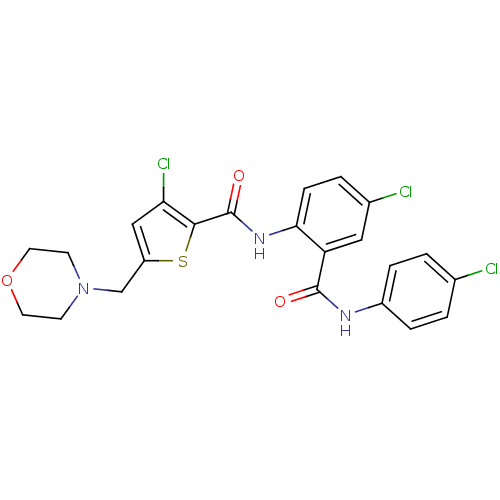
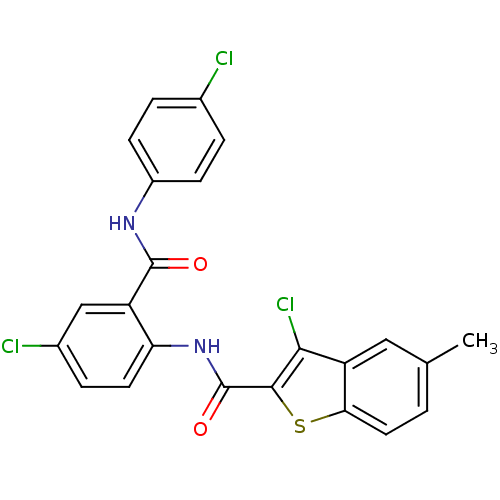
Report error Found 141 Enz. Inhib. hit(s) with all data for entry = 2130
Affinity DataKi: 0.300nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.330nM ΔG°: -53.6kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.380nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.400nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.450nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.520nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.600nM ΔG°: -52.1kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.600nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.820nM ΔG°: -51.3kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.950nM ΔG°: -51.0kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -50.9kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 1.70nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 1.90nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 2.10nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 2.10nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 2.20nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 2.70nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 3.40nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 3.5nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 4.60nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 4.80nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 5.20nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 5.60nM ΔG°: -46.6kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 5.70nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 6.20nM ΔG°: -46.4kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 7.20nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 7.5nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 9nM ΔG°: -45.5kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 9.5nM ΔG°: -45.3kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 14nM ΔG°: -44.4kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 18nM ΔG°: -43.8kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 20nM ΔG°: -43.5kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 21nM ΔG°: -43.4kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 24nM ΔG°: -43.1kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 34nM ΔG°: -42.2kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 54nM ΔG°: -41.1kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
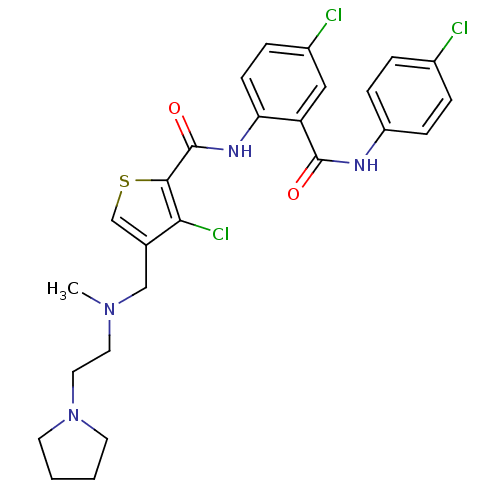
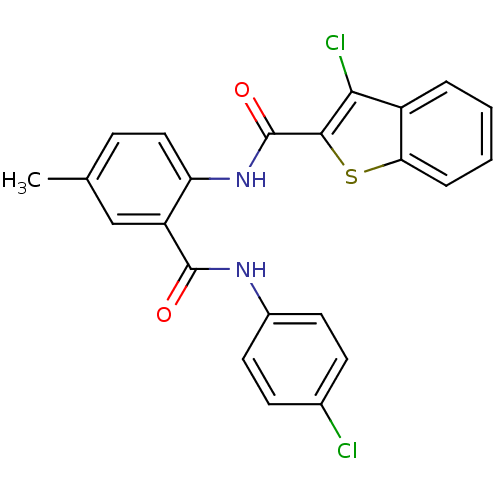
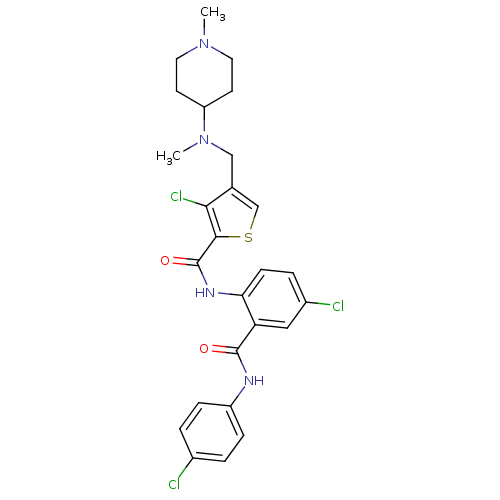
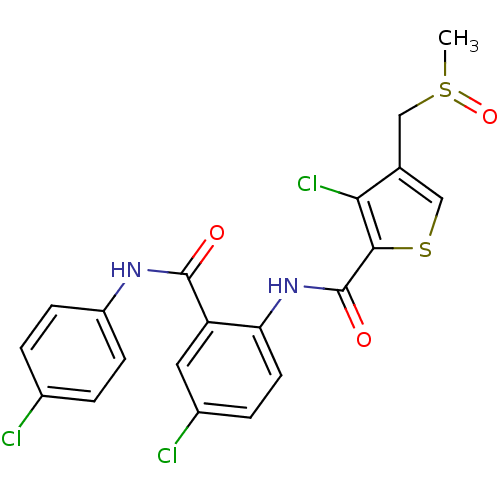
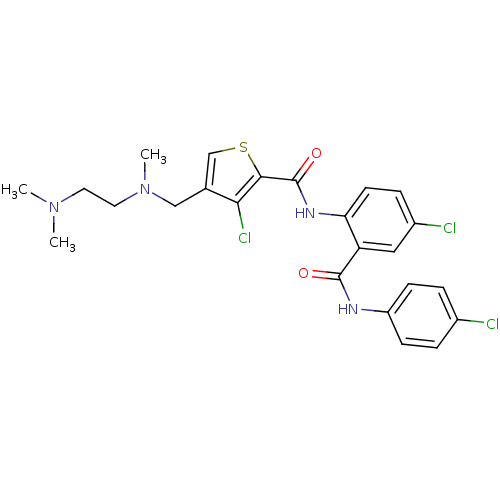
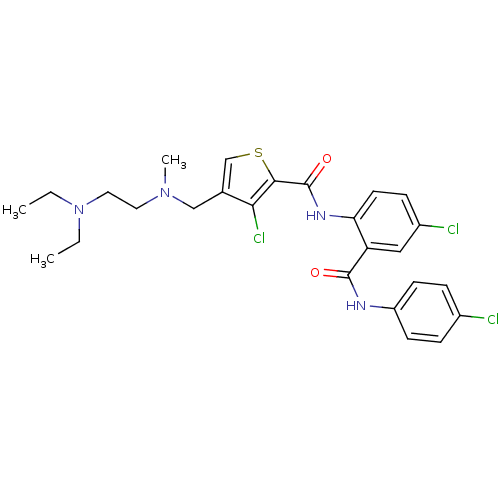

 3D Structure (crystal)
3D Structure (crystal)