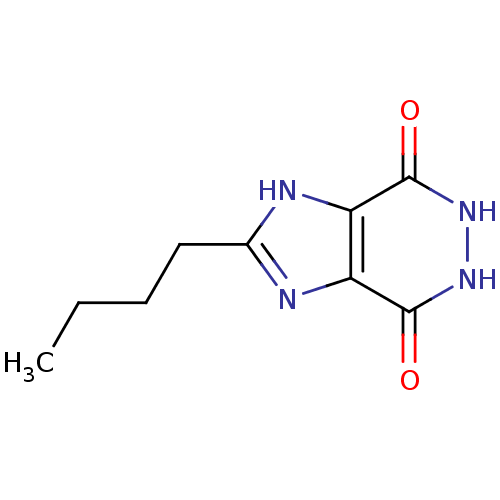
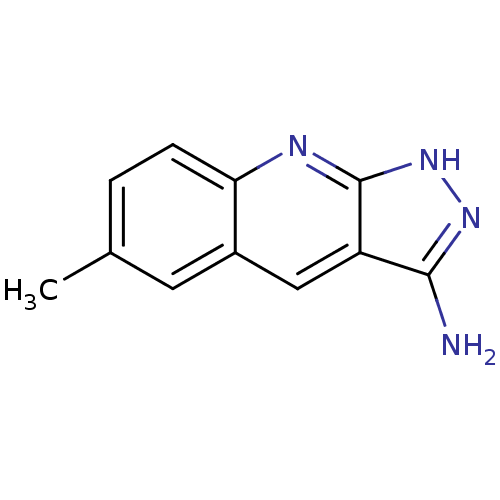
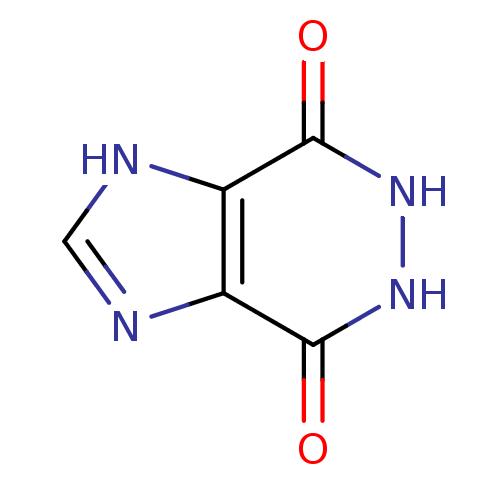
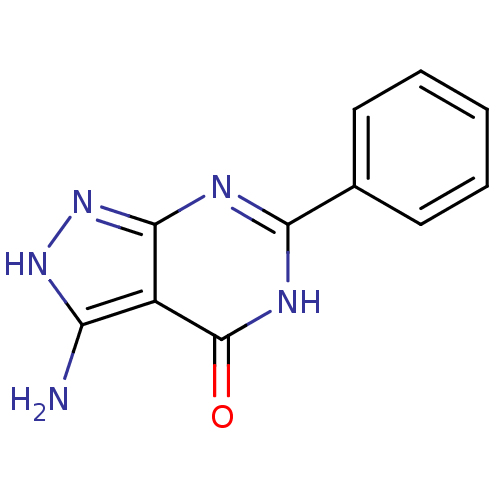
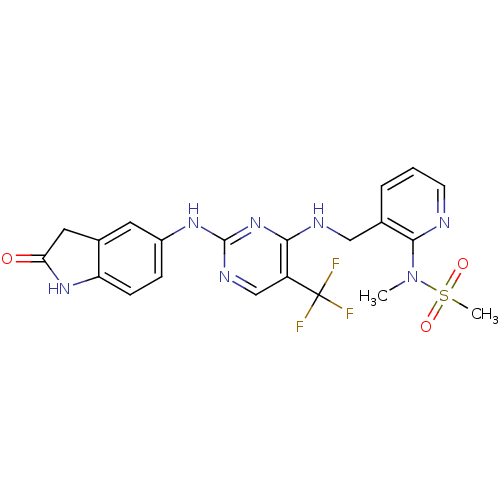
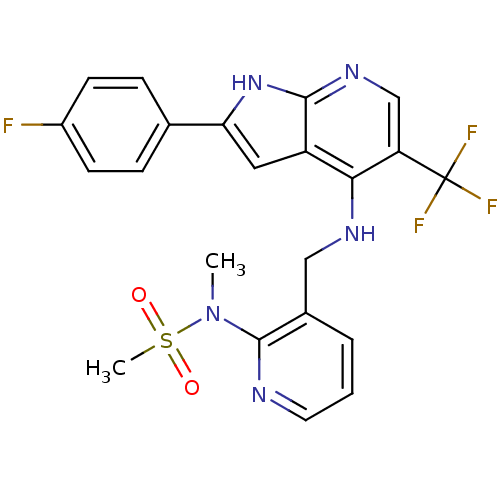
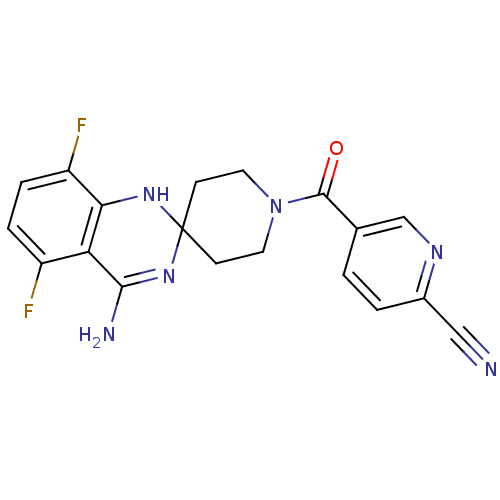
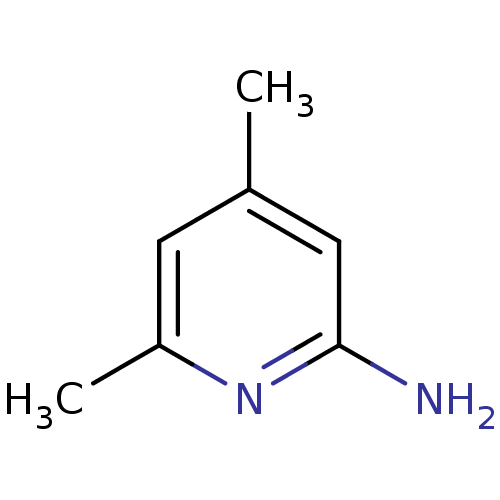
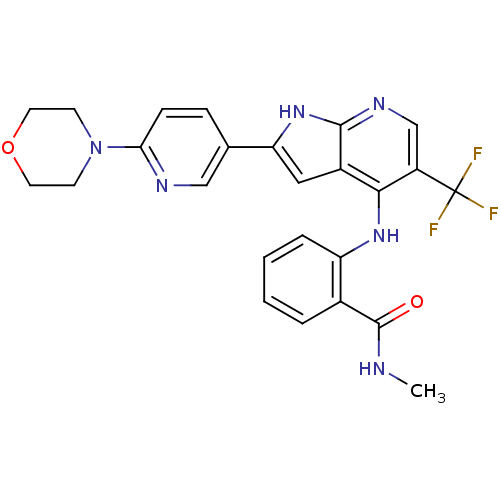
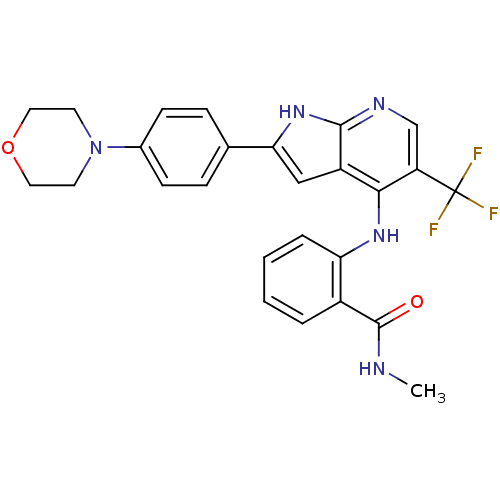
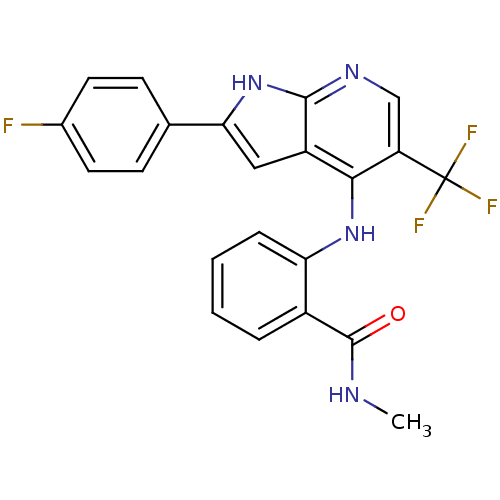
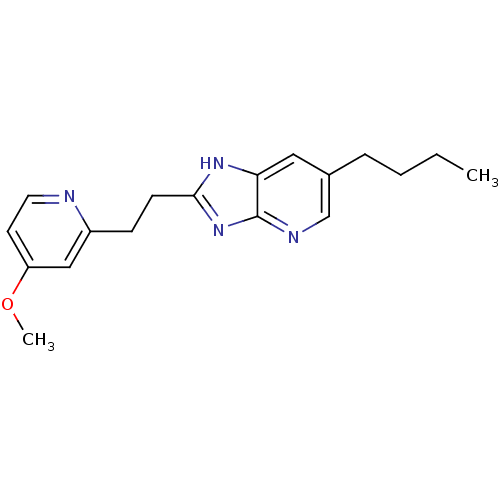
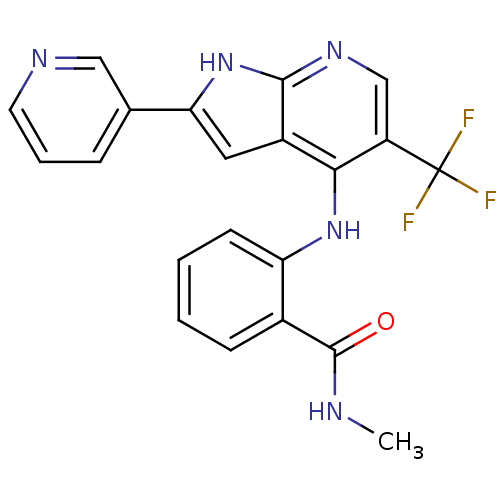
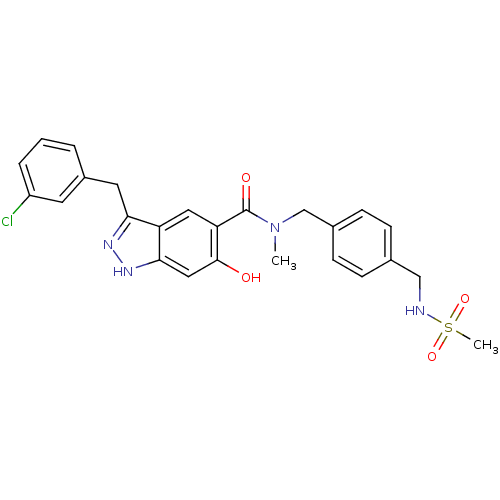
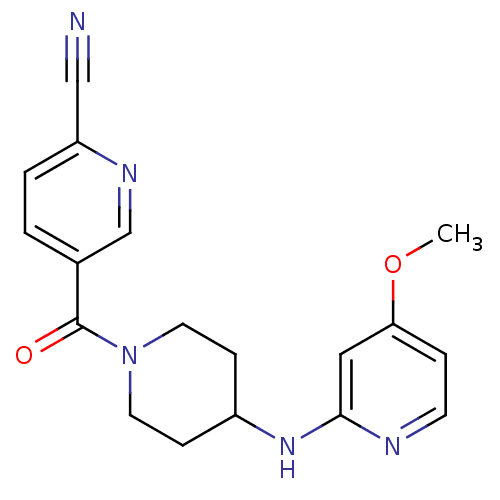
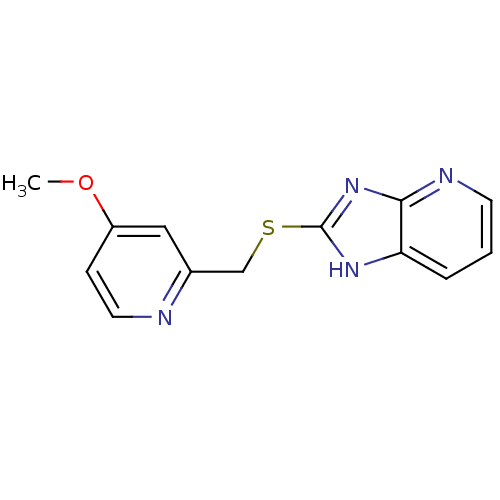
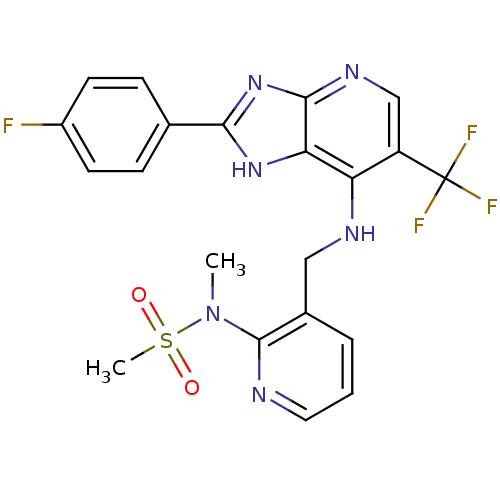
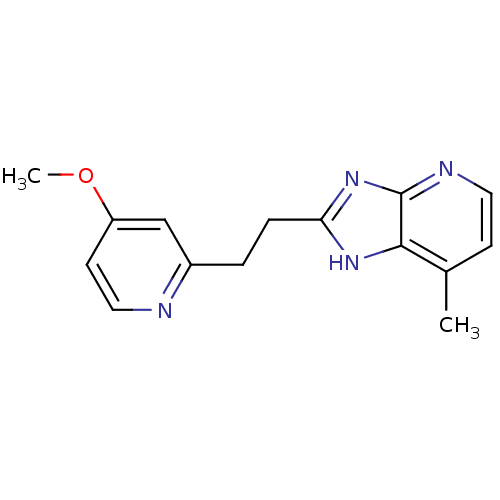
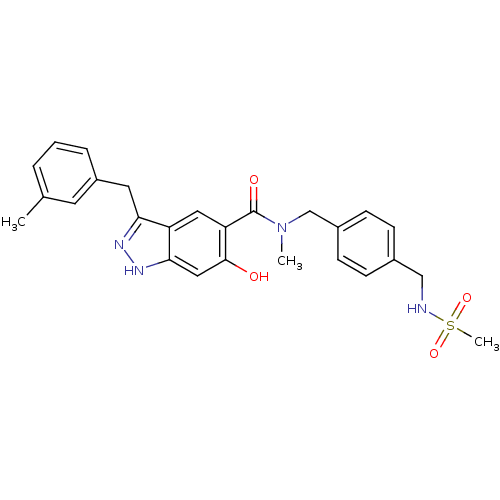
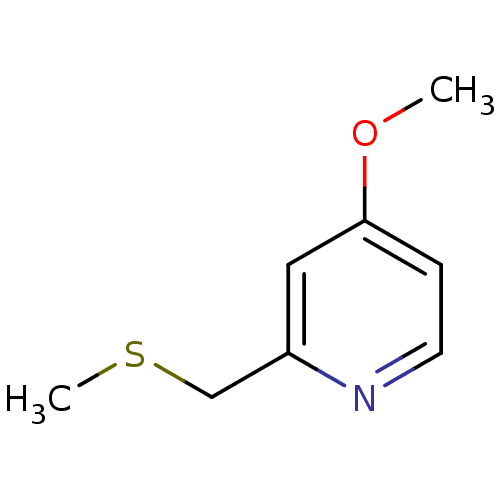
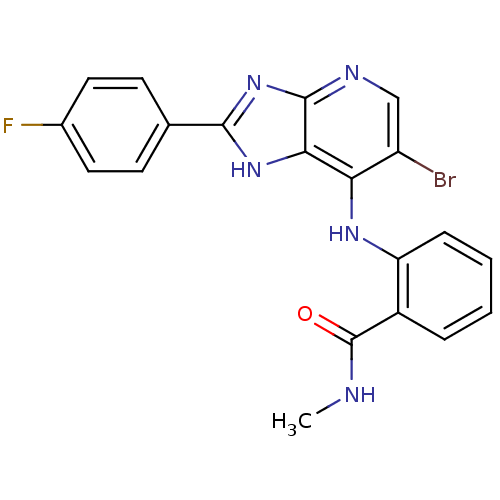
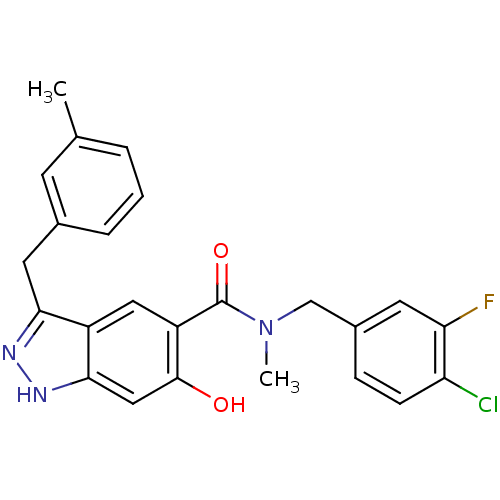
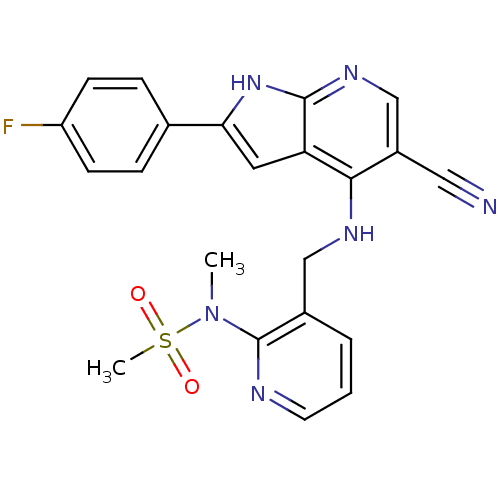
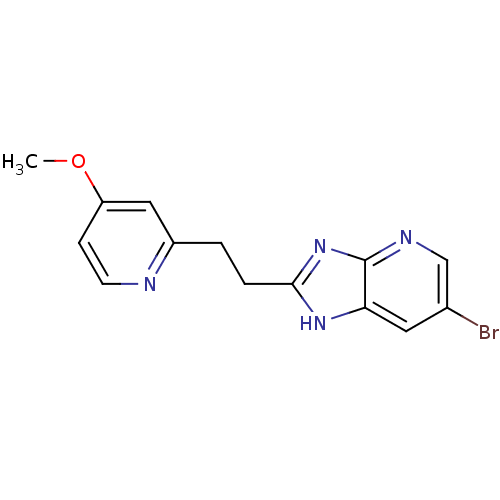
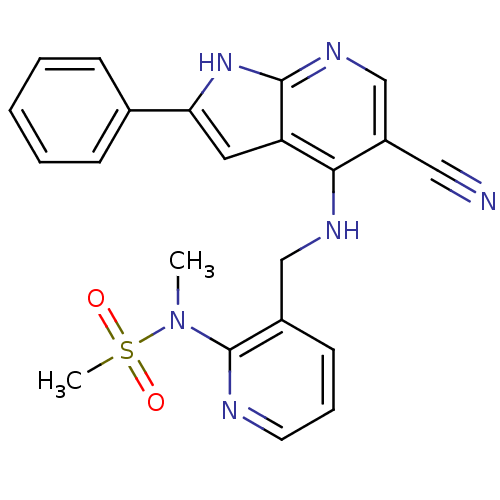
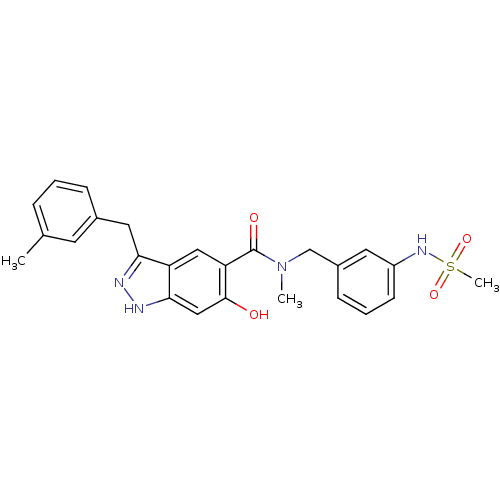
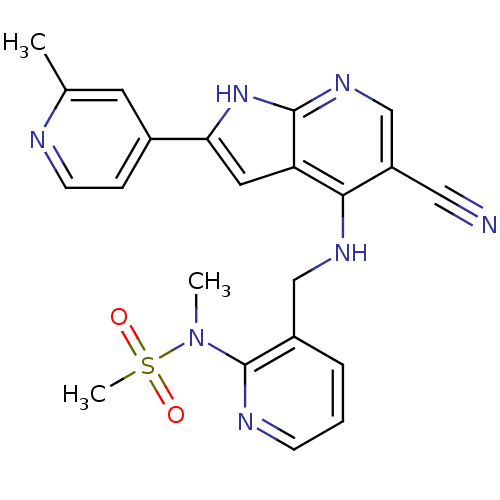
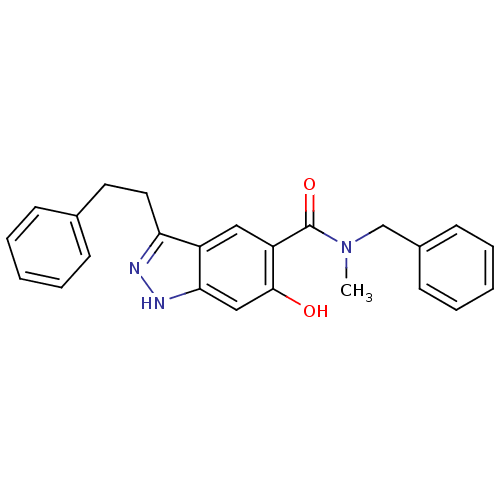
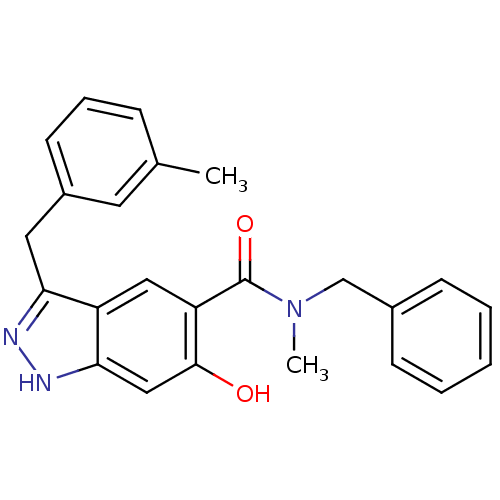
Affinity DataKi: 1.86nMAssay Description:Inhibition of human iNOS assessed as inhibition of [3H]L-arginine to [3H]L-citrulline conversion by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 18.6nMAssay Description:Inhibition of human eNOS assessed as inhibition of [3H]L-arginine to [3H]L-citrulline conversion by scintillation countingMore data for this Ligand-Target Pair
TargetQueuine tRNA-ribosyltransferase(Zymomonas mobilis)
Philipps-UniversitäT Marburg
Curated by ChEMBL
Philipps-UniversitäT Marburg
Curated by ChEMBL
Affinity DataKi: 100nMAssay Description:Inhibitory activity against eubacterial tRNA guanine transglycosylase (TGT) from Zymomonas mobilisMore data for this Ligand-Target Pair
TargetQueuine tRNA-ribosyltransferase(Zymomonas mobilis)
Philipps-UniversitäT Marburg
Curated by ChEMBL
Philipps-UniversitäT Marburg
Curated by ChEMBL
Affinity DataKi: 250nMAssay Description:Inhibitory activity against eubacterial tRNA guanine transglycosylase (TGT) from Zymomonas mobilisMore data for this Ligand-Target Pair
TargetQueuine tRNA-ribosyltransferase(Zymomonas mobilis)
Philipps-UniversitäT Marburg
Curated by ChEMBL
Philipps-UniversitäT Marburg
Curated by ChEMBL
Affinity DataKi: 300nMAssay Description:Inhibitory activity against eubacterial tRNA guanine transglycosylase (TGT) from Zymomonas mobilisMore data for this Ligand-Target Pair
TargetQueuine tRNA-ribosyltransferase(Zymomonas mobilis)
Philipps-UniversitäT Marburg
Curated by ChEMBL
Philipps-UniversitäT Marburg
Curated by ChEMBL
Affinity DataKi: 600nMAssay Description:Inhibitory activity against eubacterial tRNA guanine transglycosylase (TGT) from Zymomonas mobilisMore data for this Ligand-Target Pair
TargetQueuine tRNA-ribosyltransferase(Zymomonas mobilis)
Philipps-UniversitäT Marburg
Curated by ChEMBL
Philipps-UniversitäT Marburg
Curated by ChEMBL
Affinity DataKi: 2.70E+3nMAssay Description:Inhibitory activity against eubacterial tRNA guanine transglycosylase (TGT) from Zymomonas mobilisMore data for this Ligand-Target Pair
TargetQueuine tRNA-ribosyltransferase(Zymomonas mobilis)
Philipps-UniversitäT Marburg
Curated by ChEMBL
Philipps-UniversitäT Marburg
Curated by ChEMBL
Affinity DataKi: 3.80E+3nMAssay Description:Inhibitory activity against eubacterial tRNA guanine transglycosylase (TGT) from Zymomonas mobilisMore data for this Ligand-Target Pair
TargetQueuine tRNA-ribosyltransferase(Zymomonas mobilis)
Philipps-UniversitäT Marburg
Curated by ChEMBL
Philipps-UniversitäT Marburg
Curated by ChEMBL
Affinity DataKi: 4.70E+3nMAssay Description:Inhibitory activity against eubacterial tRNA guanine transglycosylase (TGT) from Zymomonas mobilisMore data for this Ligand-Target Pair
TargetQueuine tRNA-ribosyltransferase(Zymomonas mobilis)
Philipps-UniversitäT Marburg
Curated by ChEMBL
Philipps-UniversitäT Marburg
Curated by ChEMBL
Affinity DataKi: 5.00E+3nMAssay Description:Inhibitory activity against eubacterial tRNA guanine transglycosylase (TGT) from Zymomonas mobilisMore data for this Ligand-Target Pair
TargetQueuine tRNA-ribosyltransferase(Zymomonas mobilis)
Philipps-UniversitäT Marburg
Curated by ChEMBL
Philipps-UniversitäT Marburg
Curated by ChEMBL
Affinity DataKi: 8.10E+3nMAssay Description:Inhibitory activity against eubacterial tRNA guanine transglycosylase (TGT) from Zymomonas mobilisMore data for this Ligand-Target Pair
TargetQueuine tRNA-ribosyltransferase(Zymomonas mobilis)
Philipps-UniversitäT Marburg
Curated by ChEMBL
Philipps-UniversitäT Marburg
Curated by ChEMBL
Affinity DataKi: 8.30E+3nMAssay Description:Inhibitory activity against eubacterial tRNA guanine transglycosylase (TGT) from Zymomonas mobilisMore data for this Ligand-Target Pair
TargetQueuine tRNA-ribosyltransferase(Zymomonas mobilis)
Philipps-UniversitäT Marburg
Curated by ChEMBL
Philipps-UniversitäT Marburg
Curated by ChEMBL
Affinity DataKi: 3.60E+4nMAssay Description:Inhibitory activity against eubacterial tRNA guanine transglycosylase (TGT) from Zymomonas mobilisMore data for this Ligand-Target Pair
TargetQueuine tRNA-ribosyltransferase(Zymomonas mobilis)
Philipps-UniversitäT Marburg
Curated by ChEMBL
Philipps-UniversitäT Marburg
Curated by ChEMBL
Affinity DataKi: 3.70E+4nMAssay Description:Inhibitory activity against eubacterial tRNA guanine transglycosylase (TGT) from Zymomonas mobilisMore data for this Ligand-Target Pair
TargetQueuine tRNA-ribosyltransferase(Zymomonas mobilis)
Philipps-UniversitäT Marburg
Curated by ChEMBL
Philipps-UniversitäT Marburg
Curated by ChEMBL
Affinity DataKi: 7.20E+4nMAssay Description:Inhibitory activity against eubacterial tRNA guanine transglycosylase (TGT) from Zymomonas mobilisMore data for this Ligand-Target Pair
TargetQueuine tRNA-ribosyltransferase(Zymomonas mobilis)
Philipps-UniversitäT Marburg
Curated by ChEMBL
Philipps-UniversitäT Marburg
Curated by ChEMBL
Affinity DataKi: 8.30E+4nMAssay Description:Inhibitory activity against eubacterial tRNA guanine transglycosylase (TGT) from Zymomonas mobilisMore data for this Ligand-Target Pair
TargetQueuine tRNA-ribosyltransferase(Zymomonas mobilis)
Philipps-UniversitäT Marburg
Curated by ChEMBL
Philipps-UniversitäT Marburg
Curated by ChEMBL
Affinity DataKi: 1.56E+5nMAssay Description:Inhibitory activity against eubacterial tRNA guanine transglycosylase (TGT) from Zymomonas mobilisMore data for this Ligand-Target Pair
TargetQueuine tRNA-ribosyltransferase(Zymomonas mobilis)
Philipps-UniversitäT Marburg
Curated by ChEMBL
Philipps-UniversitäT Marburg
Curated by ChEMBL
Affinity DataKi: 2.00E+5nMAssay Description:Inhibitory activity against eubacterial tRNA guanine transglycosylase (TGT) from Zymomonas mobilisMore data for this Ligand-Target Pair
TargetQueuine tRNA-ribosyltransferase(Zymomonas mobilis)
Philipps-UniversitäT Marburg
Curated by ChEMBL
Philipps-UniversitäT Marburg
Curated by ChEMBL
Affinity DataKi: 2.49E+5nMAssay Description:Inhibitory activity against eubacterial tRNA guanine transglycosylase (TGT) from Zymomonas mobilisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.5nMAssay Description:Competitive binding affinity to FAK kinase domain (410 to 689) (unknown origin) assessed as phosphorylation of p(Glu/Tyr) in presence of ATPMore data for this Ligand-Target Pair
Affinity DataIC50: 37nMAssay Description:Inhibition of FAK (unknown origin) using biotinylated His-TEV-hsFAK(31-686)(K454R) substrate after 2 hrs by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 37nMAssay Description:Inhibition of FAK (unknown origin) using biotinylated His-TEV-hsFAK(31-686)(K454R) substrate after 2 hrs by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 37.1nMAssay Description:Inhibition of human iNOS assessed as inhibition of [3H]L-arginine to [3H]L-citrulline conversion by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 38.9nMAssay Description:Inhibition of human iNOS assessed as inhibition of [3H]L-arginine to [3H]L-citrulline conversion by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 44.7nMAssay Description:Inhibition of human eNOS assessed as inhibition of [3H]L-arginine to [3H]L-citrulline conversion by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 45nMAssay Description:Inhibition of FAK (unknown origin) using biotinylated His-TEV-hsFAK(31-686)(K454R) substrate after 2 hrs by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 45nMAssay Description:Inhibition of FAK (unknown origin) using biotinylated His-TEV-hsFAK(31-686)(K454R) substrate after 2 hrs by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 51nMAssay Description:Inhibition of FAK (unknown origin) using biotinylated His-TEV-hsFAK(31-686)(K454R) substrate after 2 hrs by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 55.0nMAssay Description:Inhibition of human iNOS assessed as inhibition of [3H]L-arginine to [3H]L-citrulline conversion by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 57nMAssay Description:Inhibition of FAK (unknown origin) using biotinylated His-TEV-hsFAK(31-686)(K454R) substrate after 2 hrs by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 60nMAssay Description:Displacement of [3H]17AAG from human recombinant Hsp90 after 30 mins by beta scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 70.8nMAssay Description:Inhibition of human iNOS assessed as inhibition of [3H]L-arginine to [3H]L-citrulline conversion by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 81.3nMAssay Description:Inhibition of human iNOS assessed as inhibition of [3H]L-arginine to [3H]L-citrulline conversion by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 83nMAssay Description:Inhibition of FAK in human HT-29 cells assessed as phosphorylation at tyrosine 397 after 45 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 107nMAssay Description:Inhibition of FAK in human HT-29 cells assessed as phosphorylation at tyrosine 397 after 45 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:Inhibition of human iNOS assessed as inhibition of [3H]L-arginine to [3H]L-citrulline conversion by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 115nMAssay Description:Inhibition of human iNOS assessed as inhibition of [3H]L-arginine to [3H]L-citrulline conversion by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Inhibition of FAK (unknown origin) using biotinylated His-TEV-hsFAK(31-686)(K454R) substrate after 2 hrs by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 145nMAssay Description:Inhibition of human iNOS assessed as inhibition of [3H]L-arginine to [3H]L-citrulline conversion by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Displacement of [3H]17AAG from human recombinant Hsp90 after 30 mins by beta scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 155nMAssay Description:Inhibition of human iNOS assessed as inhibition of [3H]L-arginine to [3H]L-citrulline conversion by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:Inhibition of FAK (unknown origin) using biotinylated His-TEV-hsFAK(31-686)(K454R) substrate after 2 hrs by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:Displacement of [3H]17AAG from human recombinant Hsp90 after 30 mins by beta scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 177nMAssay Description:Inhibition of FAK (unknown origin) using biotinylated His-TEV-hsFAK(31-686)(K454R) substrate after 2 hrs by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 186nMAssay Description:Inhibition of human iNOS assessed as inhibition of [3H]L-arginine to [3H]L-citrulline conversion by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 195nMAssay Description:Inhibition of FAK (unknown origin) using biotinylated His-TEV-hsFAK(31-686)(K454R) substrate after 2 hrs by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:Displacement of [3H]17AAG from human recombinant Hsp90 after 30 mins by beta scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 220nMAssay Description:Inhibition of FAK (unknown origin) using biotinylated His-TEV-hsFAK(31-686)(K454R) substrate after 2 hrs by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 230nMAssay Description:Displacement of [3H]17AAG from human recombinant Hsp90 after 30 mins by beta scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 250nMAssay Description:Displacement of [3H]17AAG from human recombinant Hsp90 after 30 mins by beta scintillation countingMore data for this Ligand-Target Pair

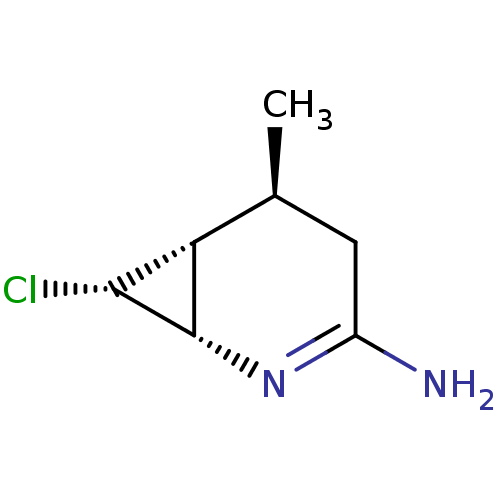
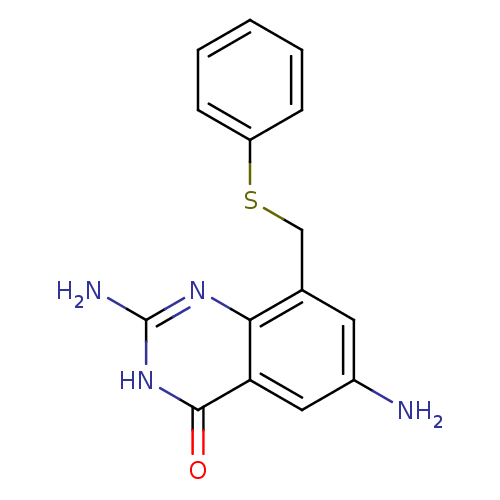
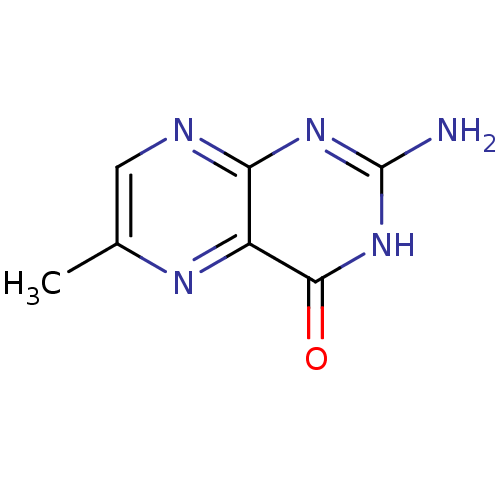
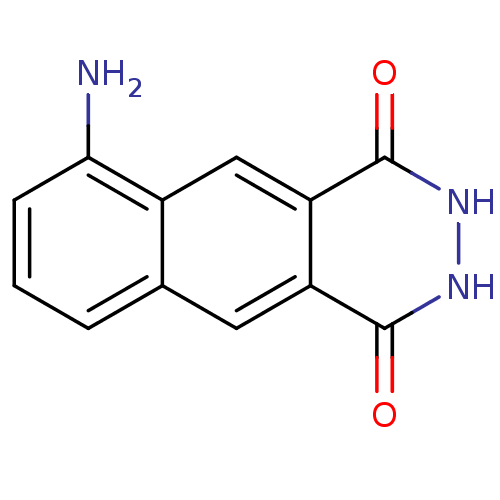
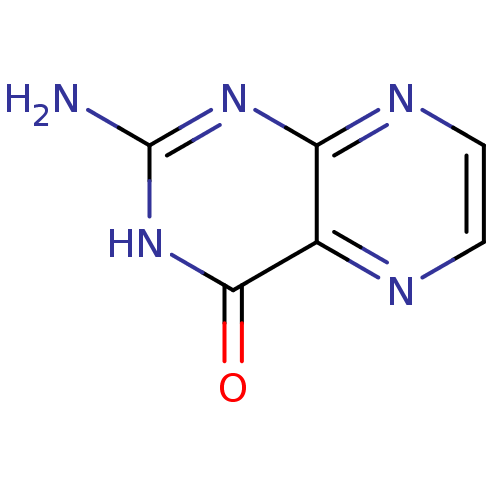
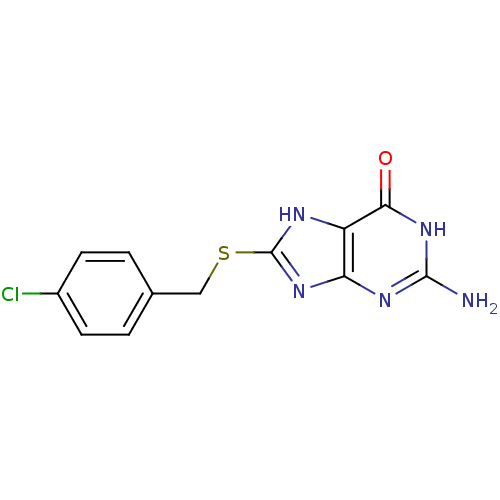
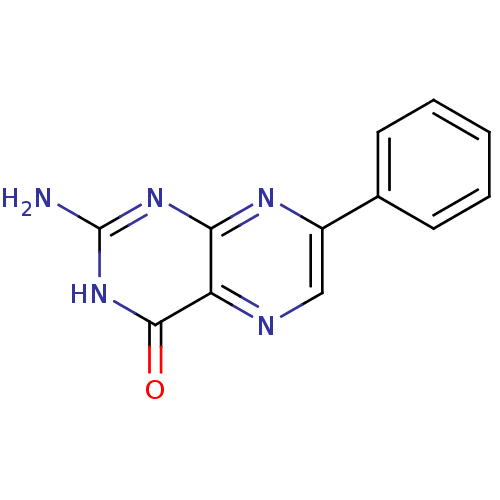
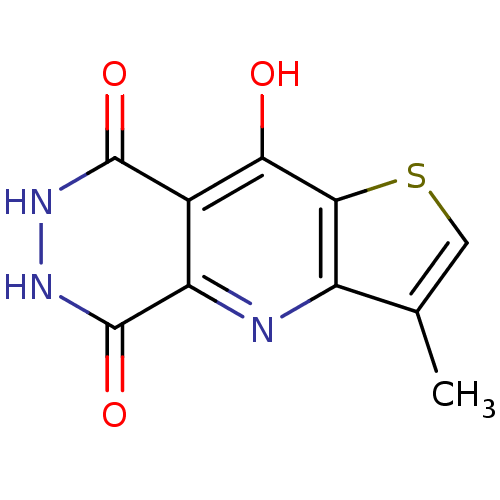
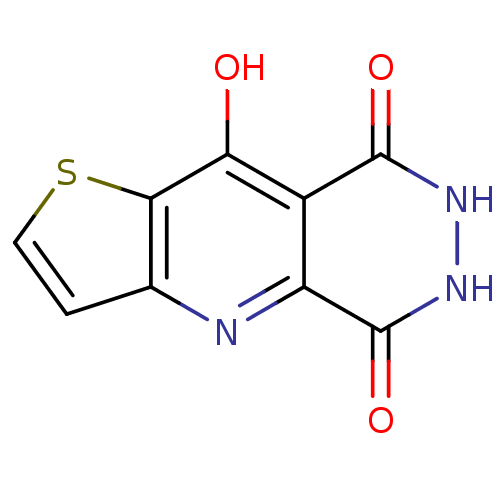
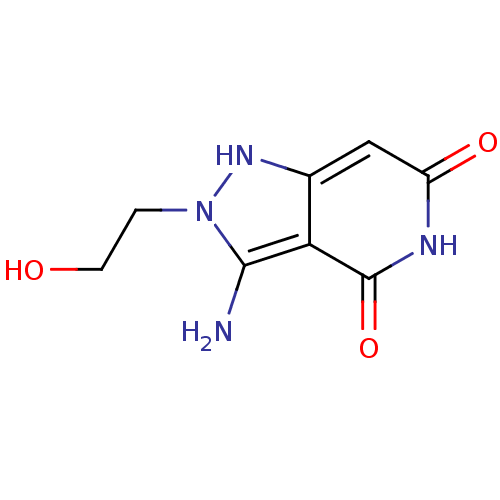
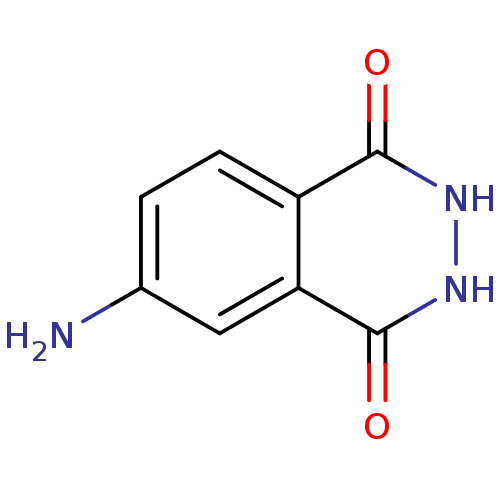
 3D Structure (crystal)
3D Structure (crystal)